 Back
BackCH. 19 p. 2- Post-Transcriptional Control and Comparison of Gene Expression in Bacteria and Eukaryotes
Study Guide - Smart Notes

Post-Transcriptional Control in Eukaryotes
Alternative Splicing – Control During RNA Processing
Alternative splicing is a major mechanism of gene regulation in eukaryotes, allowing a single gene to produce multiple protein variants. This process occurs during RNA processing, where the primary RNA transcript (pre-mRNA) is spliced in different ways to generate multiple mature mRNAs, each potentially encoding a protein with a distinct amino acid sequence.
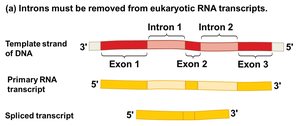
RNA Splicing: Introns (non-coding regions) are removed from the primary transcript, and exons (coding regions) are joined together to form mature mRNA.
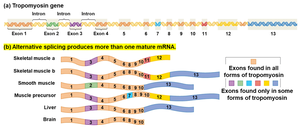
Alternative Splicing: The same pre-mRNA can be spliced in various ways, resulting in different combinations of exons in the final mRNA.
Biological Significance: In humans, over 90% of genes undergo alternative splicing, greatly expanding the diversity of proteins.
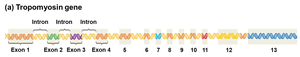
Example: The tropomyosin gene is alternatively spliced to produce different proteins in skeletal muscle, smooth muscle, and other tissues, each with unique but related functions.
Mechanism of Alternative Splicing
Alternative splicing is regulated by proteins that bind to the pre-mRNA and influence the splicing machinery. These regulatory proteins are expressed in a cell-type-specific manner, allowing different cell types to produce distinct protein isoforms from the same gene.
Splicing Regulatory Proteins: These proteins determine where small nuclear ribonucleoproteins (snRNPs) bind, influencing which exons are included or excluded.
Cellular Signals: Cells receive signals from their environment that affect the expression of splicing regulators, leading to cell-type-specific mRNA and protein production.
Example: In fruit flies, a single gene can produce tens of thousands of different mRNAs through alternative splicing.
Translational Control and RNA Interference
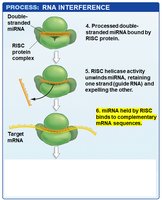
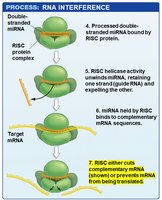
Translational control regulates gene expression by influencing the translation of mRNA into protein. One major mechanism is RNA interference (RNAi), which involves small RNAs such as microRNAs (miRNAs) that bind to target mRNAs and inhibit their translation or promote their degradation.
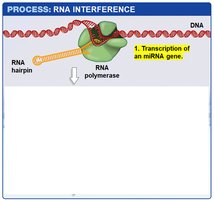
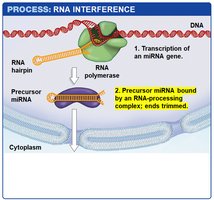
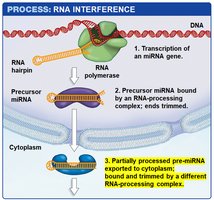
miRNA Biogenesis: miRNAs are transcribed as hairpin precursors, processed in the nucleus, exported to the cytoplasm, and further processed to form mature miRNAs.
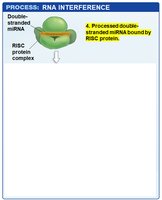
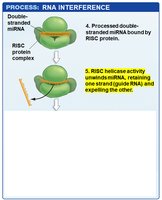
RISC Complex: The mature miRNA is loaded into the RNA-induced silencing complex (RISC), which uses the miRNA as a guide to bind complementary sequences on target mRNAs.
Outcomes: If the miRNA and mRNA are perfectly complementary, RISC cleaves the mRNA. If not, translation is inhibited without mRNA degradation.
Post-Translational Control
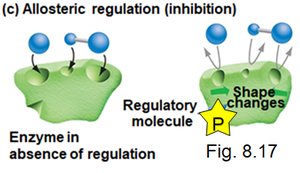
Post-translational control involves modifications to proteins after they are synthesized, affecting their activity, localization, or stability.
Activation/Deactivation: Enzymes such as kinases add phosphate groups (phosphorylation), while phosphatases remove them (dephosphorylation), altering protein function.
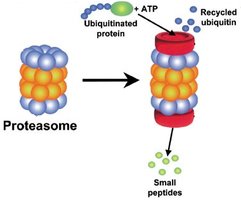
Targeted Destruction: Proteins can be tagged with ubiquitin molecules, marking them for degradation by the proteasome, a large protein complex that digests proteins into peptides.
Comparison of Gene Expression in Bacteria and Eukaryotes
Overview
Gene expression is regulated at multiple levels in both bacteria and eukaryotes, but the mechanisms and complexity differ significantly between these domains of life.
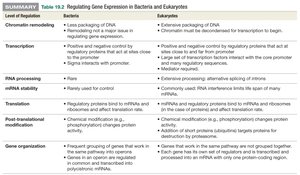
Level of Regulation | Bacteria | Eukaryotes |
|---|---|---|
Chromatin Remodeling | Less packaging of DNA; not a major issue in transcription regulation | Extensive packaging of DNA; chromatin must be decondensed for transcription to begin |
Transcription | Positive and negative control by regulatory proteins near the promoter; sigma protein required for RNA polymerase binding | Positive and negative control by regulatory proteins near and far from the promoter; many transcription factors and mediator proteins required |
RNA Processing | Rare | Extensive processing, including alternative splicing of introns |
mRNA Stability | Rarely used for regulation | Commonly used; RNA interference limits activity and lifespan of many mRNAs |
Translation | Regulatory proteins bind to mRNAs and ribosomes to affect translation rate | Small RNAs (e.g., miRNA) and regulatory proteins bind to mRNAs and ribosomes to affect translation rate |
Post-Translational Modification | Chemical modifications (e.g., phosphorylation) change protein activity | Chemical modifications (e.g., phosphorylation) and addition of ubiquitin for targeted protein destruction |
Gene Organization | Genes in the same pathway often grouped in operons and transcribed as a single mRNA | Genes in the same pathway are not grouped together; each gene has its own regulators and is transcribed separately |

Key Differences
Chromatin Structure: Eukaryotic DNA is packaged into chromatin, requiring remodeling for gene expression, unlike bacterial DNA.
RNA Processing: Eukaryotes extensively process RNA (capping, polyadenylation, splicing), while bacteria do not.
Regulatory Complexity: Eukaryotic gene expression involves more regulatory proteins and steps, allowing for greater control and diversity.
Gene Organization: Operons are common in bacteria but absent in eukaryotes, where genes are regulated individually.
