 Back
BackThe Molecular Basis of Inheritance: Structure and Replication of DNA
Study Guide - Smart Notes

Chapter 16: The Molecular Basis of Inheritance
Discovery of the Genetic Material
Early 20th-century biologists sought to identify the molecule responsible for inheritance. Chromosomes were known to carry genetic information, but their composition of both DNA and protein led many to believe proteins were the genetic material due to their complexity.
Griffith's Transformation Experiment: Demonstrated that a 'transforming principle' from dead pathogenic bacteria could make harmless bacteria pathogenic.
Avery, McCarty, and MacLeod: Identified DNA as the transforming agent, but skepticism remained due to limited knowledge of DNA's structure and function.
Bacteriophages and the Hershey-Chase Experiment
Bacteriophages (phages) are viruses that infect bacteria, composed only of DNA and protein. Hershey and Chase used radioactive labeling to show that DNA, not protein, is the genetic material injected into bacteria during infection.
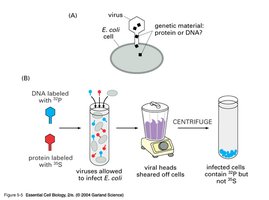
Radioactive sulfur labeled proteins; radioactive phosphorus labeled DNA.
After infection, only DNA entered bacterial cells, confirming DNA as the hereditary material.
Structure of DNA
Nucleotide Structure
DNA is a nucleic acid polymer composed of nucleotides. Each nucleotide consists of a deoxyribose sugar, a phosphate group, and a nitrogenous base (adenine, guanine, cytosine, or thymine).
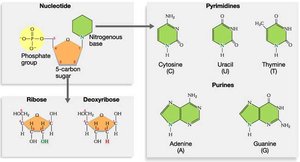
Pyrimidines: Cytosine (C), Thymine (T)
Purines: Adenine (A), Guanine (G)
DNA Polymer Structure
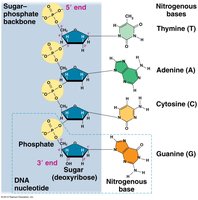
Nucleotides are linked by phosphodiester bonds, forming a sugar-phosphate backbone with nitrogenous bases as side groups. DNA strands have directionality, with a 5' end (phosphate) and a 3' end (hydroxyl).
Chargaff's Rules
Erwin Chargaff discovered that DNA composition varies between species, but within a species, the amount of adenine equals thymine and the amount of guanine equals cytosine. This provided a clue to base pairing in DNA.
Rule 1: Base composition varies between species.
Rule 2: A = T and G = C in any given species.
Discovery of the Double Helix
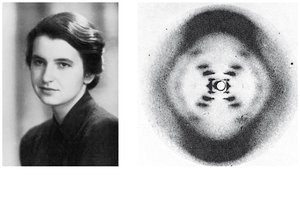
Rosalind Franklin used X-ray crystallography to reveal the helical structure of DNA. Watson and Crick, using Franklin's data, built a model of DNA as a double helix with antiparallel strands and specific base pairing.

Antiparallel strands: The two DNA strands run in opposite directions (5' to 3' and 3' to 5').
Double helix: Two strands coil around each other, stabilized by hydrogen bonds between bases.
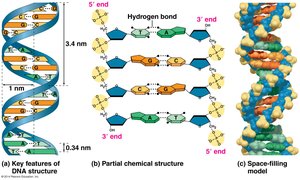
Key Features of DNA Structure
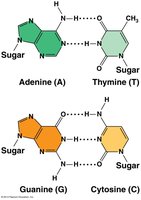
The double helix is stabilized by hydrogen bonds between complementary bases: A pairs with T (2 hydrogen bonds), and G pairs with C (3 hydrogen bonds). The structure explains Chargaff's rules and the mechanism for replication.
DNA Replication
Overview of DNA Replication
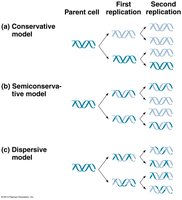
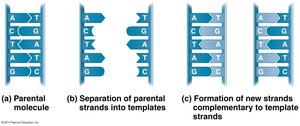
DNA replication is the process by which a cell copies its DNA before cell division. The mechanism is semi-conservative: each daughter DNA molecule consists of one parental and one newly synthesized strand.
Semi-conservative replication: Each new DNA molecule retains one old strand and one new strand.
Initiation of Replication
Replication begins at origins of replication, forming replication bubbles. In eukaryotes, there are multiple origins per chromosome, and replication proceeds bidirectionally.
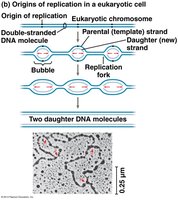
Enzymes and Proteins in DNA Replication
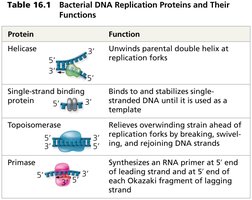
Multiple enzymes and proteins coordinate the replication process:
Protein | Function |
|---|---|
Helicase | Unwinds parental double helix at replication forks |
Single-strand binding protein | Binds and stabilizes single-stranded DNA |
Topoisomerase | Relieves overwinding strain ahead of replication forks |
Primase | Synthesizes RNA primer at 5' end of leading strand and at 5' end of each Okazaki fragment of lagging strand |
DNA polymerase III | Using parental DNA as template, synthesizes new DNA strand by adding nucleotides to an RNA primer or pre-existing DNA strand |
DNA polymerase I | Removes RNA nucleotides of primer from 5' end and replaces them with DNA nucleotides |
DNA ligase | Joins Okazaki fragments of lagging strand; joins 3' end of DNA that replaces primer to rest of leading strand DNA |
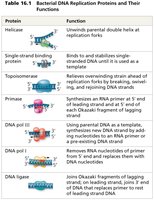

