 Back
BackTranscription and Translation: Gene Expression
Study Guide - Smart Notes

Transcription and Translation: Mechanisms of Gene Expression
Overview of the Central Dogma
The central dogma of molecular biology describes the flow of genetic information within a biological system: DNA is transcribed into RNA, which is then translated into protein. This process is fundamental to all living organisms and underlies gene expression and regulation.
Replication: DNA is copied to produce identical DNA molecules.
Transcription: DNA is used as a template to synthesize RNA.
Translation: RNA is used as a template to synthesize proteins.
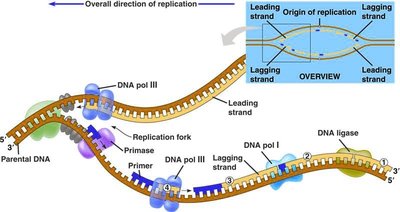
DNA Replication
Mechanism of DNA Replication
DNA replication is the process by which a cell duplicates its DNA before cell division. It is semi-conservative, meaning each new DNA molecule consists of one parental and one newly synthesized strand.
Enzymes involved: DNA polymerase III (main synthesis), primase (lays RNA primer), DNA polymerase I (replaces RNA primer with DNA), DNA ligase (joins Okazaki fragments).
Leading strand: Synthesized continuously in the 5' to 3' direction.
Lagging strand: Synthesized discontinuously as Okazaki fragments.
Error rate: Approximately one error per billion nucleotides.
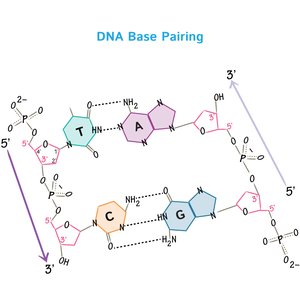
Base Pairing in DNA
Hydrogen bonds between nucleotide bases (A-T and G-C) ensure accurate DNA replication and transcription.
Adenine (A) pairs with Thymine (T) via two hydrogen bonds.
Guanine (G) pairs with Cytosine (C) via three hydrogen bonds.
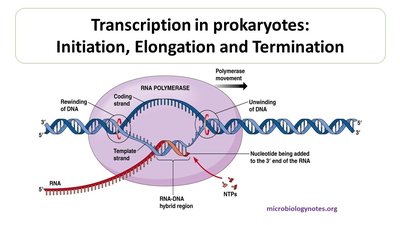
Transcription: DNA to RNA
Overview and Steps of Transcription
Transcription is the synthesis of RNA from a DNA template. It occurs in three main stages: initiation, elongation, and termination.
Initiation: RNA polymerase binds to the promoter region of DNA, unwinding the DNA strands.
Elongation: RNA polymerase moves along the template strand, synthesizing RNA in the 5' to 3' direction.
Termination: RNA polymerase releases the completed RNA transcript at a terminator sequence.
Initiation of Transcription
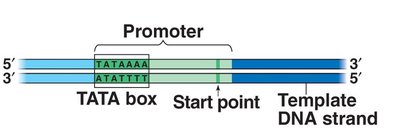
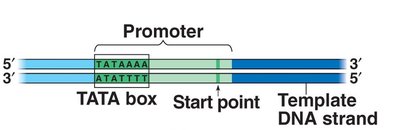
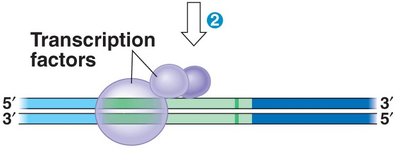
Transcription begins when RNA polymerase binds to a specific DNA sequence called the promoter. In eukaryotes, transcription factors are required for RNA polymerase II to bind and initiate transcription.
Promoter: DNA sequence where RNA polymerase attaches and initiates transcription (e.g., TATA box in eukaryotes).
Transcription factors: Proteins that help RNA polymerase recognize the promoter and regulate transcription.
Elongation of the RNA Transcript
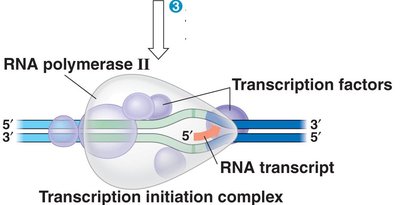
During elongation, RNA polymerase unwinds the DNA and adds complementary RNA nucleotides to the growing RNA strand. The DNA rewinds after the RNA polymerase passes.
Direction: RNA is synthesized in the 5' to 3' direction, using the 3' to 5' DNA template strand.
Base pairing: Adenine pairs with uracil (U) in RNA, and cytosine pairs with guanine.
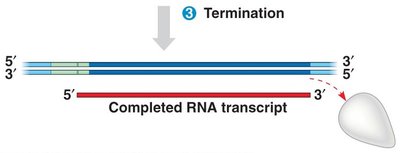
Termination of Transcription
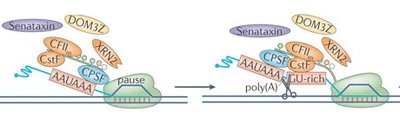
Transcription ends when RNA polymerase encounters a terminator sequence (in prokaryotes) or a polyadenylation signal (in eukaryotes). The RNA transcript is released, and the polymerase detaches from the DNA.
Prokaryotes: Terminator sequence signals the end of transcription.
Eukaryotes: Polyadenylation signal (AAUAAA) in pre-mRNA triggers cleavage and release of the transcript.
RNA Processing in Eukaryotes
Modification of Pre-mRNA
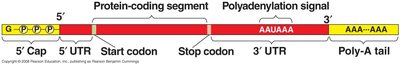
In eukaryotes, the primary RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA ready for translation.
5' Cap: Addition of a modified guanine nucleotide to the 5' end.
