 Back
BackMutation and Molecular Evolution: Genes, Genomes, and Evolutionary Analysis
Study Guide - Smart Notes

Mutation and Molecular Evolution
Introduction to Genes and Genomes
The genome comprises the full set of genes and noncoding regions of DNA, which together determine the genetic makeup and regulatory complexity of an organism. In eukaryotes, most genes are located on chromosomes, but some are found in mitochondria and chloroplasts, and these are inherited through the egg cytoplasm in sexually reproducing organisms. - Genome: The complete set of genetic material, including coding and noncoding DNA. - Noncoding DNA: DNA sequences that do not code for proteins but may have regulatory or structural functions. - Organelle genomes: Mitochondrial and chloroplast DNA, inherited maternally. 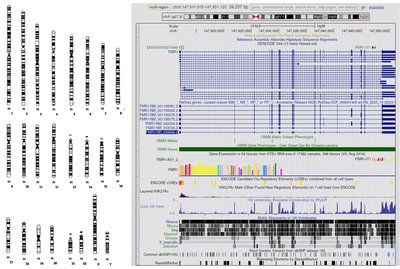
Case Study: The Orangutan
The orangutan genome illustrates the complexity and variability of genetic material. Chromosome analysis revealed differences between Sumatran and Bornean populations, particularly in chromosome 2, leading to the recognition of distinct species. - Chromosomal inversion: A segment of a chromosome is reversed end to end, affecting genetic diversity and speciation. - Homozygosity and heterozygosity: Wild orangutans are homozygous for inversion morphs, while zoo-born individuals can be heterozygous. - Conservation implications: Management of breeding in zoos to preserve genetic integrity. 

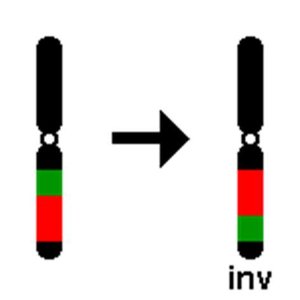

Mutation and Mutagenesis
Types and Sources of Mutation
Mutations are changes in the DNA sequence that drive molecular evolution. They can arise from endogenous sources (internal cellular processes) or exogenous sources (external environmental factors). - Endogenous mutations: Replication errors, DNA base mismatches, topoisomerase-DNA complexes, base deamination, oxidative damage, DNA methylation. - Exogenous mutations: Ionizing radiation, UV damage, alkylation, aromatic compounds, toxins (e.g., aflatoxin B1). - Impact: Mutations can alter protein structure and function, potentially affecting organismal fitness. 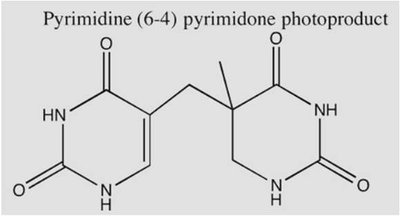
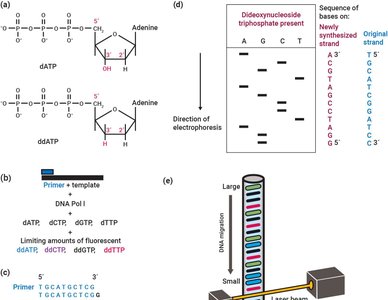
Sequencing DNA and Analyzing Genetic Material
Sanger and Illumina Sequencing
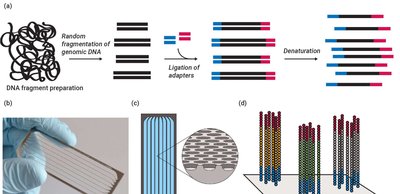
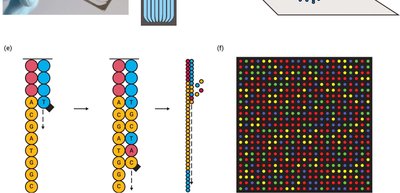
DNA sequencing technologies allow for the reading and comparison of genetic material, facilitating evolutionary studies. - Sanger sequencing: Uses dideoxynucleotides to terminate DNA synthesis, enabling sequence determination by electrophoresis. - Illumina sequencing: High-throughput method involving DNA fragmentation, adapter ligation, and parallel sequencing. - Applications: Sequence comparison, mutation detection, evolutionary analysis. 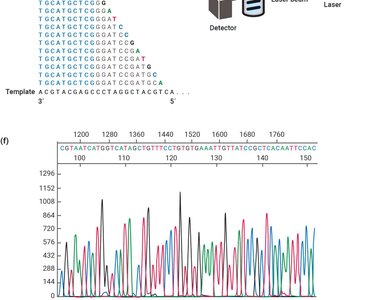
