 Back
BackThe Molecular Basis for Inheritance: DNA Structure and Replication
Study Guide - Smart Notes

The Molecular Basis for Inheritance
Introduction
The molecular basis for inheritance lies in the structure and function of DNA, which encodes genetic information and ensures its faithful transmission from one generation to the next. This chapter explores the discovery, structure, and replication of DNA, as well as its packaging within chromosomes.
Discovery of Genetic Material
Historical Foundations
Mendel’s Experiments (1865): Demonstrated that traits are inherited in predictable patterns, laying the foundation for genetics.
Rediscovery (~1900): Mendel’s work was recognized as the basis for understanding heredity.
Chromosomal Theory (1910): Thomas Hunt Morgan showed that genes are located on chromosomes, which are composed of proteins and nucleic acids.

Identifying DNA as the Genetic Material
Griffith’s Transformation Experiment (1928): Showed that a substance from dead pathogenic bacteria could transform non-pathogenic bacteria into a pathogenic form.
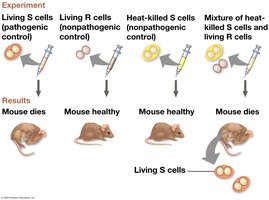
Hershey-Chase Experiment (1952): Used bacteriophages to demonstrate that DNA, not protein, is the genetic material.
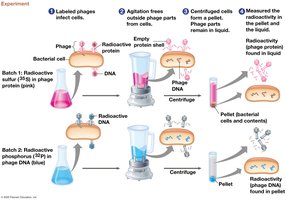
Chargaff’s Rules (1950): Found that DNA composition varies between species, but the amount of adenine (A) equals thymine (T), and guanine (G) equals cytosine (C): ,
Structure of DNA
DNA Nucleotides
DNA is a polymer composed of nucleotides, each consisting of a phosphate group, a deoxyribose sugar, and a nitrogenous base (adenine, thymine, guanine, or cytosine).
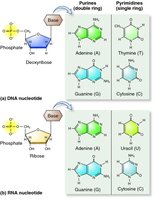
Phosphodiester Bonds: Link nucleotides together, forming the sugar-phosphate backbone.
Nitrogenous Bases: Purines (A, G) and pyrimidines (C, T).
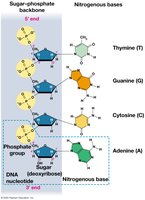
Double Helix Model
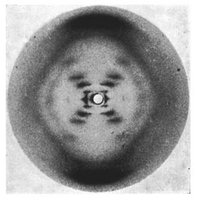
James Watson and Francis Crick (1953), using Chargaff’s rules and Rosalind Franklin’s X-ray images, proposed that DNA is a double-stranded helix with antiparallel strands.
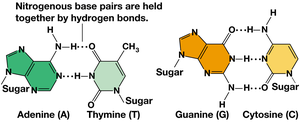
Base Pairing: A pairs with T via two hydrogen bonds; G pairs with C via three hydrogen bonds.
Directionality: DNA strands run in opposite directions (5’ to 3’ and 3’ to 5’).
Chargaff’s Rule and Base Pairing
Chargaff’s Rule: The amount of purines equals the amount of pyrimidines ().
Base Pairing Consistency: Only A-T and G-C pairs are consistent with the structure of DNA.
DNA Replication
Overview of Replication
DNA replication is the process by which a cell copies its DNA before cell division. The double helix unwinds, and each strand serves as a template for a new complementary strand.
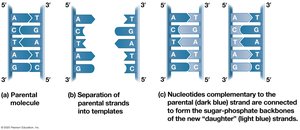
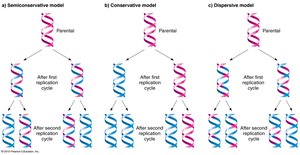
Semiconservative Model: Each new DNA molecule consists of one parental and one new strand.
