 Back
BackCH: 18.1 Genomes and Their Evolution: Sequencing, Bioinformatics, and Comparative Genomics
Study Guide - Smart Notes

Genomes and Their Evolution
Introduction to Genomics and Genome Evolution
Genomics is the comprehensive study of whole sets of genes and their interactions within an organism. This field provides critical insights into gene organization, regulation, expression, development, and evolutionary processes. The evolution of genomes is driven by mechanisms such as duplication, rearrangement, and mutation of DNA, and comparative genomics offers clues to both evolutionary history and developmental biology.
Genomics: The study of the structure, function, evolution, and mapping of genomes.
Bioinformatics: The application of computational tools to manage, analyze, and interpret biological data, especially large-scale genomic data.
Genome Evolution: Changes in genome structure and content over time, influenced by mutation, natural selection, genetic drift, and recombination.
Homology: Similarity in sequence or structure due to shared ancestry, used to infer evolutionary relationships.
The Human Genome Project and Advances in Sequencing
The Human Genome Project (HGP)
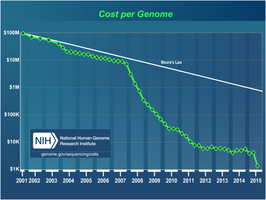
The Human Genome Project was a landmark international scientific effort to sequence the entire human genome, comprising approximately 3 billion base pairs. Initiated in 1990 and largely completed by 2003, the project fostered the development of faster and less expensive sequencing technologies, revolutionizing biological research and medicine.
Goals: Sequence the entire human genome and identify all human genes.
Approaches: Government-led effort and private initiatives (e.g., Celera Genomics).
Outcomes: Completion of the human genome sequence, development of new sequencing methods, and the foundation for personalized medicine.


Sequencing Techniques
Sequencing DNA involves several key steps, including fragmenting the DNA, cloning fragments, sequencing each fragment, and assembling the sequences using computational tools. The whole-genome shotgun approach, pioneered by J. Craig Venter, involves sequencing random DNA fragments and using software to assemble the genome.
Fragmentation: DNA is cut into overlapping fragments short enough for sequencing.
Cloning: Fragments are cloned into plasmids or vectors.
Sequencing: Each fragment is sequenced individually.
Assembly: Computer software orders the sequences into a complete genome.
Newer techniques, such as sequencing by synthesis, have dramatically increased speed and reduced costs, allowing direct sequencing without cloning.
Impact and Applications of Genome Sequencing
Genome sequencing provides a blueprint for understanding human biology, disease susceptibility, and drug metabolism. It enables personalized medicine and comparative studies with other organisms.
Comparative Genomics: Studying genome sequences across species to understand evolution and development.
Personalized Medicine: Using individual genome data to tailor medical treatments.



Metagenomics
Metagenomics is the study of genetic material recovered directly from environmental samples. This approach allows scientists to analyze the genomes of entire communities, such as those in the Sargasso Sea or the human gut, without the need to culture individual species.
Environmental DNA: DNA is extracted from a mixed community sample.
Sequencing and Assembly: Computer software sorts and assembles partial sequences into specific genomes.
Applications: Understanding microbial diversity, ecology, and function in complex environments.
Bioinformatics and Functional Genomics
Centralized Resources for Genome Analysis
Bioinformatics resources are essential for storing, searching, and analyzing the vast amounts of genomic data generated. Major databases and organizations include:
NCBI (National Center for Biotechnology Information): Provides GenBank, a comprehensive database of DNA sequences.
European Molecular Biology Laboratory and DNA Data Bank of Japan: International partners in genomic data sharing.
BGI (Beijing Genomics Institute): Major sequencing center in China.
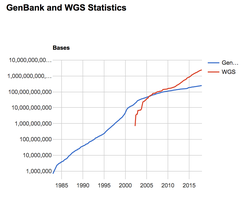
GenBank and Sequence Analysis
GenBank is a rapidly growing database, doubling in size approximately every 18 months. Online tools allow users to search for DNA or protein sequence matches, identify conserved regions, and visualize protein structures.
Sequence Alignment: Comparing DNA or protein sequences to identify similarities and infer function or evolutionary relationships.
Protein Structure: 3D models help understand protein function and interactions.
Identifying Gene Functions
Scientists deduce the function of protein-coding genes by comparing predicted amino acid sequences with known proteins, using both sequence similarity and experimental studies.
Homology-based Prediction: Similar sequences often indicate similar functions.
Biochemical Studies: Laboratory experiments confirm predicted functions.
ENCODE and Systems Biology
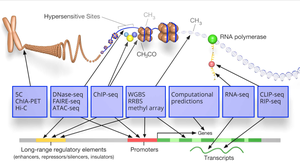
The ENCODE (Encyclopedia of DNA Elements) project has mapped protein-coding genes, noncoding RNAs, and regulatory sequences, providing a comprehensive view of genome function. Systems biology integrates data from genomics, proteomics, and other fields to model the dynamic behavior of biological systems.
Proteomics: Study of the full set of proteins (proteome) encoded by a genome.
Systems Biology: Modeling interactions among genes, proteins, and other molecules to understand cellular and organismal function.
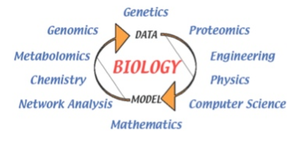
Medical Applications of Systems Biology
Systems biology approaches are used in medicine, such as The Cancer Genome Atlas project, which compares gene sequences and expression in cancer versus normal cells to identify mutations and potential therapeutic targets. Microarray technology enables the simultaneous analysis of thousands of genes.
Genome Size, Gene Number, and Gene Density
Variation in Genome Size
Genome sizes vary widely among organisms. Bacteria and archaea typically have genomes ranging from 1 to 6 million base pairs (Mb), while eukaryotes, especially plants and animals, have much larger genomes. However, there is no consistent relationship between genome size and organismal complexity (phenotype).
Bacteria/Archaea: 1–6 Mb
Plants/Animals: Often >100 Mb; humans have ~3,000 Mb
No direct correlation: Genome size does not predict gene number or complexity.
Gene Number and Density
The number of genes is not directly proportional to genome size. For example, the nematode C. elegans has 100 Mb and 20,100 genes, while humans have 3,000 Mb and about 21,000 genes. Humans and other mammals have low gene density, with large amounts of noncoding DNA and many introns within genes. Alternative splicing allows a single gene to produce multiple polypeptides, increasing protein diversity without increasing gene number.
Gene Density: Number of genes per unit length of DNA; lowest in mammals.
Noncoding DNA: Includes introns and intergenic regions; plays roles in regulation and genome structure.
Alternative Splicing: Mechanism by which different proteins are produced from the same gene.