Poly-A Tail: Addition of 50-250 adenine nucleotides to the 3' end.
Functions: Facilitate export from the nucleus, protect mRNA from degradation, and assist ribosome binding.
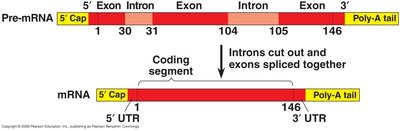
RNA Splicing: Removal of Introns
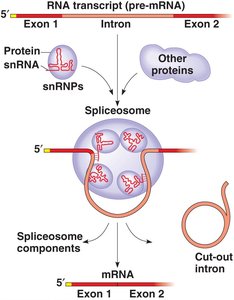
RNA splicing removes non-coding regions (introns) from pre-mRNA and joins coding regions (exons) together. This process is catalyzed by the spliceosome, a complex of proteins and small nuclear RNAs (snRNAs).
Introns: Non-coding sequences removed from pre-mRNA.
Exons: Coding sequences that remain in mature mRNA.
Spliceosome: Complex that catalyzes splicing, involving snRNPs and snRNAs.
Ribozymes: RNA molecules with catalytic activity, such as those in the spliceosome.
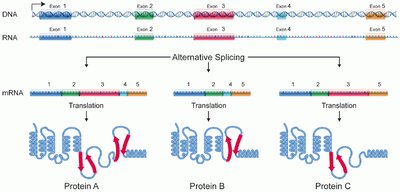
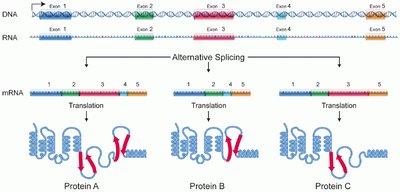
Alternative Splicing
Alternative splicing allows a single gene to code for multiple proteins by varying the combination of exons included in the final mRNA. This increases protein diversity and is crucial for processes such as immune system function.
Example: Antibody diversity is generated through alternative splicing in immune cells.
Translation: RNA to Protein
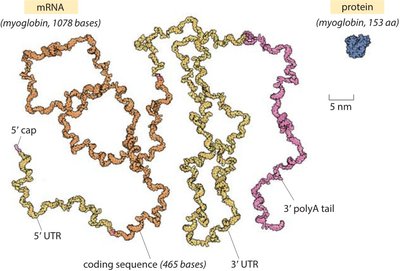
Genetic Code and Codons
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. The genetic code is read in sets of three nucleotides (codons), each specifying a particular amino acid.
Codon: Sequence of three mRNA nucleotides that codes for an amino acid.
tRNA: Transfer RNA molecules bring amino acids to the ribosome, matching codons with their anticodons.
rRNA: Ribosomal RNA forms the core of the ribosome and catalyzes peptide bond formation.
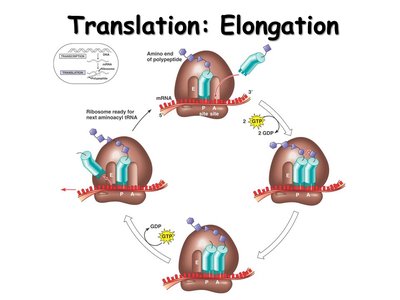
Mechanism of Translation
Translation occurs in three stages: initiation, elongation, and termination. Accurate translation requires correct matching of tRNA and amino acids, and proper codon-anticodon pairing.
Initiation: Ribosome assembles around the start codon of mRNA.
Elongation: Amino acids are added one by one to the growing polypeptide chain.
Termination: Translation ends at a stop codon, releasing the completed polypeptide.
Error rate: About one incorrect amino acid per thousand codons translated.
Prokaryotic vs. Eukaryotic Gene Expression
There are key differences in gene expression between prokaryotes and eukaryotes, particularly in the timing and location of transcription and translation.
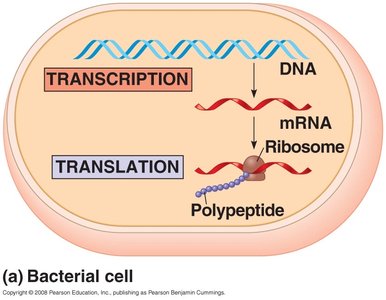
Prokaryotes: Transcription and translation occur simultaneously in the cytoplasm.
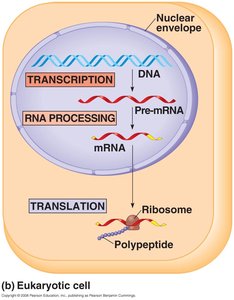
Eukaryotes: Transcription occurs in the nucleus; translation occurs in the cytoplasm after RNA processing.
Summary Table: Error Rates in Genetic Information Flow
Process | Template | Product | Error Rate |
|---|---|---|---|
Replication | DNA | DNA | 1 per 1,000,000,000 |
Transcription | DNA | RNA | 1 per 1,000,000 |
Translation | RNA | Protein | 1 per 1,000 |
Key Terms and Concepts
Promoter: DNA sequence where transcription begins.
Terminator: DNA sequence signaling the end of transcription (prokaryotes).
Polyadenylation signal: Sequence in eukaryotic pre-mRNA signaling cleavage and poly-A tail addition.
Spliceosome: Complex responsible for removing introns from pre-mRNA.
Codon: Three-nucleotide sequence in mRNA specifying an amino acid.
Anticodon: Three-nucleotide sequence in tRNA complementary to an mRNA codon.
Wobble: Flexibility in base pairing at the third position of the codon, allowing some tRNAs to pair with multiple codons.
Further Reading and Practice
Review Chapter 17: Gene Expression (Transcription & Translation).
Practice reading scientific literature and solving problems related to transcription and translation.
