Back
BackThe Molecular Basis of Inheritance: DNA Structure, Function, and Replication
Study Guide - Smart Notes

The Molecular Basis of Inheritance
Discovery of DNA as the Genetic Material
Early in the 20th century, the identity of the genetic material was a major question in biology. Initially, proteins were considered more likely candidates than DNA due to their structural complexity and diversity. However, a series of experiments with bacteria and viruses established DNA as the hereditary material.
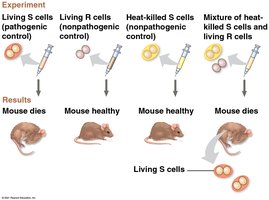
Griffith's Transformation Experiment (1928): Frederick Griffith discovered that non-pathogenic bacteria could be transformed into pathogenic forms by exposure to heat-killed pathogenic bacteria, suggesting the transfer of a 'transforming principle.'
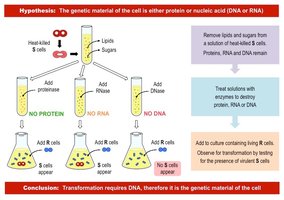
Avery, McCarty, and MacLeod (1944): Identified DNA as the 'transforming principle' by systematically eliminating proteins, RNA, and DNA from bacterial extracts and showing only DNA was required for transformation.
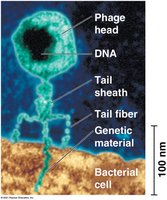
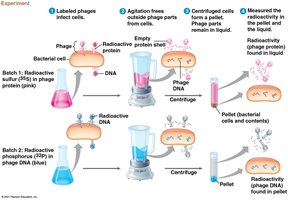
Hershey-Chase Experiment (1952): Used bacteriophages labeled with radioactive isotopes to demonstrate that DNA, not protein, enters bacterial cells and directs viral replication.
Chargaff's Rules and DNA Composition
Erwin Chargaff's experiments revealed two key findings about DNA composition:
The base composition of DNA varies between species.
In any species, the number of adenine (A) bases equals thymine (T), and the number of guanine (G) equals cytosine (C).
These findings provided important clues for the structure of DNA.
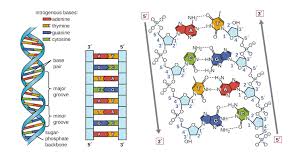
DNA Structure: The Double Helix
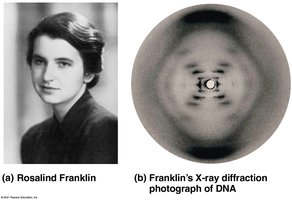
After DNA was established as the genetic material, the next challenge was to determine its structure. Rosalind Franklin and Maurice Wilkins used X-ray crystallography to reveal the helical structure of DNA. James Watson and Francis Crick built a model of DNA as a double helix with antiparallel sugar-phosphate backbones and specific base pairing.
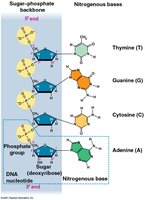
DNA is a polymer of nucleotides, each consisting of:
A nitrogenous base (A, T, G, or C)
A deoxyribose sugar
A phosphate group
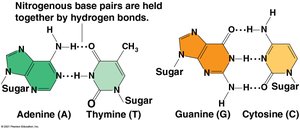
Base pairing: A pairs with T via two hydrogen bonds; G pairs with C via three hydrogen bonds.
Antiparallel strands: The two DNA strands run in opposite directions (5' to 3' and 3' to 5').
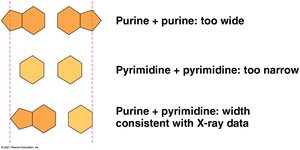
Uniform width: Pairing a purine (A or G) with a pyrimidine (T or C) maintains a constant helix width.

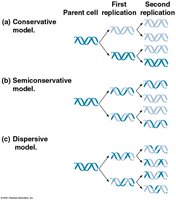
Models of DNA Replication
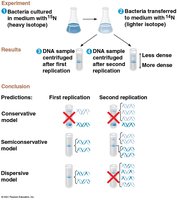
Three models were proposed for DNA replication: conservative, semiconservative, and dispersive. The semiconservative model, supported by Meselson and Stahl's experiments, states that each new DNA molecule consists of one parental and one newly synthesized strand.
Semiconservative replication: Each daughter DNA molecule contains one old strand and one new strand.
Mechanism of DNA Replication
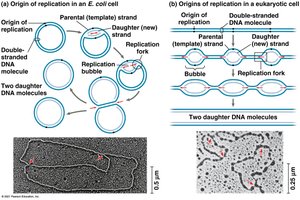
DNA replication begins at origins of replication, where the DNA strands are separated to form replication bubbles. In bacteria, there is a single origin; in eukaryotes, there are multiple origins per chromosome. Replication proceeds bidirectionally until the entire molecule is copied.
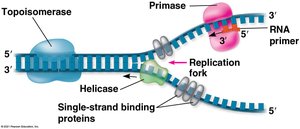
Replication fork: A Y-shaped region where new DNA strands are synthesized.
Key enzymes and proteins:
Helicase: Unwinds the DNA double helix.
Single-strand binding proteins: Stabilize unwound DNA.
Topoisomerase: Relieves strain ahead of the replication fork.
Primase: Synthesizes short RNA primers needed for DNA polymerase to begin synthesis.
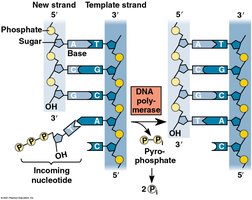
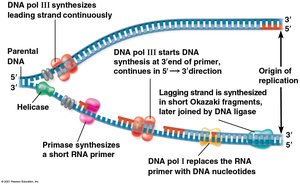
DNA polymerase III: Adds nucleotides to the 3' end of the primer, synthesizing new DNA in the 5' to 3' direction.
Leading strand: Synthesized continuously toward the replication fork.
Lagging strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments, later joined by DNA ligase.
Proofreading and Repair of DNA
DNA polymerases proofread newly synthesized DNA, correcting errors. Additional repair mechanisms, such as mismatch repair and nucleotide excision repair, fix errors and damage caused by environmental factors.
Mismatch repair: Enzymes correct errors missed by DNA polymerase proofreading.
Nucleotide excision repair: Damaged DNA segments are removed and replaced.
Telomeres and Telomerase
Linear eukaryotic chromosomes have repetitive sequences called telomeres at their ends. Telomeres protect genes from erosion during replication but shorten with each cell division. The enzyme telomerase extends telomeres in germ cells, maintaining chromosome integrity across generations.
Telomeres: Repetitive DNA sequences at chromosome ends that protect coding DNA.
Telomerase: Enzyme that extends telomeres in germ cells, preventing loss of genetic information.
Telomere shortening is associated with aging and may limit cell division, acting as a tumor-suppressing mechanism.

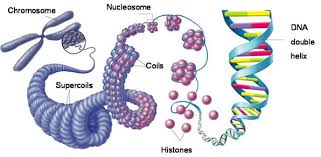
DNA Packing in the Cell
DNA is highly compacted to fit within the cell. In bacteria, DNA is supercoiled and located in the nucleoid. In eukaryotes, DNA is wrapped around histone proteins to form nucleosomes, further coiled and folded into chromatin. Chromatin exists as loosely packed euchromatin or highly condensed heterochromatin.
Histones: Proteins responsible for the first level of DNA packing in eukaryotic chromatin.
Euchromatin: Loosely packed, transcriptionally active chromatin.
Heterochromatin: Densely packed, transcriptionally inactive chromatin (e.g., centromeres, telomeres).