 Back
BackCH. 18- Gene Regulation in Bacteria: Mechanisms and the lac Operon
Study Guide - Smart Notes

Gene Regulation in Bacteria
Overview: Gene Regulation & Information Flow
Gene regulation allows bacteria to adapt to changing environments by turning genes on or off as needed. This process is essential for cellular efficiency and survival, especially in competitive environments such as the gut. Gene expression refers to the synthesis and activation of a gene product, and its regulation impacts cellular fitness by optimizing resource use and response time.
Model organism: Escherichia coli (E. coli) is commonly used to study gene regulation.
Necessity of regulation: Expressing all genes at all times is energetically costly and can lead to cellular chaos.
Fitness outcomes: Regulation allows bacteria to compete for space and nutrients efficiently, responding to environmental signals such as nutrient availability.
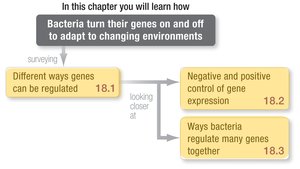
Mechanisms of Regulation: Information Flow
Information flows from DNA to RNA to protein, with regulatory controls possible at multiple steps. The main types of gene regulation are:
Transcriptional control: Determines if and when mRNA is produced from DNA.
Translational control: Regulates whether an mRNA is translated into protein.
Post-translational control: Modifies proteins after they are made, affecting their activity.

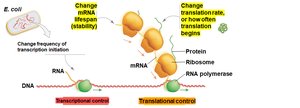
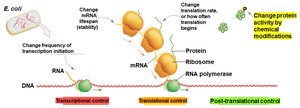
Tradeoff: Transcriptional control is most resource-efficient but slowest; post-translational control is fastest but least efficient; translational control is intermediate.
Regulated vs. constitutive expression: Regulated genes can be turned on/off or expressed at varying levels; constitutive genes are always on.
Lactose Metabolism in E. coli: A Model System
Pathway and Enzymes
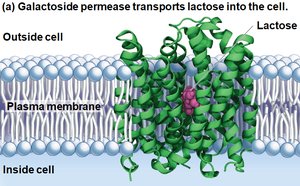
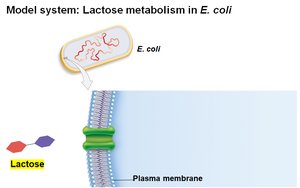
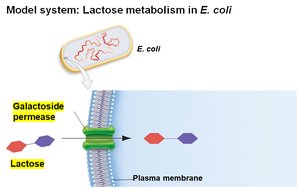
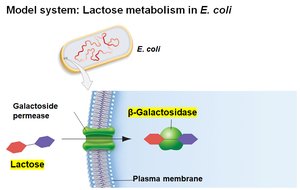
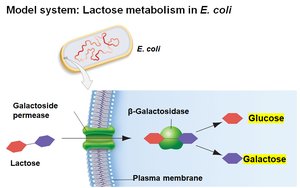
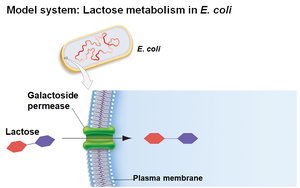
E. coli can metabolize various sugars, but prefers glucose. When glucose is depleted, it can use lactose, which requires:
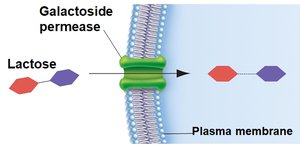
Transport of lactose into the cell by galactoside permease
Conversion of lactose to glucose and galactose by β-galactosidase
Experimental Analysis: Regulation by Glucose and Lactose
Researchers hypothesized that E. coli would not produce β-galactosidase when glucose is present, even if lactose is available. This was tested using different growth media:
Treatment 1: Glucose only
Treatment 2: Glucose + lactose
Treatment 3: Lactose only
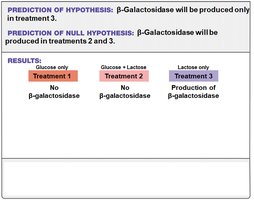
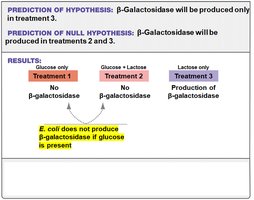
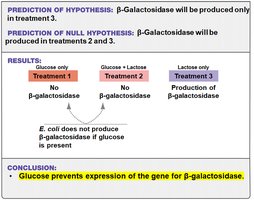
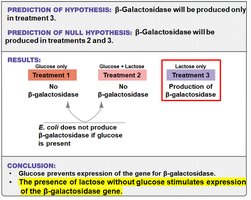

