 Back
BackProtein Structure and Function: Study Notes
Study Guide - Smart Notes

Protein Structure and Function
3.1 Amino Acids and Their Polymerization
Amino acids are the building blocks of proteins, each sharing a common core structure but differing in their side chains (R-groups). Their chemical properties and interactions are fundamental to protein structure and function.
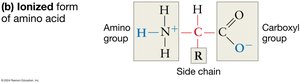
Structure of Amino Acids: Each amino acid consists of a central (alpha) carbon atom bonded to a hydrogen atom, an amino group (–NH2), a carboxyl group (–COOH), and a variable R-group (side chain).
Ionization in Water: In aqueous solutions, the amino group tends to gain a proton (becoming –NH3+), and the carboxyl group loses a proton (becoming –COO–), which helps amino acids stay in solution and affects their reactivity.
R-Groups (Side Chains): The R-group is the variable part of an amino acid, making each of the 20 standard amino acids unique. R-groups can contain functional groups that participate in chemical reactions or may be composed solely of carbon and hydrogen.
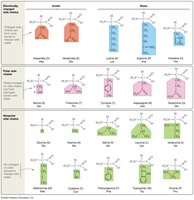
Classification by Polarity and Charge: Amino acids are classified based on the properties of their R-groups:
Charged (acidic or basic): Side chains with a negative charge are acidic (lost a proton), while those with a positive charge are basic (gained a proton).
Uncharged Polar: Side chains with an oxygen atom (highly electronegative) form polar covalent bonds.
Nonpolar: Side chains lacking charge and oxygen are nonpolar (e.g., methionine).
Polymerization of Amino Acids: Amino acids link via peptide bonds, which form between the carboxyl group of one amino acid and the amino group of another through a condensation reaction (releasing water).
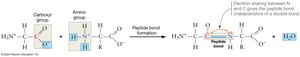
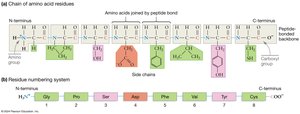
Peptide-Bonded Backbone:
R-group orientation: Side chains extend outward, allowing interactions.
Directionality: The chain has an N-terminus (free amino group) and a C-terminus (free carboxyl group); sequences are written from N- to C-terminus.
Flexibility: The peptide bond itself is rigid, but single bonds adjacent to it can rotate, allowing flexibility in the chain.
Oligopeptides and Polypeptides: Chains with fewer than 50 amino acids are called oligopeptides or peptides; longer chains are polypeptides. A protein is a complete, functional polypeptide.
3.2 What Do Proteins Look Like?
Protein structure is hierarchical and determines function. Proteins are the most diverse class of biological molecules in terms of size, shape, and chemical properties.
Primary Structure: The unique sequence of amino acids in a protein. Even a single amino acid change can drastically alter protein function (e.g., sickle-cell hemoglobin).
Secondary Structure: Local folding patterns stabilized by hydrogen bonds between backbone atoms. The main types are:
Alpha-helix (α-helix)
Beta-pleated sheet (β-sheet)
Tertiary Structure: The overall 3D shape of a polypeptide, resulting from interactions among R-groups:
Hydrogen bonding (between polar side chains)
Hydrophobic interactions (nonpolar side chains cluster away from water)
van der Waals interactions (weak attractions between hydrophobic side chains)
Covalent bonding (e.g., disulfide bonds between cysteine residues)
Ionic bonding (between oppositely charged side chains)
Quaternary Structure: The arrangement of multiple polypeptide subunits in a protein. Subunits may be identical (homodimers) or different (heterodimers). Some proteins form large complexes called macromolecular machines (e.g., ribosomes).
Level of Structure | Stabilizing Bonds/Interactions | Description |
|---|---|---|
Primary | Peptide bonds | Linear sequence of amino acids |
Secondary | Hydrogen bonds | Alpha-helices and beta-sheets |
Tertiary | Hydrogen, hydrophobic, van der Waals, covalent, ionic | 3D folding of a single polypeptide |
Quaternary | Same as tertiary (between subunits) | Multiple polypeptides forming a functional protein |
3.3 Folding and Function
Proper protein folding is essential for function. Folding is driven by chemical interactions and is often spontaneous, resulting in a stable, low-energy structure. Misfolded proteins can lose function or become harmful.
Molecular Chaperones: Specialized proteins that assist in the proper folding of other proteins and prevent inappropriate interactions (e.g., heat shock proteins like Hsp90).
Protein Flexibility: Many proteins are dynamic and can adopt multiple conformations until they bind specific molecules or are activated.
Prions: Infectious, misfolded proteins that can induce normal proteins to adopt the abnormal conformation (e.g., PrP in mad cow disease).
3.4 Protein Functions Are as Diverse as Protein Structures
Proteins perform a vast array of functions in cells, reflecting their structural diversity.
Catalysis: Enzymes accelerate chemical reactions by binding substrates at their active sites and orienting them for reaction.
Structure: Proteins provide support and shape to cells and tissues.
Movement: Motor proteins move cells or molecules within cells.
Signaling: Proteins transmit signals between cells.
Transport: Proteins facilitate the movement of substances across membranes or throughout the body.
Defense: Antibodies and other proteins protect against pathogens.
Why Are Enzymes Good Catalysts? Enzymes are highly specific, binding substrates at their active sites and lowering the activation energy required for reactions.
