 Back
BackThe Molecular Basis of Inheritance: Structure, Function, and Replication of DNA
Study Guide - Smart Notes

The Molecular Basis of Inheritance
Introduction
This chapter explores the experimental evidence and molecular mechanisms underlying the role of DNA as the genetic material, its structure, the process of DNA replication, and the organization of DNA within chromosomes. Understanding these concepts is fundamental to modern genetics and molecular biology.
Evidence for DNA as the Genetic Material
Griffith's Transformation Experiment
Frederick Griffith's 1928 experiment with Streptococcus pneumoniae bacteria provided the first evidence for a "transforming principle" that could transfer genetic information between cells.
S Strain (smooth): Virulent, causes death in mice.
R Strain (rough): Non-virulent, harmless to mice.
Heat-killed S Strain: Harmless alone.
Mix of Heat-killed S Strain and R Strain: Causes death; R strain transformed into virulent form.
Conclusion: A heritable substance from the dead S cells transformed the R cells.
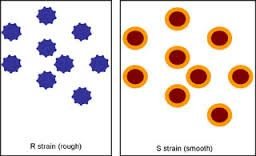
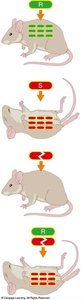
Transformation: The process by which a cell takes up external DNA from its environment.
Hershey-Chase Experiment
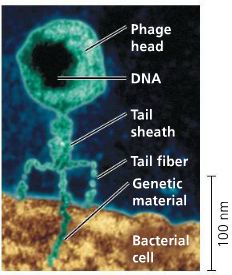
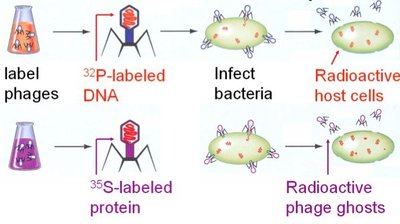
Alfred Hershey and Martha Chase (1952) used bacteriophages (viruses that infect bacteria) to determine whether DNA or protein is the genetic material.
Phages labeled with radioactive sulfur (35S): Labels protein coat.
Phages labeled with radioactive phosphorus (32P): Labels DNA.
Results: Only 32P (DNA) entered bacterial cells and directed viral replication, not 35S (protein).
Conclusion: DNA is the genetic material.
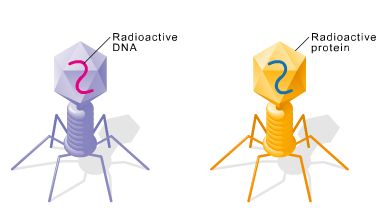
Chargaff's Rules
Erwin Chargaff discovered that the composition of DNA varies between species, but the amount of adenine (A) always equals thymine (T), and the amount of guanine (G) equals cytosine (C).
Chargaff's Rule: %A = %T and %G = %C in DNA.
Implication: Base pairing is specific and fundamental to DNA structure.

Source of DNA | Adenine (%) | Guanine (%) | Cytosine (%) | Thymine (%) |
|---|---|---|---|---|
Sea urchin | 32.8 | 17.7 | 17.7 | 32.1 |
Salmon | 29.7 | 20.8 | 20.4 | 29.1 |
Wheat | 28.1 | 21.8 | 22.7 | 27.4 |
E. coli | 24.7 | 26.0 | 25.7 | 23.6 |
Human | 30.4 | 19.6 | 19.6 | 30.1 |
Ox | 29.0 | 21.2 | 21.2 | 28.6 |
Structure of DNA
Discovery of the Double Helix
Rosalind Franklin used X-ray crystallography to reveal the helical structure of DNA. James Watson and Francis Crick (1953) built the first accurate model of DNA's double helix.
Double Helix: Two antiparallel strands of nucleotides twisted around each other.
Backbone: Sugar-phosphate on the outside; nitrogenous bases on the inside.
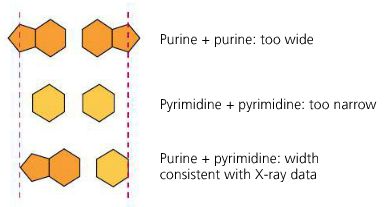
Base Pairing: Purines (A, G) pair with pyrimidines (T, C): A-T and G-C.
Dimensions: 3.4 nm per turn; 0.34 nm per nucleotide.

