 Back
BackPhylogeny and the Tree of Life: Study Notes
Study Guide - Smart Notes

Phylogeny and the Tree of Life
Introduction to Phylogeny and Systematics
Phylogeny is the evolutionary history of a species or group of related species. Systematics is the scientific discipline focused on classifying organisms and determining their evolutionary relationships. Understanding phylogeny helps biologists reconstruct the evolutionary pathways that have led to the diversity of life on Earth.
Phylogeny: The evolutionary history and relationships among species or groups.
Systematics: The study of biological diversity in an evolutionary context, encompassing taxonomy and phylogenetics.
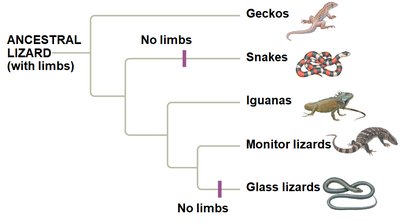
Example: Phylogenetic analysis reveals that legless lizards and snakes evolved from different lineages of lizards with limbs.

Binomial Nomenclature and Hierarchical Classification
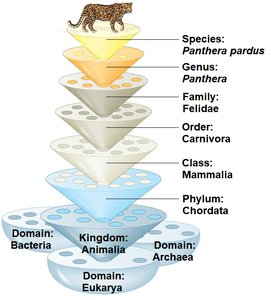
Modern taxonomy uses a two-part naming system (binomial nomenclature) and organizes species into a hierarchy of increasingly broad categories. This system, developed by Carolus Linnaeus, remains the foundation of biological classification.
Binomial Nomenclature: Each species is assigned a two-part Latin name: the genus (capitalized) and the specific epithet (lowercase), both italicized (e.g., Homo sapiens).
Hierarchical Classification: Species are grouped into a hierarchy: Domain, Kingdom, Phylum, Class, Order, Family, Genus, Species.
Linking Classification and Phylogeny
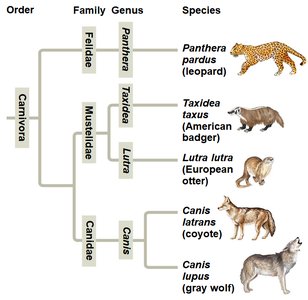
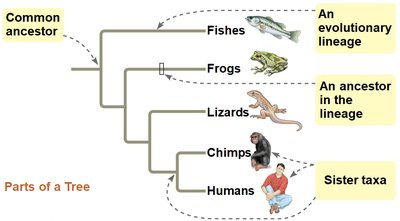
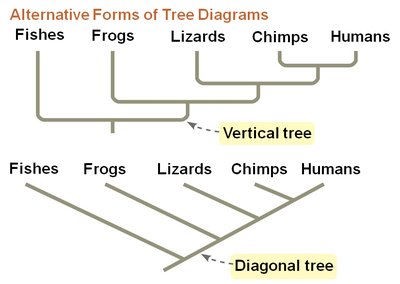
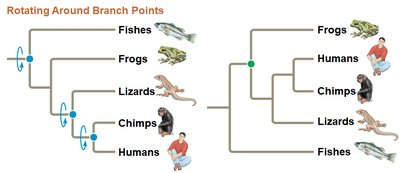
Phylogenetic trees visually represent hypotheses about evolutionary relationships. The tips of the branches represent taxa, and branch points (nodes) indicate common ancestors. Trees can be drawn in various orientations without changing the relationships they depict.
Branch Point (Node): Represents the divergence of two evolutionary lineages from a common ancestor.
Sister Taxa: Groups that share an immediate common ancestor not shared by any other group.
Tree Orientation: Trees can be drawn vertically, horizontally, or diagonally; the relationships remain unchanged.
Inferring Phylogenies: Morphological and Molecular Data
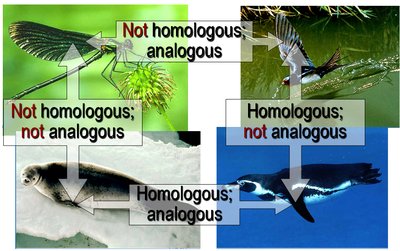
Systematists use morphological, genetic, and biochemical data to infer phylogenies. Similarities due to shared ancestry are called homologies, while similarities due to convergent evolution are analogies.
Homology: Similarity resulting from common ancestry (e.g., vertebrate forelimbs).
Analogy: Similarity due to convergent evolution, not shared ancestry (e.g., wings of birds and insects).

