 Back
BackLEC 18: The Structure and Replication of DNA: Discovery, Evidence, and Models
Study Guide - Smart Notes

DNA Structure and the Double Helix
Discovery and Key Contributors
The discovery of the double-helical structure of DNA was a pivotal moment in molecular biology. This breakthrough was achieved through the combined efforts of several scientists, most notably Rosalind Franklin, James Watson, Francis Crick, and Maurice Wilkins. X-ray diffraction data provided crucial evidence for the helical nature of DNA, while model-building and chemical analysis led to the elucidation of its double-stranded structure.
Rosalind Franklin produced high-quality X-ray diffraction images that revealed the helical structure of DNA.
James Watson and Francis Crick constructed a physical model of DNA that fit all available data, including base pairing and helical geometry.
Maurice Wilkins contributed to the X-ray diffraction studies and shared the Nobel Prize with Watson and Crick in 1962.

X-ray Diffraction Evidence
X-ray diffraction patterns of DNA fibers, such as those produced by Rosalind Franklin, showed an X-shaped pattern characteristic of a helical structure. The data indicated that:
DNA is helical (X-pattern in diffraction image).
It is likely double-stranded.
The sugar-phosphate backbone is on the outside, with bases on the inside.
The distance between the two strands is uniform.

Key Features of DNA Structure
The Double Helix Model
Watson and Crick's model of DNA described it as a right-handed double helix with two antiparallel strands. The sugar-phosphate backbones run in opposite directions (5' to 3' and 3' to 5'), and the nitrogenous bases are paired in the interior of the helix.
Antiparallel orientation: The two DNA strands run in opposite directions.
Right-handed helix: The helix twists in a right-handed direction.
Uniform width: The pairing of bases ensures a consistent diameter of the helix.
Major and minor grooves: The twisting of the helix creates alternating wide (major) and narrow (minor) grooves, important for protein binding.
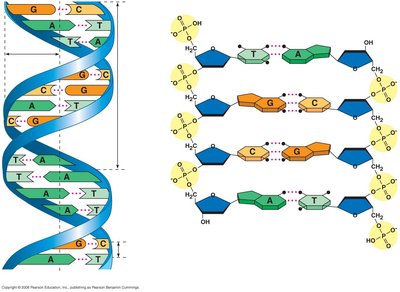
Nucleotide Structure and Base Pairing
Each nucleotide in DNA consists of a deoxyribose sugar, a phosphate group, and a nitrogenous base. The four bases are adenine (A), thymine (T), guanine (G), and cytosine (C). Base pairing is highly specific:
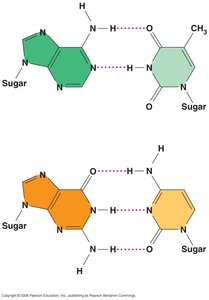
Adenine (A) pairs with Thymine (T) via two hydrogen bonds.
Guanine (G) pairs with Cytosine (C) via three hydrogen bonds.
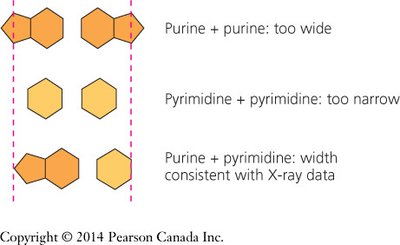
Pairing a purine (A or G) with a pyrimidine (T or C) maintains the uniform width of the helix.
Example: The sequence 5'-ATGC-3' on one strand will pair with 3'-TACG-5' on the complementary strand.
Chargaff's Rules
Erwin Chargaff discovered that in any DNA sample, the amount of adenine equals thymine (A = T), and the amount of guanine equals cytosine (G = C). This provided key evidence for specific base pairing.
Chargaff's rules: A = T and G = C in double-stranded DNA.
This explained the uniform width and the specificity of base pairing in the double helix.
DNA Replication
Models of DNA Replication
Three models were originally proposed for DNA replication:
Conservative model: The parental double helix remains intact, and an entirely new copy is made.
Dispersive model: Each strand of both daughter molecules contains a mixture of old and new DNA.
Semi-conservative model (Watson & Crick model): Each daughter DNA molecule consists of one parental strand and one newly synthesized strand.
Meselson-Stahl Experiment
Matthew Meselson and Franklin Stahl designed an experiment to distinguish between the three models of DNA replication. They used isotopes of nitrogen (15N and 14N) to label DNA and density-gradient ultracentrifugation to separate DNA based on its density.
Method: E. coli were grown in 15N (heavy) medium, then switched to 14N (light) medium. DNA was extracted at various time points and analyzed.
Results: After one generation in 14N, DNA had intermediate density (one band), supporting the semi-conservative model. After two generations, two bands appeared (one light, one intermediate), further confirming the model.
Model | Prediction after 1st Generation | Prediction after 2nd Generation |
|---|---|---|
Conservative | One heavy, one light band | One heavy, one light band |
Dispersive | One intermediate band | One lighter intermediate band |
Semi-conservative | One intermediate band | One light, one intermediate band |
Conclusion: DNA replication is semi-conservative, with each new molecule containing one old and one new strand.
Summary Table: Key Features of DNA Structure
Feature | Description |
|---|---|
Helical Structure | Right-handed double helix |
Strand Orientation | Antiparallel (5' to 3' and 3' to 5') |
Base Pairing | A-T (2 H-bonds), G-C (3 H-bonds) |
Backbone | Sugar-phosphate on outside |
Bases | On inside, paired specifically |
Replication | Semi-conservative |
Additional info: The discovery of the double helix and the mechanism of DNA replication laid the foundation for modern molecular genetics, enabling advances in biotechnology, medicine, and evolutionary biology.
