 Back
BackChapter 11 Study Guide
Study Guide - Smart Notes

Biochemical Identification of the Genetic Material
Criteria for Genetic Material
Genetic material must fulfill four essential criteria: information storage, replication, transmission, and variation. Early researchers hypothesized that chromosomes carried genetic information, but the molecular identity was unclear until the mid-20th century.
Information: Must contain instructions for the structure and function of an organism.
Replication: Must be accurately copied for transmission to offspring.
Transmission: Must be passed from parent to offspring and between cells during division.
Variation: Must be capable of change to account for genetic diversity.
Griffith’s Bacterial Transformation
Frederick Griffith's experiments with Streptococcus pneumoniae in the 1920s demonstrated the phenomenon of transformation, where genetic material from dead bacteria could transfer to live bacteria, conferring new traits.
S strain (smooth): Secretes a capsule, virulent (kills mice).
R strain (rough): No capsule, non-virulent (mice survive).
Heat-killed S strain: Non-virulent.
Live R + heat-killed S: Mice die; live S bacteria recovered, indicating transformation.
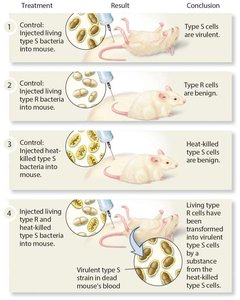
Conclusion: A 'transforming principle' from the dead S strain converted R strain into virulent S type.
Avery, MacLeod, and McCarty’s Experiment
In the 1940s, Avery, MacLeod, and McCarty identified DNA as the transforming principle by systematically eliminating proteins and RNA as candidates. Only DNase (which degrades DNA) prevented transformation, confirming DNA as the genetic material.
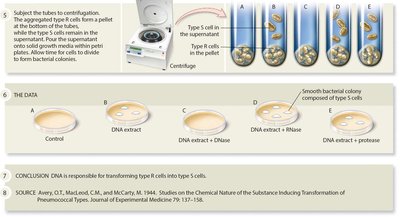
Conclusion: DNA is responsible for transforming R cells into S cells.
Nucleic Acid Structure
Levels of DNA Structure
Nucleotides: Building blocks of DNA and RNA.
Strand: Linear polymer of nucleotides.
Double helix: Two strands of DNA wound together.
Chromosomes: DNA associated with proteins, forming a complex structure.
Genome: Complete set of genetic material in an organism.
DNA and RNA Nucleotide Structure
Both DNA and RNA are polymers of nucleotides, each consisting of a phosphate group, a pentose sugar, and a nitrogenous base. DNA contains deoxyribose and the bases adenine (A), guanine (G), cytosine (C), and thymine (T). RNA contains ribose and uracil (U) instead of thymine.
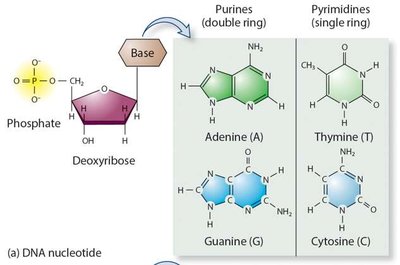
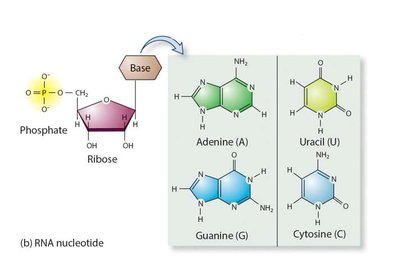
Purines: Adenine (A), Guanine (G) – double ring structure.
Pyrimidines: Cytosine (C), Thymine (T, in DNA), Uracil (U, in RNA) – single ring structure.
Nucleotide Numbering and DNA Strand Structure
The carbons in the sugar are numbered 1' to 5'. The base attaches to the 1' carbon, and the phosphate group attaches to the 5' carbon. DNA strands have directionality, with a 5' phosphate end and a 3' hydroxyl end. Nucleotides are joined by phosphodiester bonds, forming a sugar-phosphate backbone.
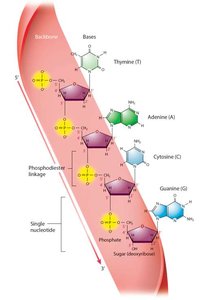
Example: 5’ – TACG – 3’
Solving the Structure of DNA
Watson, Crick, and the Double Helix
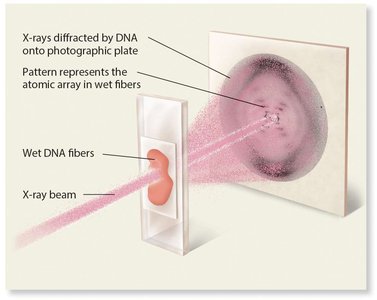
James Watson and Francis Crick, using model-building and X-ray diffraction data from Rosalind Franklin, proposed the double helix structure of DNA in 1953. Their model explained how DNA could store genetic information and replicate accurately.