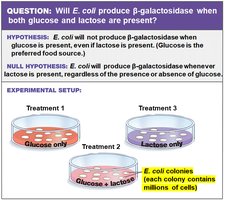
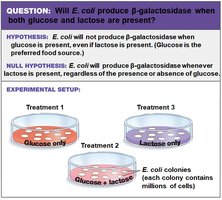





Conclusion: Glucose prevents expression of the gene for β-galactosidase. Lactose alone stimulates its expression.
Negative and Positive Control of Transcription
Mechanisms of Control
Gene expression can be regulated by two main mechanisms:
Negative control: A repressor protein binds DNA and blocks transcription.
Positive control: An activator protein binds DNA and increases transcription.
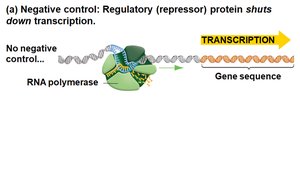
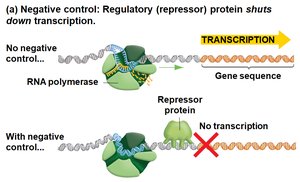
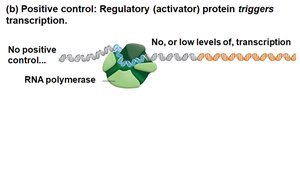
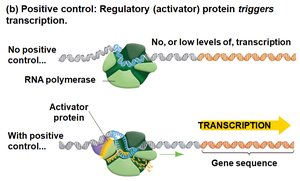
Genetic Analysis of Lactose Metabolism
Mutational analysis identified three classes of mutants affecting lactose metabolism:
Observed Phenotype | Interpretation | Genotype |
|---|---|---|
Cells cannot cleave lactose, even in the presence of inducer (lactose). | No β-galactosidase; gene for β-galactosidase is defective. | lacZ- |
Cells cannot accumulate lactose. | No membrane protein (galactoside permease) to import lactose; gene for permease is defective. | lacY- |
Cells can cleave lactose even if lactose is absent as an inducer. | Constitutive expression of lacZ and lacY; regulatory gene that shuts down both is defective. | lacI- |

