 Back
BackChapter 14
Study Guide - Smart Notes

Gene Regulation
Overview of Gene Regulation
Gene regulation is the process by which cells control the expression and timing of gene activity, ensuring proteins are produced at the correct time and in appropriate amounts. This regulation is essential for cellular efficiency and specialization.
Constitutive genes are unregulated and expressed at constant levels, providing essential cellular functions.
Regulated genes are expressed only when needed, conserving energy and resources.
Gene Regulation in Eukaryotes
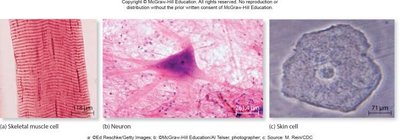
In multicellular eukaryotes, gene regulation is crucial for cell differentiation, allowing cells with identical genomes to develop into specialized types by expressing different sets of proteins (proteomes).
Different cell types (e.g., muscle, neuron, skin) arise from differential gene expression.
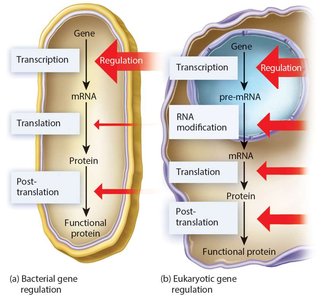
Levels of Gene Regulation
Gene expression in eukaryotes can be regulated at multiple stages:
Transcription
RNA processing (e.g., splicing, capping, polyadenylation)
Translation
Post-translation (protein modification and degradation)
Additional eukaryotic controls: chromatin remodeling, mRNA stability
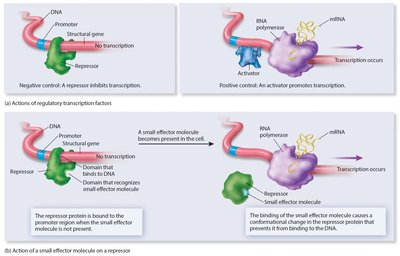
Regulation of Transcription in Eukaryotes
Transcriptional regulation involves activators, repressors, and complex interactions with DNA and chromatin structure.
Activator proteins stimulate RNA polymerase to initiate transcription.
Repressor proteins inhibit RNA polymerase activity.
Small effector molecules, protein-protein interactions, and covalent modifications modulate these regulators.
Chromatin remodeling and DNA methylation further influence gene accessibility.
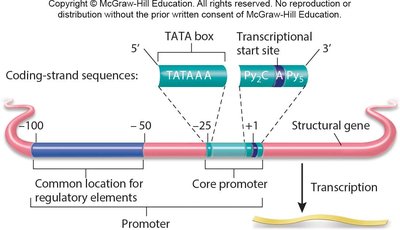
Promoter Structure and Regulatory Elements
Most eukaryotic promoters contain three key features:
TATA box: A conserved DNA sequence (5’–TATAAAA–3’) about 25 base pairs upstream of the transcription start site, determining where transcription begins.
Transcriptional start site: The location where RNA synthesis initiates.
Regulatory/response elements: DNA sequences recognized by regulatory proteins (e.g., enhancers and silencers) that modulate transcription initiation.
Transcription Initiation Complex
Transcription initiation in eukaryotes requires:
RNA polymerase II
Five general transcription factors (GTFs)
A large protein complex called mediator, which facilitates interactions between transcription factors and regulatory elements, controlling the rate of transcription initiation.
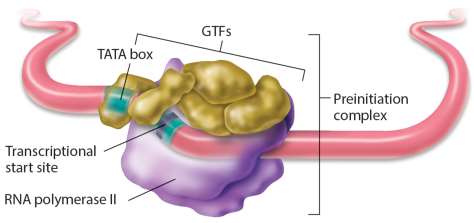
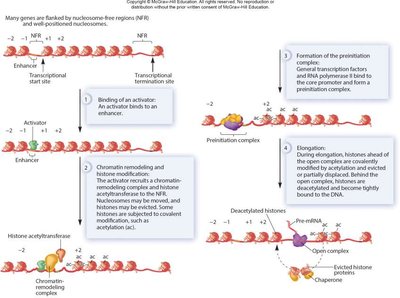
Mechanisms of Transcriptional Regulation
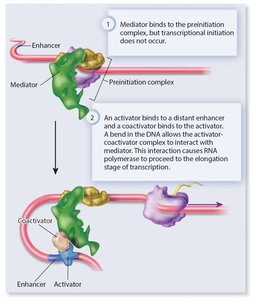
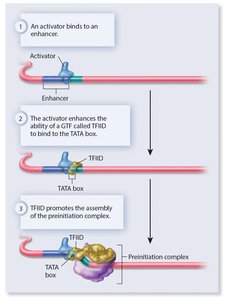
Activators and repressors can regulate RNA polymerase II by binding to GTFs (such as TFIID) or by interacting with the mediator complex.
Regulatory proteins can also recruit chromatin-modifying enzymes to alter DNA accessibility.
Chromatin Structure and Gene Regulation
In eukaryotes, DNA is packaged into chromatin, which can exist in open (accessible) or closed (inaccessible) conformations, affecting gene expression.
Transcription is generally repressed in tightly packed chromatin and facilitated in loosely packed chromatin.
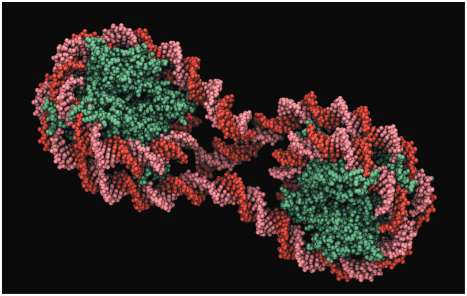
Chromatin Remodeling
Chromatin remodeling complexes can reposition, evict, or replace nucleosomes, altering DNA accessibility.
Histone acetyltransferases (HATs) add acetyl groups to histones, loosening DNA-histone interactions and promoting transcription.
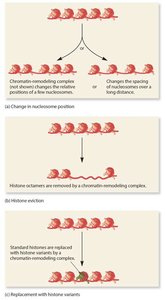
Nucleosome-Free Regions (NFRs)
NFRs are typically found at the beginning and end of genes, facilitating access for transcription machinery.

