 Back
BackGene Regulation in Prokaryotes and Eukaryotes: Operons, Chromatin, and Transcriptional Control
Study Guide - Smart Notes

Regulation of Gene Expression in Bacteria
Feedback Inhibition and Transcriptional Regulation
Prokaryotic cells, such as bacteria, regulate gene expression to adapt to environmental changes and conserve resources. Two primary mechanisms are feedback inhibition and regulation of gene transcription.
Feedback inhibition: The end product of a metabolic pathway inhibits an enzyme involved earlier in the pathway, preventing overproduction.
Transcriptional regulation: Cells adjust enzyme production by controlling the expression of genes encoding those enzymes.
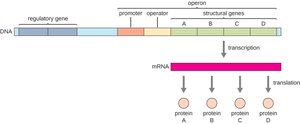
Operons: Coordinated Gene Regulation
In bacteria, a group of functionally related genes can be regulated together as a unit called an operon. This allows efficient and coordinated control of gene expression.
Operator: A DNA segment acting as an "on-off switch," located within or near the promoter.
Promoter: The DNA region where RNA polymerase binds to initiate transcription.
Structural genes: Genes encoding proteins with related functions, transcribed together as a single mRNA.
Repressor: A protein that binds to the operator to block transcription by RNA polymerase.
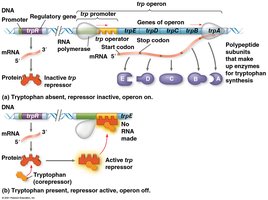
The trp Operon: A Repressible System
The trp operon in Escherichia coli controls the synthesis of tryptophan, an essential amino acid. It is an example of a repressible operon, typically active but can be turned off when tryptophan is abundant.
When tryptophan levels are high, tryptophan acts as a corepressor, activating the repressor protein, which binds to the operator and blocks transcription.
When tryptophan is scarce, the repressor is inactive, and the operon is transcribed, allowing tryptophan synthesis.
Repressible operons usually regulate anabolic (biosynthetic) pathways.
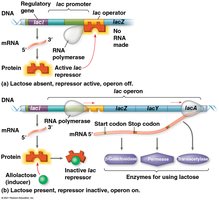
The lac Operon: An Inducible System
The lac operon enables bacteria to metabolize lactose when it is available. It is an inducible operon, usually off but can be turned on in the presence of an inducer.
The lac repressor is active by default, keeping the operon off.
When lactose is present, its isomer allolactose acts as an inducer, inactivating the repressor and allowing transcription.
A regulatory gene (lacI) encodes the repressor protein.
Inducible operons typically regulate catabolic (degradative) pathways.
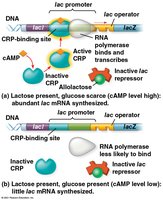
Negative and Positive Control of Operons
Both the trp and lac operons are subject to negative control (switched off by active repressors). Some operons, such as the lac operon, are also regulated by positive control mechanisms.
cAMP-CRP complex: When glucose is low, cyclic AMP (cAMP) binds to the cAMP receptor protein (CRP), which then binds to the promoter, enhancing transcription of the lac operon.
When glucose is abundant, cAMP levels drop, CRP detaches, and transcription decreases.
Gene Regulation in Eukaryotes
Chromatin Structure and Epigenetic Regulation
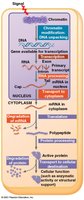
In eukaryotes, gene expression is regulated at multiple levels, including chromatin structure, transcription, RNA processing, translation, and protein modification.
Heterochromatin: Densely packed chromatin, generally transcriptionally inactive.
Euchromatin: Loosely packed chromatin, accessible for transcription.
Histone acetylation: Addition of acetyl groups to histone tails opens chromatin, promoting transcription.
DNA methylation: Addition of methyl groups condenses chromatin, reducing transcription; can cause long-term gene silencing.
Post-Transcriptional Regulation
Gene expression can also be regulated after transcription through alternative RNA splicing, mRNA stability, and translational control.
Alternative RNA splicing: Produces different mRNAs (and thus proteins) from the same gene by varying exon/intron inclusion.
mRNA stability: The lifespan of mRNA in the cytoplasm affects protein synthesis; eukaryotic mRNAs are generally more stable than prokaryotic mRNAs.
UTR sequences: Untranslated regions at the 3' end of mRNA influence stability and translation efficiency.
Protein degradation: Proteins are marked for degradation by ubiquitin and destroyed by proteasomes.
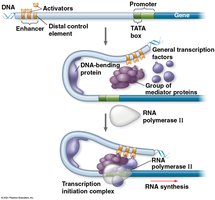
Transcriptional Control in Eukaryotes
Transcription of eukaryotic genes requires the assembly of a complex of proteins at the promoter, including general and specific transcription factors.
General transcription factors: Required for transcription of all protein-coding genes; help RNA polymerase II bind to the promoter.
Enhancers: Distal control elements that bind activator proteins, increasing transcription rates.
Mediator proteins: Facilitate interactions between activators, repressors, and the transcription machinery.
Repressors: Inhibit gene expression by blocking activator binding or interfering with the transcription complex.
Summary Table: Comparison of trp and lac Operons
Feature | trp Operon | lac Operon |
|---|---|---|
Type | Repressible | Inducible |
Default State | On | Off |
Regulation | Turned off by corepressor (tryptophan) | Turned on by inducer (allolactose) |
Pathway Type | Anabolic (biosynthetic) | Catabolic (degradative) |