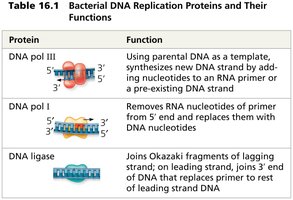
Mechanism of DNA Synthesis
DNA polymerases add nucleotides only to the 3' end of a growing strand, synthesizing DNA in the 5' to 3' direction. The leading strand is synthesized continuously, while the lagging strand is synthesized discontinuously as Okazaki fragments.
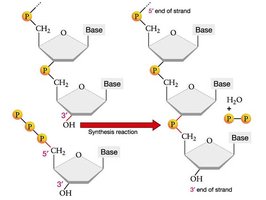
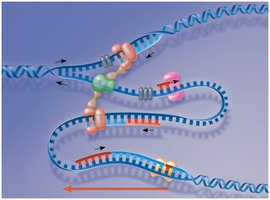
Leading strand: Synthesized continuously toward the replication fork.
Lagging strand: Synthesized in short Okazaki fragments away from the replication fork, later joined by DNA ligase.
Proofreading and Repair of DNA
Proofreading and Mismatch Repair
DNA polymerases proofread each nucleotide as it is added. Mismatch repair enzymes correct errors missed during replication. DNA can also be damaged by environmental factors, requiring repair mechanisms such as nucleotide excision repair.

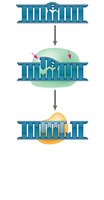
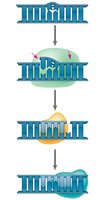
Nuclease: Cuts out damaged DNA segments.
DNA polymerase: Fills in the gap with correct nucleotides.
DNA ligase: Seals the remaining nick in the DNA backbone.
Evolutionary Significance of DNA Repair
Although proofreading and repair mechanisms are highly accurate, some errors (mutations) persist. These mutations are a source of genetic variation, which is essential for evolution by natural selection.
Replication of Chromosome Ends: Telomeres
Telomeres and Replication Limits
Linear DNA molecules in eukaryotic chromosomes have special nucleotide sequences called telomeres at their ends. Due to the limitations of DNA polymerase, the ends of DNA cannot be fully replicated, leading to gradual shortening with each cell division.
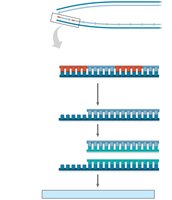
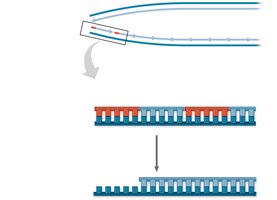
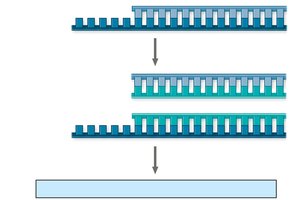
Telomeres: Protect genes near the ends of DNA molecules from erosion.
Telomerase: Enzyme that extends telomeres in germ cells, preventing loss of essential genetic information.
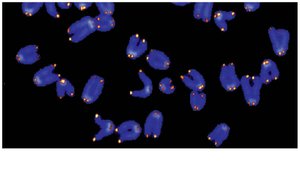
Shortening of telomeres is associated with aging and may protect against cancer by limiting cell divisions. However, cancer cells often reactivate telomerase, allowing unlimited division.
Chromosome Structure and DNA Packing
Organization of Chromosomal DNA
Bacterial chromosomes are circular and supercoiled, while eukaryotic chromosomes are linear and highly organized with proteins into chromatin. Chromatin structure allows efficient packing of DNA within the nucleus.
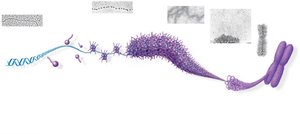
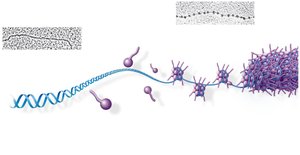

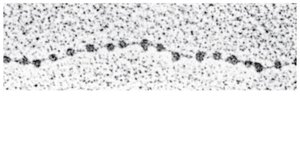
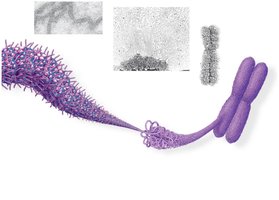


Chromatin Structure and Function
Chromatin can exist in a loosely packed form (euchromatin) or a highly condensed form (heterochromatin). Euchromatin is accessible for gene expression, while heterochromatin is transcriptionally inactive. Chromatin structure changes during the cell cycle and can be regulated by chemical modifications of histones.
Euchromatin: Loosely packed, transcriptionally active.
Heterochromatin: Densely packed, transcriptionally inactive (e.g., centromeres, telomeres).
Additional info: Chemical modifications of histones, such as acetylation and methylation, play a key role in regulating chromatin structure and gene expression.
