 Back
BackLEC 21: Transcription and RNA Processing in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcription: Overview and Enzymology
RNA Polymerase and the Transcription Process
Transcription is the process by which RNA is synthesized from a DNA template. The enzyme responsible for this process is RNA polymerase (RNA pol), which catalyzes the formation of phosphodiester bonds between RNA nucleotides, synthesizing RNA in the 5′→3′ direction. RNA polymerase has intrinsic DNA helicase activity, allowing it to unwind the DNA helix during transcription. Unlike DNA polymerase, RNA polymerase does not require a primer to initiate synthesis.
RNA polymerase types in eukaryotes:
RNA pol I: synthesizes rRNA (ribosomal RNA)
RNA pol II: synthesizes mRNA (messenger RNA)
RNA pol III: synthesizes tRNA (transfer RNA)
Bacteria and archaea: possess a single RNA polymerase for all transcription.
Reactants: ATP, GTP, CTP, and UTP (uridine triphosphate).

Promoters and Initiation of Transcription
Promoter Structure and Function
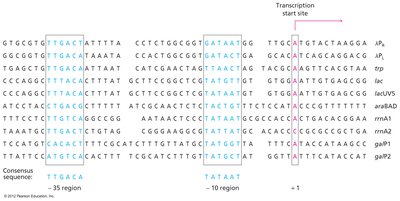
Promoters are specific DNA sequences located upstream of the gene to be transcribed. They serve as binding sites for RNA polymerase and associated proteins, determining the site, timing, and frequency of transcription initiation. In bacteria, promoters contain two consensus sequences: the Pribnow box (−10 element) and the −35 element. In eukaryotes, the TATA box is a common promoter element located 25–35 base pairs upstream of the transcription start site.
Pribnow box (−10 element): 5′-TATAAT-3′
−35 element: 5′-TTGACA-3′
TATA box (eukaryotes): 5′-TATAAA-3′
Initiation is the rate-limiting step in gene expression.
Recruitment of RNA Polymerase
RNA polymerase requires additional proteins to bind promoters:
Bacteria: Sigma (σ) factor binds to the −10 and −35 elements, recruiting RNA polymerase. Sigma dissociates after initiation.
Eukaryotes: General transcription factors (GTFs) bind to promoter regions and recruit RNA polymerase II, forming the transcription initiation complex. Each RNA polymerase type has its own set of GTFs.

Stages of Transcription
1. Initiation
Initiation involves the recruitment of RNA polymerase, unwinding of the DNA helix, and the beginning of RNA synthesis at the +1 site (first base transcribed). The promoter sequence determines the orientation of RNA polymerase binding and the direction of transcription.
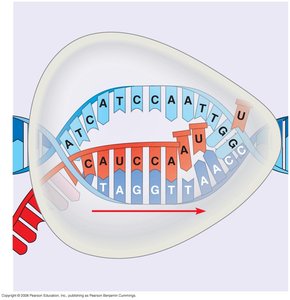
2. Elongation
During elongation, RNA polymerase moves downstream along the DNA template, synthesizing RNA in the 5′→3′ direction. The transcription bubble closes as the enzyme progresses, and the DNA rewinds behind the polymerase.
3. Termination
Termination signals the end of transcription and differs between prokaryotes and eukaryotes:
Bacteria:
Intrinsic termination: A GC-rich region forms a stem-loop structure in the RNA, causing RNA polymerase to dissociate.
Extrinsic (rho-dependent) termination: The rho protein binds to the transcript and unwinds the RNA-DNA duplex, releasing RNA polymerase.
Eukaryotes:
RNA pol II transcribes a polyadenylation signal (AAUAAA), which is recognized by proteins that cleave the transcript and add a poly-A tail.
RNA pol I and III have distinct termination mechanisms involving protein binding or less well-understood processes.

Simultaneous Transcription and Regulation
Multiple RNA Polymerases on a Single Gene
In both prokaryotes and eukaryotes, multiple RNA polymerase molecules can transcribe a single gene simultaneously, allowing rapid production of RNA. The efficiency of promoter recruitment determines the amount of RNA synthesized.
RNA Processing in Eukaryotes
Post-Transcriptional Modifications
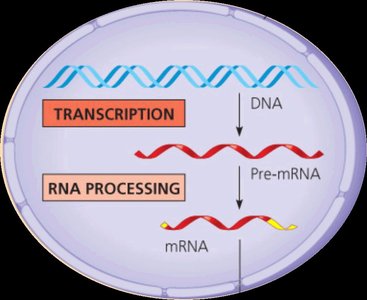
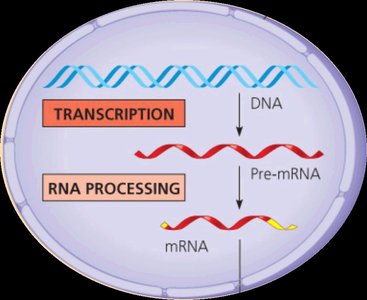
In eukaryotes, the initial RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA. These modifications increase transcript stability and facilitate export and translation.
1. Addition of a 5′ cap: A modified guanosine is added to the 5′ end via a 5′–5′ phosphate linkage.
2. Addition of a poly-A tail: 50–250 adenine nucleotides are added to the 3′ end after cleavage at the polyadenylation signal.
3. RNA splicing: Introns (non-coding regions) are removed, and exons (coding regions) are joined together.
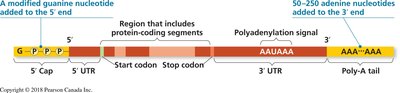
Functions of the 5′ Cap and Poly-A Tail
Facilitate export of mRNA from the nucleus
Protect mRNA from degradation by exonucleases
Assist ribosome binding for translation initiation
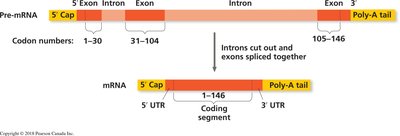
RNA Splicing and the Spliceosome
RNA splicing removes introns from pre-mRNA and joins exons together. This process is catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs) and proteins. The spliceosome recognizes specific sequences at intron-exon boundaries, cuts the RNA, releases the intron, and ligates the exons.
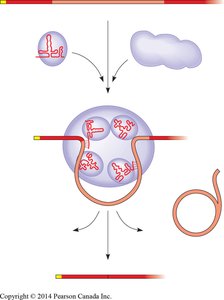
Significance of Introns and Alternative Splicing
Some introns contain regulatory sequences that influence gene expression.
Alternative splicing allows a single gene to produce multiple protein isoforms by varying the combination of exons included in the mature mRNA.
Human genes have an average of 8 introns, and the number can vary widely.
Example: Alternative splicing can result in different proteins being produced in liver cells versus fibroblasts from the same gene.
Additional info: Some evidence suggests that introns may have evolved from mobile self-splicing elements known as group II introns.