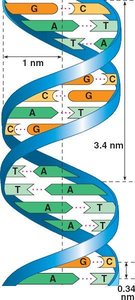
Hydrogen Bonds: Hold the base pairs together (A-T: 2 bonds, G-C: 3 bonds).
DNA Replication
Models of Replication
Three models were proposed for DNA replication: conservative, semiconservative, and dispersive. The semiconservative model was confirmed by the Meselson-Stahl experiment.
Semiconservative: Each daughter DNA has one parental and one new strand.
Conservative: Parental molecule conserved; daughter is entirely new.
Dispersive: Each strand is a mix of old and new DNA.

Meselson-Stahl Experiment
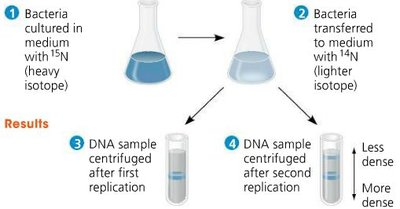
Matthew Meselson and Franklin Stahl (1958) used isotopes of nitrogen (15N and 14N) to demonstrate semiconservative replication.
Bacteria grown in 15N medium, then transferred to 14N medium.
After replication, DNA showed intermediate density, consistent with semiconservative model.

Origins of Replication
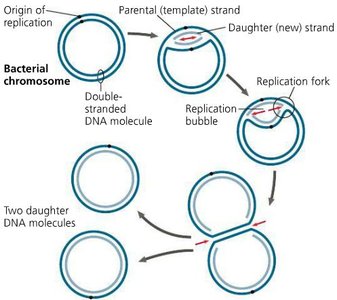
Replication begins at specific sites called origins of replication.
Prokaryotes: Single origin per circular chromosome.
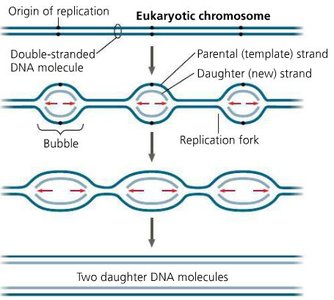
Eukaryotes: Multiple origins per linear chromosome; replication bubbles fuse.
Mechanism of DNA Replication
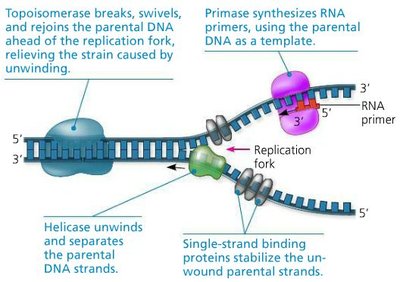
DNA replication is carried out by a complex of enzymes and proteins.
Helicase: Unwinds the DNA double helix.
Single-strand binding proteins: Stabilize unwound DNA.
Primase: Synthesizes RNA primers.
DNA Polymerase III: Adds nucleotides to the 3' end of the primer; synthesizes new DNA in the 5' to 3' direction.
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork in Okazaki fragments.
DNA Polymerase I: Replaces RNA primers with DNA.
DNA Ligase: Joins Okazaki fragments.
Proofreading and DNA Repair
Proofreading by DNA Polymerase
DNA polymerase proofreads each nucleotide as it is added, reducing the error rate to about 1 in 1010 nucleotides.
Mismatch Repair: Other enzymes correct errors missed by DNA polymerase.
Nucleotide Excision Repair: Damaged DNA is cut out by nucleases and replaced by DNA polymerase and ligase.
Mutations: Permanent changes in DNA sequence; can be caused by replication errors or environmental factors (e.g., UV, X-rays).
Somatic mutations: Passed to daughter cells in the body.
Germ-line mutations: Passed to offspring.
Telomeres and Replication of Chromosome Ends
Telomeres
Telomeres are repetitive, non-coding sequences at the ends of eukaryotic chromosomes that protect genes from erosion during replication.
Each round of replication shortens telomeres.
Telomerase is an enzyme that extends telomeres in germ cells, stem cells, and some cancer cells.
Shortened telomeres are associated with aging; high telomerase activity is linked to cancer cell immortality.
Chromatin Structure and Chromosome Packing
Levels of Chromatin Packing
DNA is packaged with proteins to form chromatin, which condenses to form chromosomes during cell division.
Histones: Positively charged proteins around which DNA wraps, forming nucleosomes ("beads on a string").
Nucleosome: Core of 8 histone proteins with DNA wrapped twice around it.
Euchromatin: Less condensed, transcriptionally active.
Heterochromatin: Highly condensed, transcriptionally inactive.
Metaphase Chromosome: Most condensed form, visible during cell division.
Chromatin structure is dynamic and plays a key role in gene regulation and genome stability.