The lac Operon Model
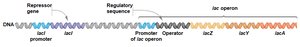
The lac operon is a set of adjacent genes (lacZ, lacY, lacA) transcribed together from a single promoter and regulated by the lacI gene product (repressor protein). The operon is controlled by the presence or absence of lactose:
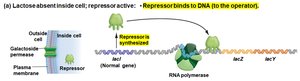
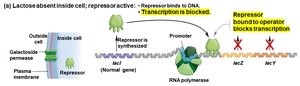
When lactose is absent: The repressor binds the operator, blocking transcription.
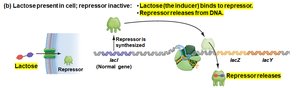
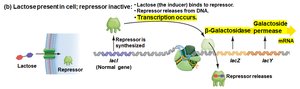
When lactose is present: Lactose binds the repressor, causing it to release from the operator, allowing transcription.
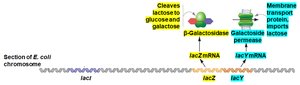
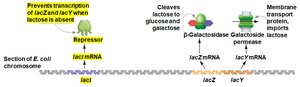
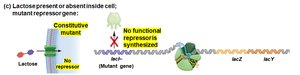
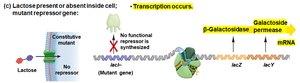
Super-repressor mutants: Repressor cannot bind inducer (lactose), so operon is always off.
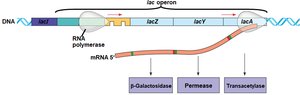
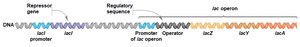
Operon Structure and Regulation
The lac operon consists of structural genes (lacZ, lacY, lacA) downstream of a single promoter and operator. The lacI gene encodes the repressor protein, which is constitutively expressed. The operon is an example of coordinated gene regulation in bacteria.
Positive Control: CAP Regulation
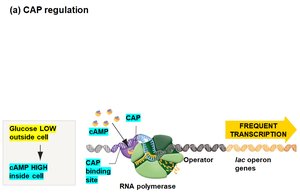
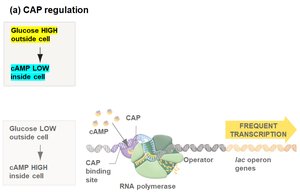
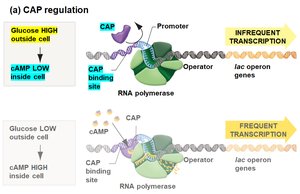
Transcription of the lac operon is also regulated by the catabolite activator protein (CAP), which requires cyclic AMP (cAMP) to bind DNA and enhance transcription. Glucose inhibits cAMP synthesis, thus reducing CAP activity and lac operon transcription.
Low glucose: High cAMP, CAP binds DNA, frequent transcription.
High glucose: Low cAMP, CAP does not bind DNA, infrequent transcription.
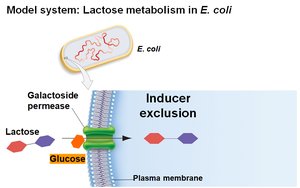
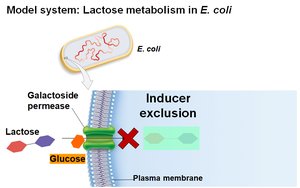
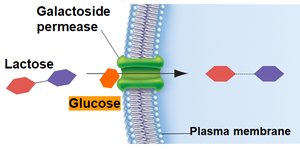
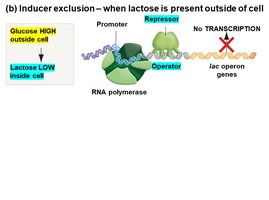
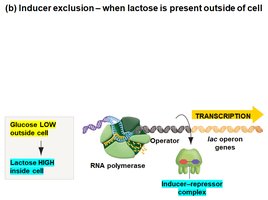
Inducer Exclusion
Glucose also regulates the lac operon by inhibiting lactose transport (inducer exclusion). When glucose is present, galactoside permease activity is reduced, preventing lactose from entering the cell and thus keeping the operon off.
Significance of the lac Operon
The lac operon is a classic example of gene regulation in bacteria, demonstrating both negative and positive control mechanisms. Many bacterial genes are regulated by repressors (negative control) and activators (positive control), with gene expression controlled by direct interactions between regulatory proteins and DNA sequences. This system also highlights the importance of post-translational control for rapid cellular responses.
Global Gene Regulation
Regulons
Global gene regulation involves the coordinated control of multiple genes or operons, often in response to environmental changes. A regulon is a set of genes or operons with the same regulatory sequences, controlled by a single type of regulatory protein. Regulation can be positive or negative.
Example: The SOS regulon, which is negatively controlled and activated in response to DNA damage.
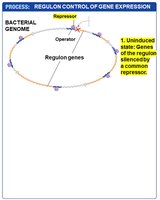
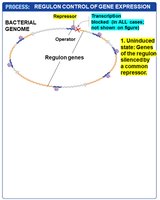
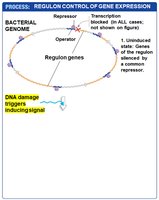
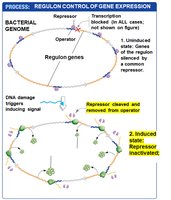
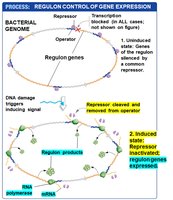
Regulons allow bacteria to rapidly adjust large sets of genes in response to environmental stress or signals.
Additional info: The lac operon and regulon models are foundational for understanding gene regulation in prokaryotes and have informed research in molecular biology, biotechnology, and medicine.