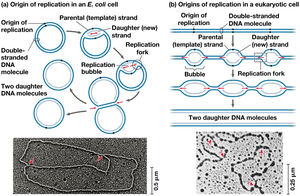
Initiation of Replication
Origins of Replication: Specific sequences where replication begins, forming replication bubbles.
Replication Fork: Y-shaped region where DNA is unwound and new strands are synthesized.
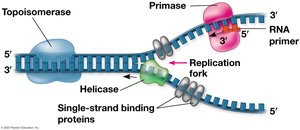
Enzymes Involved in Replication
Helicase: Unwinds the DNA double helix.
Single-Strand Binding Proteins: Stabilize unwound DNA.
Primase: Synthesizes short RNA primers to provide a starting point for DNA synthesis.
DNA Polymerase III: Adds nucleotides to the 3’ end of the primer, synthesizing new DNA in the 5’ to 3’ direction.
DNA Polymerase I: Removes RNA primers and replaces them with DNA.
Ligase: Seals nicks between Okazaki fragments on the lagging strand.
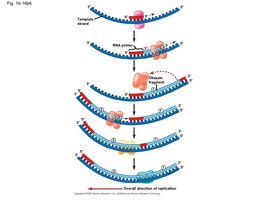
Leading and Lagging Strands
Leading Strand: Synthesized continuously in the direction of the replication fork.
Lagging Strand: Synthesized discontinuously as Okazaki fragments, which are later joined together.
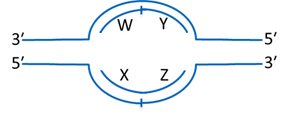
Replication Bubble and Strand Identification
In a replication bubble, the leading and lagging strands can be identified based on the direction of synthesis and the orientation of the template strands.
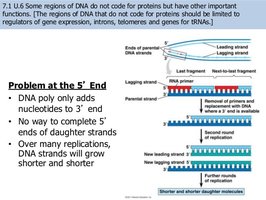
Replication of Linear Chromosomes and Telomeres
Problem at the 5’ End: DNA polymerase cannot complete the 5’ ends of linear chromosomes, leading to progressive shortening with each replication.
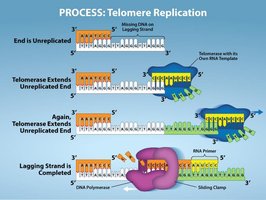
Telomerase: An enzyme that extends telomeres, preventing loss of genetic information in germ cells and some stem cells.
Proofreading and DNA Repair
Proofreading by DNA Polymerase
DNA Polymerase I: Proofreads newly synthesized DNA, removing and replacing incorrect nucleotides.
DNA Repair Mechanisms
Mismatch Repair: Proteins detect and remove incorrectly paired bases after replication.
Nucleotide Excision Repair: Removes damaged DNA segments, such as thymine dimers caused by UV light, and fills the gap with correct nucleotides.
Evolutionary Significance of Mutations
Although DNA repair mechanisms are highly accurate, some errors persist, resulting in mutations. These mutations are the source of genetic variation and drive evolution by natural selection.
Chromosome Structure and DNA Packaging
Bacterial and Eukaryotic Chromosomes
Bacterial Chromosomes: Double-stranded, circular DNA molecules with associated proteins.
Eukaryotic Chromosomes: Linear DNA molecules packaged with proteins into chromatin, found in the nucleus.
Chromatin Organization
Histones: Proteins responsible for the first level of DNA packing. DNA wraps around histone octamers to form nucleosomes.
Nucleosome: The basic unit of chromatin, consisting of DNA wound around histone proteins.
Chromatin States: Euchromatin (less condensed, transcriptionally active) and heterochromatin (highly condensed, transcriptionally inactive).
Higher-Order Chromatin Structure
During interphase, chromatin exists as a 10-nm fiber, which is further compacted into looped domains. Chromosomes are dynamic and can be remodeled for gene expression and other processes.
Histone modifications regulate chromatin structure and gene accessibility.