Substitutions and Sequence Comparison
Sequence comparison between species reveals the minimum number of differences, but actual substitution events may be underestimated due to multiple, coincident, parallel, or back substitutions. - Substitution types: Single, multiple, coincident, parallel, and back (reversion) substitutions. - Sequence alignment: Essential for identifying homologous positions and evolutionary relationships. 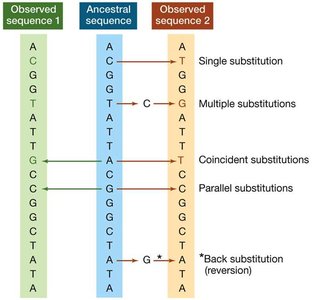
Types of DNA Mutations
Transitions and Transversions
DNA mutations can be classified as transitions (purine to purine or pyrimidine to pyrimidine) or transversions (purine to pyrimidine or vice versa). Transitions are more frequent, with a typical rate of 2:1 or 3:1 compared to transversions. - Purines: Adenine (A) and Guanine (G) - Pyrimidines: Thymine (T) and Cytosine (C) - Transition: R for R or Y for Y - Transversion: R for Y or Y for R 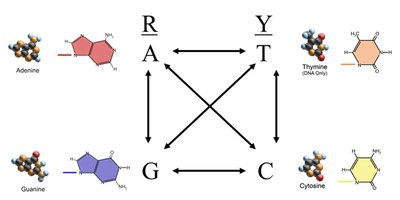
Genomes and Evolutionary Analysis
Homology and Sequence Alignment
Homologous features are inherited from a common ancestor. Sequence alignment is used to compare genes and proteins, identifying deletions, insertions, and constructing similarity matrices. - Homology: Shared ancestry of genes or proteins. - Sequence alignment: Computer algorithms align long sequences and generate similarity matrices. 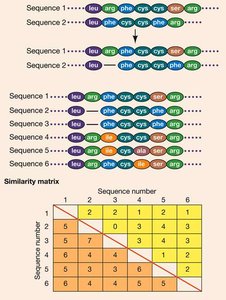
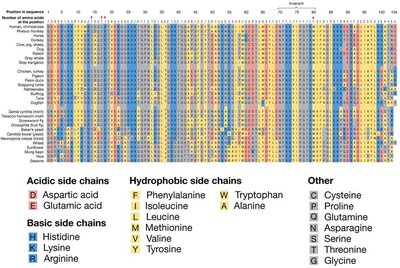
Synonymous and Nonsynonymous Substitutions
Definitions and Evolutionary Implications
Substitutions in protein-coding regions can be synonymous (silent, no amino acid change) or nonsynonymous (missense, amino acid change). Synonymous substitutions occur more rapidly and are often selectively neutral, while nonsynonymous substitutions may be deleterious or neutral. - Synonymous substitution: No change in amino acid, often neutral. - Nonsynonymous substitution: Change in amino acid, may affect protein function. 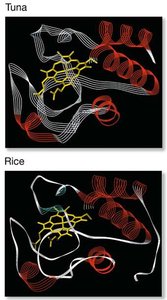
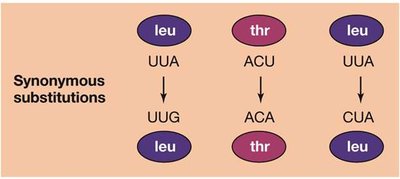
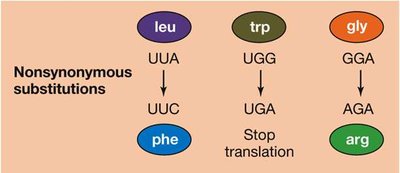
Distinguishing Evolutionary Processes
Comparative Genomics and Selection
Comparing gene sequences among species reveals the history and timing of substitutions. Lysozyme gene analysis in foregut fermenters and non-fermenters illustrates different modes of selection and evolutionary convergence. - Gene sequence comparison: Determines rates of synonymous and nonsynonymous substitutions. - Evolutionary convergence: Similar adaptations in unrelated lineages. 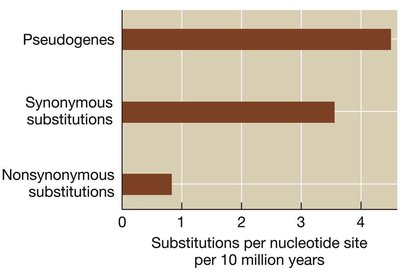
