 Back
BackRegulation of Gene Expression in Eukaryotes
Study Guide - Smart Notes

Regulation of Gene Expression in Eukaryotes
Overview of Eukaryotic Gene Expression
Gene expression in eukaryotic cells is a highly regulated process that determines when, where, and how much of each protein is produced. Regulation occurs at multiple stages, from chromatin structure to post-translational modifications, allowing cells to respond to internal and external signals and to specialize for distinct functions.
Differential gene expression enables cell specialization in multicellular organisms.
Abnormal gene expression can lead to diseases, including cancer.
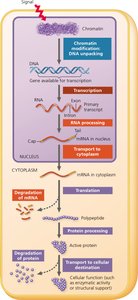
Stages of Gene Expression Regulation
Gene expression can be regulated at several key stages, each providing opportunities for control:
Chromatin modification: DNA packaging affects accessibility for transcription.
Transcription: Initiation and rate of mRNA synthesis are tightly controlled.
RNA processing: Includes capping, polyadenylation, and splicing of pre-mRNA.
mRNA transport and degradation: Stability and localization of mRNA affect protein synthesis.
Translation: Initiation and efficiency of protein synthesis from mRNA.
Protein processing and degradation: Modifications and turnover of proteins regulate their activity and lifespan.
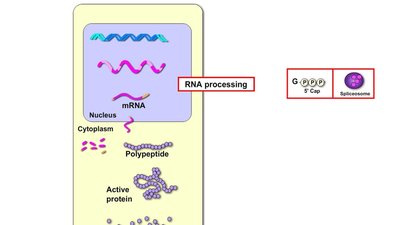
Regulation of Chromatin Structure
Chromatin Organization and Gene Accessibility
DNA in eukaryotic cells is packaged with histone proteins into chromatin. The basic unit, the nucleosome, consists of DNA wrapped around histone octamers. Chromatin structure influences gene expression by controlling the accessibility of DNA to transcription machinery.
Heterochromatin: Highly condensed, generally transcriptionally inactive.
Euchromatin: Less condensed, accessible for transcription.
Histone Modifications and DNA Methylation
Chemical modifications to histones and DNA play a direct role in regulating gene expression:
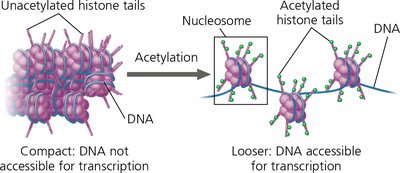
Histone acetylation (addition of acetyl groups) loosens chromatin structure, making DNA more accessible for transcription.
Histone deacetylation leads to tighter packing and reduced transcription.
DNA methylation (addition of methyl groups to cytosine bases) is associated with gene silencing and can be inherited through cell divisions.
Epigenetic Inheritance
Epigenetic inheritance refers to the transmission of traits by mechanisms not involving changes in the DNA sequence, such as chromatin modifications. These changes can be reversible and responsive to environmental factors.
Example: Maternal diet can affect DNA methylation patterns and gene expression in offspring, as shown in coat color variation in mice.

Regulation of Transcription Initiation
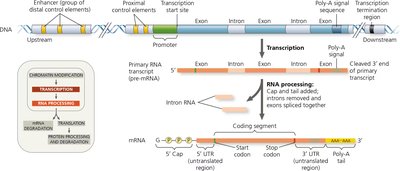
Organization of a Typical Eukaryotic Gene
Eukaryotic genes contain regulatory DNA sequences that control transcription:
Promoter: Site where RNA polymerase binds to initiate transcription.
Control elements: Short DNA sequences that serve as binding sites for transcription factors; can be proximal (near promoter) or distal (enhancers, far from promoter).
Polyadenylation signal: Marks the end of transcription and site for poly-A tail addition.
Transcription Factors
General transcription factors: Required for transcription of all protein-coding genes; assemble at the promoter.
Specific transcription factors: Bind to control elements (enhancers or silencers) to increase (activators) or decrease (repressors) transcription of particular genes.