Chargaff’s Rules and Base Pairing
Erwin Chargaff found that in DNA, the amount of adenine equals thymine, and the amount of guanine equals cytosine. This base pairing (A=T, G≡C) is essential for the double helix structure and for accurate replication.
Organism | Adenine (%) | Thymine (%) | Guanine (%) | Cytosine (%) |
|---|---|---|---|---|
Escherichia coli | 26.0 | 23.9 | 24.9 | 25.2 |
Streptococcus pneumoniae | 29.8 | 31.6 | 20.5 | 18.0 |
Saccharomyces cerevisiae | 31.7 | 32.6 | 18.3 | 17.4 |
Turtle | 28.7 | 27.9 | 22.0 | 21.3 |
Salmon | 29.7 | 29.1 | 20.2 | 21.0 |
Chicken | 28.0 | 28.8 | 21.6 | 21.6 |
Human | 30.3 | 30.3 | 19.5 | 19.9 |

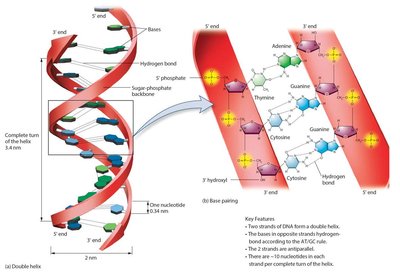
Features of the DNA Double Helix
Double-stranded, antiparallel strands
Right-handed helix
Sugar-phosphate backbone on the outside, bases on the inside
Stabilized by hydrogen bonds between complementary bases
Approximately 10 nucleotides per helical turn
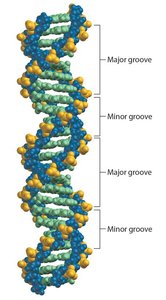
Major and Minor Grooves
The DNA double helix has major and minor grooves, which are important for protein binding and gene regulation. The major groove is wider and more accessible to proteins that regulate gene expression.
DNA Replication
Models of DNA Replication
Three models were proposed for DNA replication: semiconservative, conservative, and dispersive. The semiconservative model, where each daughter DNA has one old and one new strand, was supported by experimental evidence.
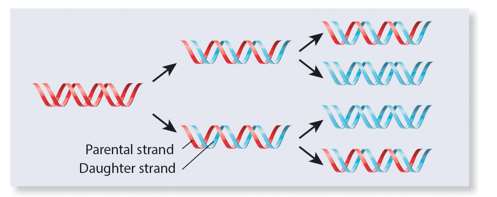
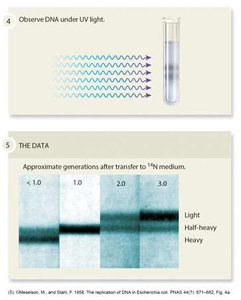
The Meselson–Stahl Experiment
Meselson and Stahl used isotopic labeling and density gradient centrifugation to show that DNA replication is semiconservative. After one generation in light nitrogen, DNA had intermediate density; after two generations, both light and intermediate DNA were present.
Molecular Mechanism of DNA Replication
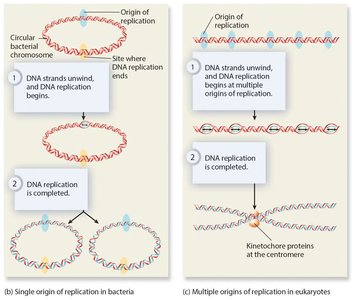
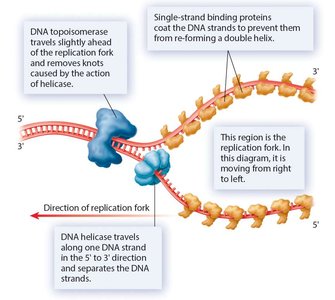
Replication begins at origins of replication, forming replication bubbles and forks. Bacteria have a single origin; eukaryotes have multiple origins. DNA polymerase synthesizes new DNA strands using parental strands as templates, following the AT/GC base-pairing rule.
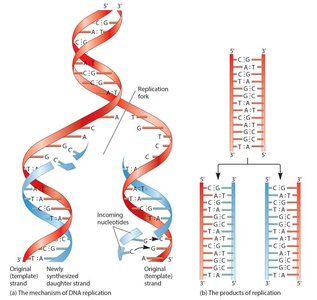
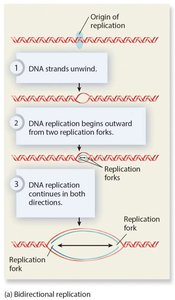
DNA Polymerase and the Role of Primers
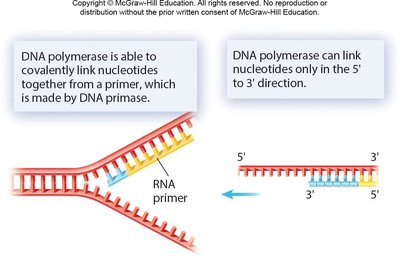
DNA polymerase cannot initiate synthesis on a bare template; it requires a short RNA primer made by primase. DNA polymerase adds nucleotides in the 5’ to 3’ direction, replacing RNA primers with DNA later.
