Back
BackCh 18- Regulation of Gene Expression in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Regulation of Gene Expression
Introduction
Gene expression is the process by which information from a gene is used to synthesize functional gene products, such as proteins. Regulation of gene expression allows cells to respond to environmental changes and maintain homeostasis. In both prokaryotes and eukaryotes, gene expression is tightly controlled at multiple levels, including transcription, RNA processing, translation, and protein modification or degradation.
Bacterial Gene Regulation: The Operon Model
Feedback Inhibition and Gene Regulation
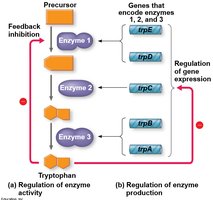
Bacteria can regulate enzyme production through feedback inhibition or by controlling gene expression. Feedback inhibition involves the end product of a metabolic pathway inhibiting an enzyme involved earlier in the pathway, while gene regulation involves controlling the transcription of genes encoding enzymes.
The Operon Concept
An operon is a cluster of functionally related genes regulated together and transcribed as a single mRNA. It consists of:
Promoter: Site where RNA polymerase binds to initiate transcription.
Operator: A DNA segment acting as an on/off switch, usually located within or near the promoter.
Structural genes: Genes encoding proteins with related functions.
The operon can be switched off by a repressor protein, which binds to the operator and blocks RNA polymerase. A corepressor is a small molecule that cooperates with the repressor to turn the operon off.
Repressible and Inducible Operons
There are two main types of negative gene regulation in bacteria:
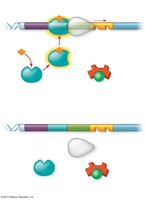
Repressible operons (e.g., trp operon): Usually on; can be turned off when a repressor binds to the operator in the presence of a corepressor (e.g., tryptophan).
Inducible operons (e.g., lac operon): Usually off; can be turned on when an inducer inactivates the repressor, allowing transcription.
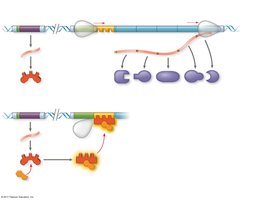
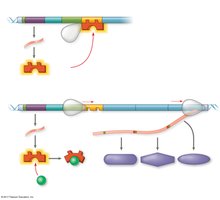
Inducible enzymes typically function in catabolic pathways, while repressible enzymes are involved in anabolic pathways.
Positive Gene Regulation
In addition to negative control, some operons are regulated by activator proteins. For example, the catabolite activator protein (CAP) enhances transcription of the lac operon when glucose is scarce and cAMP levels are high.
Regulation of Gene Expression in Eukaryotes
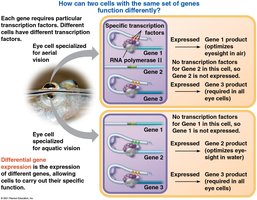
Differential Gene Expression
All cells in a multicellular organism contain the same genome, but different cell types express different sets of genes. This phenomenon is called differential gene expression and is responsible for cell specialization.
Levels of Gene Regulation
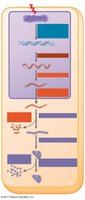
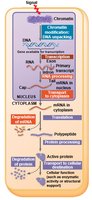
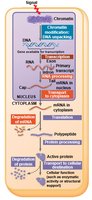
Gene expression in eukaryotes is regulated at multiple stages:
Chromatin modification
Transcription
RNA processing
mRNA transport and degradation
Translation
Protein processing and degradation
Regulation of Chromatin Structure
The structure of chromatin affects gene accessibility. Genes in highly compacted heterochromatin are usually not expressed. Chemical modifications to histones and DNA can influence chromatin structure and gene expression:
Histone acetylation: Addition of acetyl groups to histone tails loosens chromatin, making DNA accessible for transcription.
DNA methylation: Addition of methyl groups to DNA bases (often cytosine) generally reduces transcription.
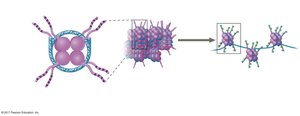
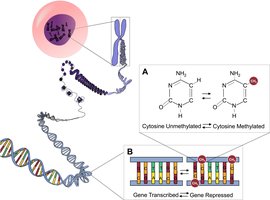
These modifications can be inherited by daughter cells, a phenomenon known as epigenetic inheritance.
Organization of a Typical Eukaryotic Gene
Eukaryotic genes contain multiple regulatory elements:
Promoter: Site for RNA polymerase binding.
Proximal control elements: Located near the promoter.
Distal control elements (enhancers): Can be far from the gene or within introns; bind specific transcription factors.
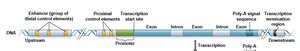
Transcription Initiation and Control Elements
Transcription of protein-coding genes requires general transcription factors and specific transcription factors. The combination of control elements and available activators determines the expression of each gene.
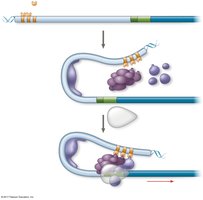
Combinatorial Control of Gene Activation
Gene expression is regulated by the specific combination of control elements and transcription factors present in a cell. This allows for precise and cell-type-specific gene expression.
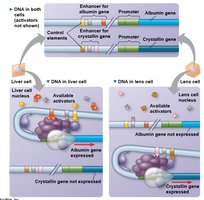
Mechanisms of Post-Transcriptional Regulation
Gene expression can also be regulated after transcription:
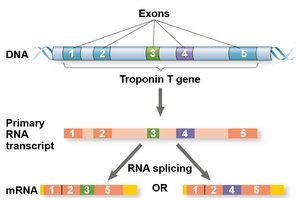
Alternative RNA splicing: Different mRNAs can be produced from the same primary transcript by including or excluding certain exons.
mRNA degradation: The stability and lifespan of mRNA molecules affect protein synthesis.
Translational control: Regulatory proteins can block translation initiation.
Protein processing and degradation: Proteins may be chemically modified, cleaved, or marked for degradation by ubiquitin and destroyed by proteasomes.
Summary Table: Key Mechanisms of Eukaryotic Gene Regulation
Level | Mechanism | Effect |
|---|---|---|
Chromatin modification | Histone acetylation, DNA methylation | Alters DNA accessibility for transcription |
Transcription | Control elements, transcription factors | Initiates or represses gene transcription |
RNA processing | Alternative splicing | Generates different mRNAs from one gene |
mRNA degradation | Poly-A tail length, UTR sequences | Determines mRNA stability and lifespan |
Translation | Regulation of initiation factors | Controls rate of protein synthesis |
Protein processing & degradation | Ubiquitin tagging, proteasome degradation | Regulates protein activity and longevity |
Summary Diagram