Cladistics and Phyletic Groupings
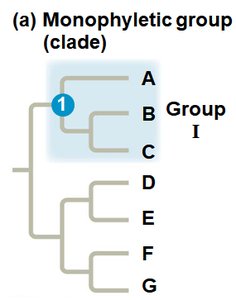
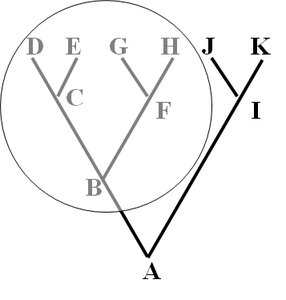
Cladistics is a method of systematics that groups organisms by common ancestry. A clade is a group consisting of an ancestor and all its descendants. Clades can be monophyletic, paraphyletic, or polyphyletic.
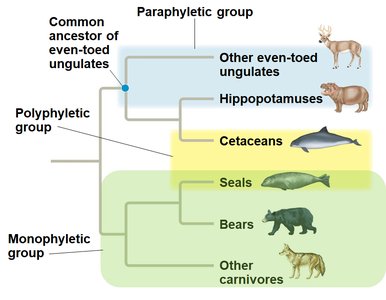
Monophyletic Group (Clade): Includes an ancestor and all its descendants.
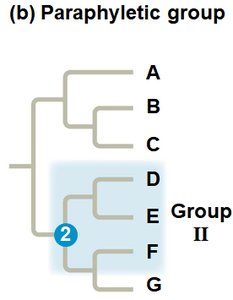
Paraphyletic Group: Includes an ancestor and some, but not all, descendants.
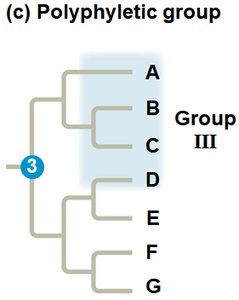
Polyphyletic Group: Includes distantly related species but not their most recent common ancestor.
Shared Ancestral and Shared Derived Characters
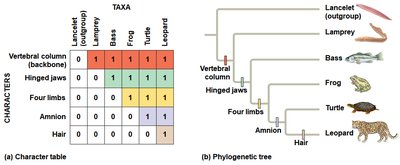
Characters used in cladistics can be ancestral (originated in an ancestor) or derived (unique to a clade). Outgroups help distinguish between shared ancestral and shared derived characters.
Shared Ancestral Character: A trait present in the ancestor of a group.
Shared Derived Character: An evolutionary novelty unique to a particular clade.
Outgroup: A species or group closely related to the ingroup but diverged before the lineage of the ingroup.
Phylogenetic Trees and Branch Lengths
Branch lengths in phylogenetic trees can represent the number of genetic changes or the passage of time, depending on the data and method used.
Genetic Changes: Longer branches indicate more genetic changes.
Time: Branch lengths can be proportional to time, calibrated with fossil records.
Tree-Building Methods: Parsimony and Likelihood
Systematists use principles such as maximum parsimony (fewest evolutionary events) and maximum likelihood (most probable sequence of events) to select the best phylogenetic tree.
Maximum Parsimony: The simplest explanation (fewest changes) is preferred.
Maximum Likelihood: Considers probability of DNA changes and evolutionary events.
Phylogenetic Trees as Hypotheses
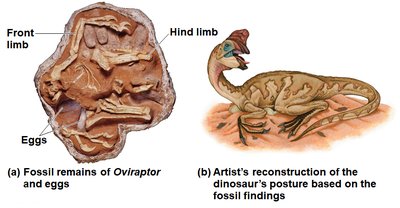
Phylogenetic trees are hypotheses that can be tested with additional data. Phylogenetic bracketing allows scientists to infer characteristics of extinct ancestors based on shared traits in living descendants.
Genomes and Evolutionary History
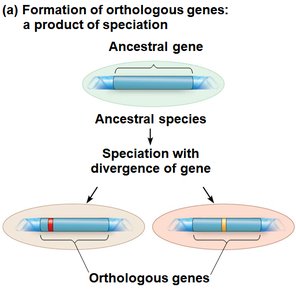
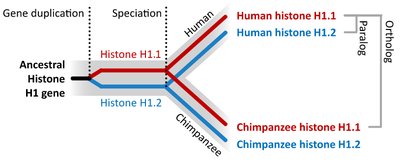
Comparing DNA and other molecules helps trace evolutionary history. Genes can be orthologous (between species) or paralogous (within a species due to duplication).
Orthologous Genes: Homologous genes found in different species, diverged after a speciation event.
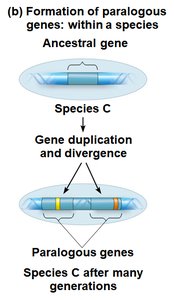
Paralogous Genes: Homologous genes within a species, diverged after gene duplication.
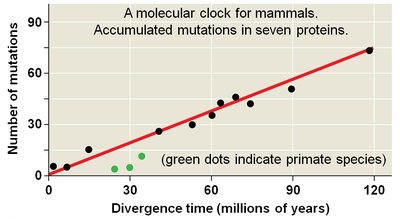
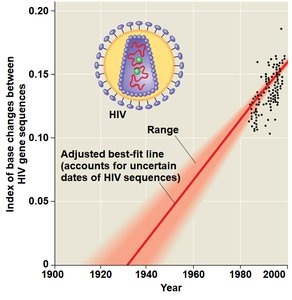
Molecular Clocks
Molecular clocks use the constant rate of genetic change in some genes to estimate the timing of evolutionary events. They are calibrated with fossil data and can be used to date divergences beyond the fossil record.
Orthologous Genes: Substitutions proportional to time since last common ancestor.
Paralogous Genes: Substitutions proportional to time since gene duplication.
Limitations: Not all genes evolve at the same rate; natural selection and other factors can affect clock accuracy.
The Tree of Life and Horizontal Gene Transfer
Modern systematics recognizes three domains of life: Bacteria, Archaea, and Eukarya. Horizontal gene transfer (HGT) complicates the reconstruction of evolutionary relationships, as genes can move between distantly related lineages.
Three-Domain System: Bacteria, Archaea, and Eukarya, supported by molecular data.
Horizontal Gene Transfer: The movement of genes between genomes, common in prokaryotes and some eukaryotes.
Impact: HGT can blur the lines of descent, making the tree of life more like a network.