DNA Methylation
DNA methylation (addition of –CH3 groups to cytosines in CpG islands) generally represses transcription by condensing chromatin or blocking activator binding.
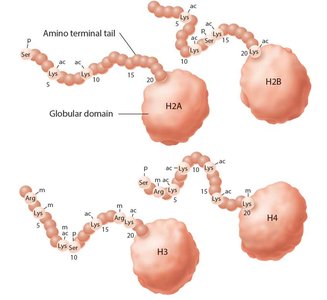
Histone Modification and the Histone Code
Histone modifications (acetylation, methylation, phosphorylation) influence chromatin structure and gene expression.
The "histone code" refers to specific combinations of modifications that determine chromatin state and gene activity.
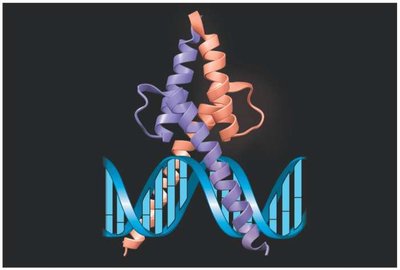
Transcription Factor-DNA Recognition
Transcription factors recognize specific DNA sequences by interacting with exposed bases in the major and minor grooves of the DNA double helix.
Regulation of RNA Processing and Translation
Gene expression in eukaryotes is also regulated at the level of RNA processing and translation, allowing for rapid and diverse protein production.
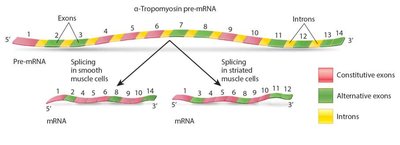
Alternative splicing enables a single gene to produce multiple mRNA variants and thus different proteins, increasing proteome diversity without increasing genome size.
Mutation, DNA Repair, and Cancer
Mutation
A mutation is a heritable change in the genetic material, serving as a source of genetic variation but often being harmful. DNA repair systems exist to correct mutations, and failures in these systems can lead to diseases such as cancer.
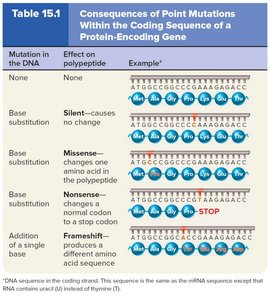
Types of Point Mutations
Silent mutation: No change in amino acid sequence.
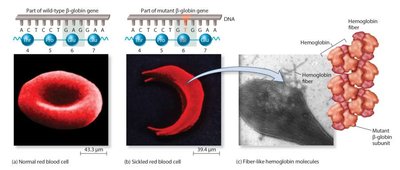
Missense mutation: Changes one amino acid; may or may not affect protein function (e.g., sickle-cell disease).
Nonsense mutation: Converts a codon to a stop codon, truncating the protein.
Frameshift mutation: Insertion or deletion (not in multiples of three) alters downstream amino acid sequence.
Mutations Outside Coding Sequences
Mutations in promoters, regulatory elements, splice sites, or untranslated regions can affect gene expression, mRNA processing, or translation efficiency.
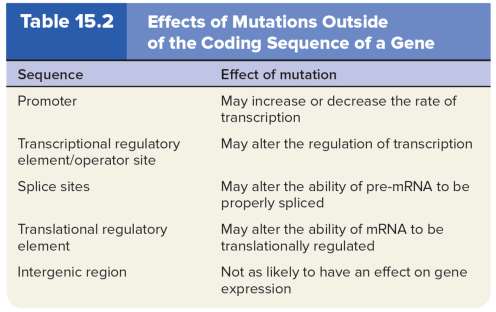

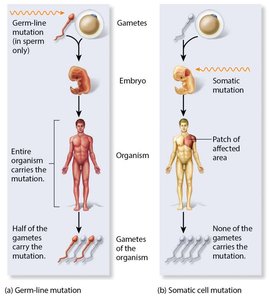
Germ-Line vs. Somatic Mutations
Germ-line mutations occur in gametes and are heritable.
Somatic mutations occur in body cells and can lead to genetic mosaics but are not passed to offspring.
Causes of Mutation
Spontaneous mutations: Result from natural biological processes (e.g., DNA replication errors, metabolic byproducts).
Induced mutations: Caused by environmental agents (mutagens) such as chemicals or radiation.