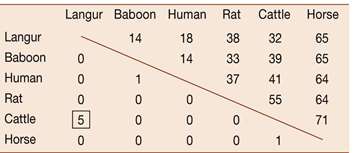


Experimental Evolution
Studying Evolution in Microorganisms
Experimental evolution uses organisms with short generation times, such as bacteria and viruses, to study adaptive radiations and substitution rates. The bacterium Pseudomonas fluorescens was used to test the hypothesis that heterogeneous environments promote phenotypic diversity. - Experimental evolution: Laboratory studies of evolutionary processes. - Adaptive radiation: Diversification in response to environmental heterogeneity. 
Genome Size and Complexity
Variation and Noncoding Elements
Genome size varies greatly among organisms, with some correlation to complexity. However, when only protein and RNA coding regions are considered, variation is reduced. Noncoding elements include satellite DNA, retroviral insertions, retrotransposons, and lateral gene transfers. - Genome size: Total DNA content, variable across taxa. - Noncoding elements: Satellite DNA, retrotransposons (LINES and SINES), retroviral insertions, lateral gene transfers. 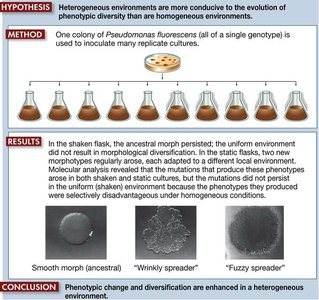
Gene Families and Evolution
Gene Duplication and Divergence
Successive rounds of duplication and mutation can result in gene families, such as the globin gene family. Hemoglobin and myoglobin illustrate functional divergence, with hemoglobin carrying O2, CO2, and H+, and myoglobin storing O2 in muscle tissue. - Gene family: Group of related genes arising from duplication events. - Divergence rate: About one substitution every two million years.
Concerted Evolution
In some gene families, genes evolve together (concerted evolution), as seen in ribosomal operons, tRNAs, and histone genes.
Gene Conversion and Evolutionary Relationships
Biased and Unbiased Gene Conversion
Gene conversion can be biased (favoring certain sequences) or unbiased (substitutions fixed or lost randomly).
Gene Trees and Homology
Gene trees illustrate evolutionary relationships among gene family members. - Orthologs: Genes related by speciation events. - Paralogs: Genes related by duplication events.
Term | Definition |
|---|---|
Genome | Complete set of genetic material |
Mutation | Change in DNA sequence |
Transition | Purine to purine or pyrimidine to pyrimidine |
Transversion | Purine to pyrimidine or vice versa |
Synonymous substitution | Silent, no amino acid change |
Nonsynonymous substitution | Missense, amino acid change |
Gene family | Group of related genes from duplication |
Ortholog | Gene related by speciation |
Paralog | Gene related by duplication |
Additional info: Some explanations and definitions were expanded for clarity and completeness, and tables were reconstructed for academic context.