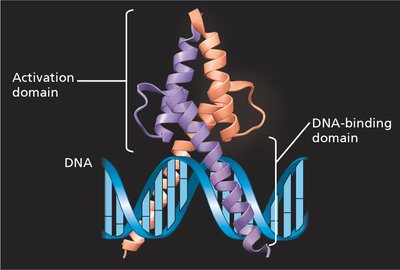
Transcription factors often have distinct DNA-binding domains and activation domains.
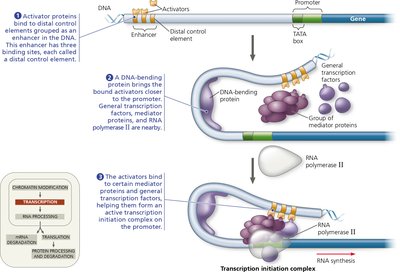
Enhancers and Mediator Proteins
Enhancers are distal control elements that can greatly increase transcription rates. Activators bound to enhancers interact with mediator proteins and general transcription factors at the promoter, facilitating assembly of the transcription initiation complex.
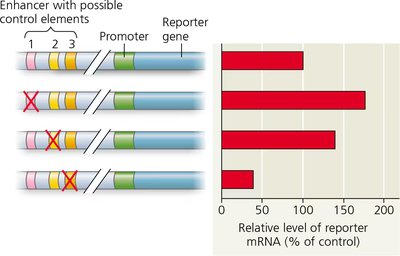
Experimental Analysis of Control Elements
Deletion experiments help identify functional control elements within enhancers by measuring changes in reporter gene expression.
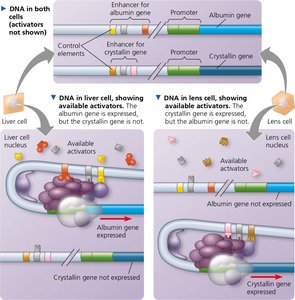
Combinatorial Control and Cell-Type Specificity
Gene expression in eukaryotes is often regulated by the specific combination of control elements present in each gene's enhancer. Different cell types express different sets of transcription factors, allowing for cell-type specific gene expression.
Example: Liver and lens cells express different proteins due to the presence of different activators, even though both cell types contain the same genes.
Coordinate Control of Dispersed Genes
Unlike prokaryotic operons, eukaryotic genes with related functions are often scattered across the genome. Coordinate expression is achieved by shared control elements recognized by the same transcription factors, often in response to external signals such as hormones.
Post-Transcriptional Regulation
RNA Processing and Alternative Splicing
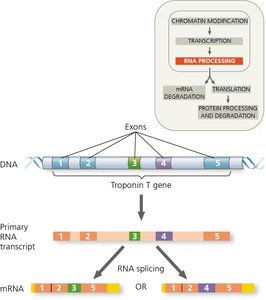
Alternative RNA splicing allows a single gene to produce multiple mRNA variants, increasing protein diversity. Regulatory proteins determine which exons are included in the final mRNA.
Example: The troponin T gene can be spliced in different ways to produce proteins with distinct functions in muscle contraction.
Regulation of mRNA Translation and Degradation
Translation initiation can be blocked by proteins binding to mRNA, or globally regulated by modifying translation factors.
The stability (life span) of mRNA affects how much protein is produced; mRNA degradation is a key regulatory step.
Protein Processing and Degradation
Proteins often require chemical modifications (e.g., phosphorylation, glycosylation) or cleavage to become active.
Selective degradation of proteins, often marked by ubiquitin, controls protein levels and activity in the cell.
Summary Table: Key Regulatory Steps in Eukaryotic Gene Expression
Stage | Regulatory Mechanism | Effect |
|---|---|---|
Chromatin modification | Histone acetylation, DNA methylation | Alters DNA accessibility for transcription |
Transcription initiation | Transcription factors, enhancers, repressors | Controls rate and timing of mRNA synthesis |
RNA processing | Alternative splicing, capping, polyadenylation | Generates mRNA diversity, stability |
mRNA degradation | Regulatory RNAs, mRNA stability factors | Determines mRNA life span and protein output |
Translation | Initiation factors, regulatory proteins | Controls protein synthesis from mRNA |
Protein processing/degradation | Cleavage, chemical modification, ubiquitination | Activates/inactivates proteins, regulates protein levels |