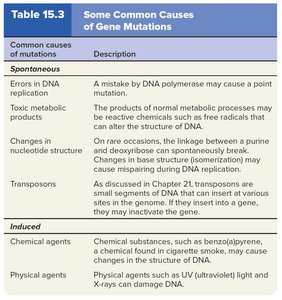
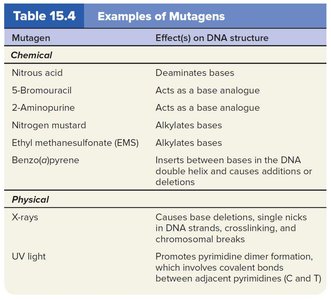
Mechanisms of Mutagenesis
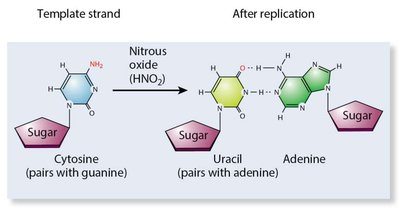
Chemical mutagens can deaminate bases, alkylate nucleotides, or insert base analogues.
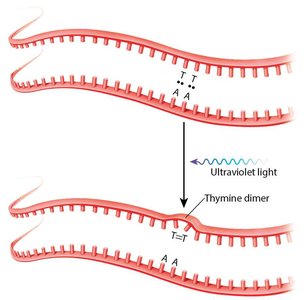
Physical mutagens such as X-rays and UV light can cause strand breaks or thymine dimers, respectively.
DNA Repair Mechanisms
Direct repair: Enzyme directly reverses the DNA damage.
Nucleotide excision repair (NER): Damaged DNA segment is removed and replaced using the undamaged strand as a template.
Methyl-directed mismatch repair: Mismatched bases are recognized, and the incorrect DNA segment is excised and replaced.
Cancer: Genetic Basis and Progression
Cancer is a disease of uncontrolled cell division, often resulting from the accumulation of multiple genetic mutations. Most cancers are not inherited but arise from somatic mutations, often due to exposure to carcinogens (mutagens that increase cancer risk).
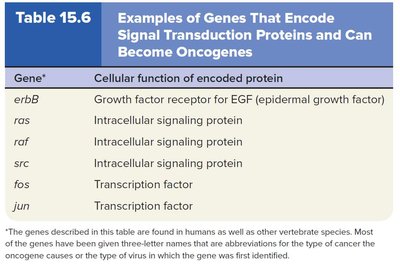
Oncogenes and Proto-Oncogenes
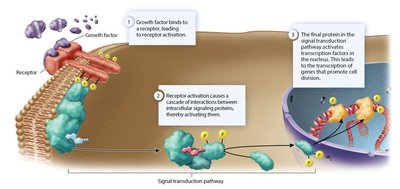
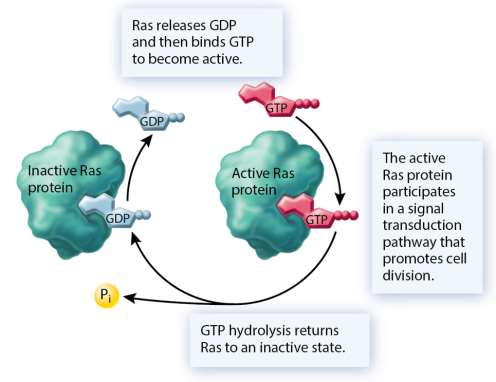
Proto-oncogenes are normal genes that promote cell division; mutations can convert them into oncogenes that drive cancer by causing excessive or unregulated cell proliferation.
Common mechanisms: missense mutations, gene amplifications, chromosomal translocations, retroviral insertions.
Tumor-Suppressor Genes
These genes normally inhibit cell division or promote DNA repair. Loss of their function removes critical cell cycle checkpoints, increasing cancer risk.
Examples: p53 (G1 checkpoint, apoptosis), Rb (retinoblastoma protein, inhibits E2F transcription factor).

Genetic Changes in Cancer
Cancer typically involves multiple genetic alterations, including mutations, chromosomal abnormalities, and epigenetic changes (e.g., abnormal methylation).
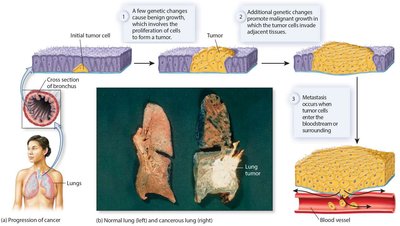
Additional info: Cancer progression is a multistep process, often beginning with benign growths that accumulate further mutations, eventually leading to malignancy and metastasis.
