 Back
BackGene Regulation in Bacteria: The Operon Model and Its Mechanisms
Study Guide - Smart Notes

Gene Regulation in Bacteria
Introduction to Bacterial Gene Regulation
Bacteria must efficiently regulate gene expression to adapt to changing environments and conserve resources. This regulation often occurs at the level of transcription, allowing bacteria to produce only the proteins necessary for survival under specific conditions.
Selective Advantage: Bacteria that regulate gene expression efficiently have a survival advantage.
Example: Escherichia coli (E. coli) in the human colon adjusts its metabolism based on nutrient availability, such as the presence or absence of tryptophan.
Levels of Metabolic Pathway Regulation
Regulation of Enzyme Activity and Production
Bacteria can control metabolic pathways at two main levels: by regulating the activity of existing enzymes (a rapid response) and by regulating the production of enzymes (a longer-term response).
Feedback Inhibition: The end product of a pathway (e.g., tryptophan) inhibits the activity of the first enzyme in the pathway, preventing overproduction.
Regulation of Gene Expression: Cells can repress or activate the transcription of genes encoding pathway enzymes, adjusting enzyme levels as needed.
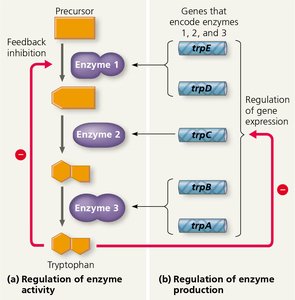
The Operon Model
Structure and Function of Operons
An operon is a cluster of functionally related genes regulated together and transcribed as a single mRNA. The operon model explains how bacteria coordinate the expression of genes involved in a common pathway.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Operator: DNA segment within or near the promoter that acts as an on-off switch, controlling RNA polymerase access to the genes.
Regulatory Gene: Encodes a repressor protein that can bind to the operator and block transcription.
Repressible and Inducible Operons
Repressible Operons: The trp Operon
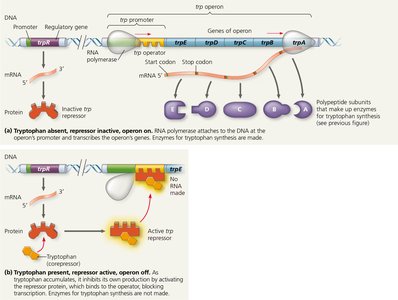
The trp operon in E. coli is a classic example of a repressible operon, which is usually active but can be turned off (repressed) when the end product (tryptophan) is abundant.
trp Repressor: Encoded by the trpR gene, this protein is inactive alone but becomes active when bound by tryptophan (the corepressor).
Mechanism: When tryptophan is present, it binds to the repressor, activating it. The active repressor binds to the operator, blocking transcription of the operon.
Feedback: This prevents unnecessary synthesis of tryptophan when it is already available.
Inducible Operons: The lac Operon
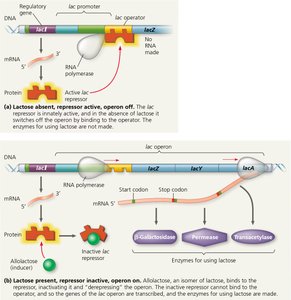
The lac operon is an example of an inducible operon, which is usually off but can be turned on (induced) in the presence of a specific substrate (lactose).
lac Repressor: Encoded by the lacI gene, this protein is active by default and binds to the operator, preventing transcription.
Inducer (Allolactose): When lactose is present, a small amount is converted to allolactose, which binds to the repressor and inactivates it, allowing transcription of the operon.
Enzymes Produced: The operon encodes enzymes for lactose metabolism, including β-galactosidase, permease, and transacetylase.
Negative and Positive Gene Regulation
Negative Regulation
Both the trp and lac operons are subject to negative regulation, where an active repressor protein binds to the operator to block transcription.
trp Operon: Repressed by the active trp repressor (with tryptophan bound).
lac Operon: Repressed by the active lac repressor (in the absence of allolactose).
Positive Regulation: The Role of CRP (CAP) in the lac Operon
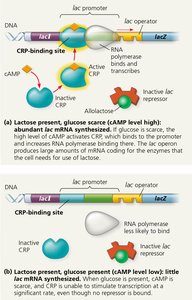
Positive regulation occurs when a regulatory protein increases transcription. In the lac operon, the cAMP receptor protein (CRP, also called CAP) acts as an activator in response to low glucose levels.
cAMP: Accumulates when glucose is scarce and binds to CRP, activating it.
Active CRP: Binds to a site near the lac promoter, increasing RNA polymerase affinity and enhancing transcription of the lac operon.
Dual Control: The lac operon is regulated both negatively (by the lac repressor) and positively (by CRP).
Summary Table: Comparison of trp and lac Operons
Feature | trp Operon | lac Operon |
|---|---|---|
Type of Regulation | Repressible (usually on) | Inducible (usually off) |
Regulatory Molecule | Tryptophan (corepressor) | Allolactose (inducer) |
Repressor Activity | Inactive alone, active with tryptophan | Active alone, inactive with allolactose |
Pathway Type | Anabolic (biosynthetic) | Catabolic (degradative) |
Positive Regulation | No | Yes (by CRP/cAMP) |
Key Terms and Concepts
Operon: A unit of genetic function found in bacteria and phages, consisting of a promoter, an operator, and a coordinately regulated cluster of genes whose products function in a common pathway.
Repressor: A protein that inhibits gene transcription by binding to the operator and blocking RNA polymerase.
Inducer: A molecule that inactivates the repressor to turn on transcription (e.g., allolactose for the lac operon).
Corepressor: A small molecule that cooperates with a repressor protein to switch an operon off (e.g., tryptophan for the trp operon).
Activator: A protein that binds to DNA and stimulates transcription (e.g., CRP for the lac operon).
Practice Questions
How does binding of the trp corepressor to the trp repressor alter repressor function and transcription? How does binding of the lac inducer alter the function of the lac repressor?
Describe the binding of repressors and activators to the lac operon, and the effect on transcription, when both lactose and glucose are scarce.
What would happen if a mutation in E. coli changed the lac operator so that the active repressor could not bind? How would this affect the cell’s production of β-galactosidase?