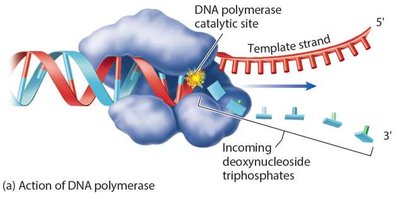
Energy for DNA Synthesis
DNA polymerization is driven by the hydrolysis of high-energy deoxyribonucleoside triphosphates (dNTPs), making the reaction exergonic.
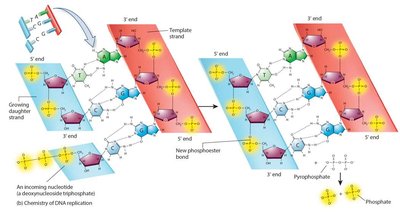
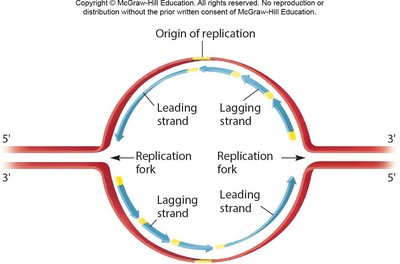
Leading and Lagging Strands; Okazaki Fragments
The leading strand is synthesized continuously, while the lagging strand is synthesized discontinuously as Okazaki fragments. DNA ligase joins these fragments after RNA primers are replaced with DNA.

Accuracy of DNA Replication
Hydrogen bonding between correct base pairs is more stable.
DNA polymerase active site excludes mismatched pairs.
Proofreading activity of DNA polymerase removes errors.
DNA Polymerase Diversity
Multiple DNA polymerases exist in both prokaryotes and eukaryotes, with specialized functions for replication and repair. For example, E. coli has five DNA polymerases, while humans have at least twelve, designated by Greek letters.
Replicating the Ends of Linear Chromosomes: Telomeres and Telomerase
The End Replication Problem
On the lagging strand, the inability to replace the final RNA primer leaves a single-stranded overhang, leading to progressive chromosome shortening. Telomeres, composed of repetitive sequences, protect genes from loss.
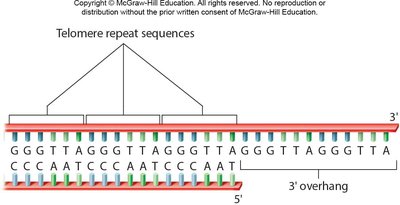
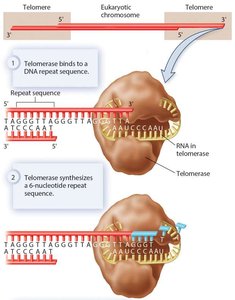
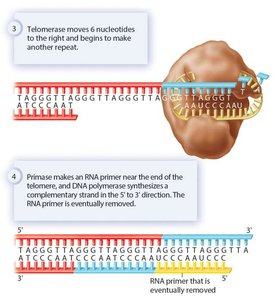
Role of Telomerase
Telomerase extends the 3' end of the lagging strand using an RNA template, allowing complete replication of chromosome ends. This enzyme is active in germ cells and some stem cells, but not in most somatic cells.
Molecular Structure of Eukaryotic Chromosomes
Chromatin Organization and DNA Compaction
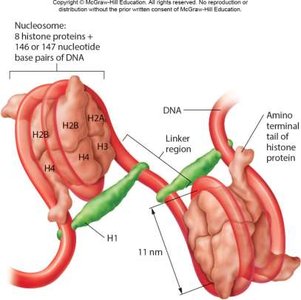
Eukaryotic chromosomes are highly compacted to fit within the nucleus. DNA wraps around histone proteins to form nucleosomes, which further fold into 30-nm fibers and higher-order structures such as radial loop domains.
Nucleosome: DNA wrapped around histone octamer.
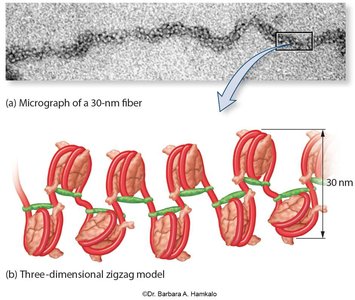
30-nm fiber: Zigzag folding of nucleosomes.
Radial loop domains: Interaction with nuclear matrix, organizing chromosomes into discrete territories.
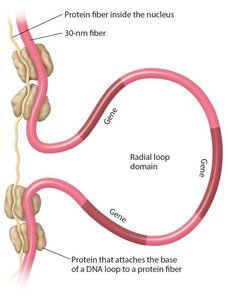
During cell division, chromosomes become even more compacted, with heterochromatin being highly condensed and euchromatin less so.
