 Back
BackTranscription, RNA Processing, and Translation: Study Notes
Study Guide - Smart Notes

Transcription, RNA Processing, and Translation
Overview of Transcription
Transcription is the process by which RNA polymerases synthesize an RNA copy of the genetic instructions stored in DNA. This process is fundamental to gene expression and is the first step in the central dogma of molecular biology.
RNA polymerases use ribonucleoside triphosphates (NTPs) to build RNA molecules complementary to one DNA strand.
Only one DNA strand serves as the template strand; the other is the non-template (coding) strand, which matches the RNA sequence except that RNA contains uracil (U) instead of thymine (T).
RNA polymerases perform template-directed synthesis in the 5' to 3' direction.
Unlike DNA polymerases, RNA polymerases do not require a primer to begin transcription.
Bacteria have a single RNA polymerase, while eukaryotes have at least three distinct types.
Initiation of Transcription in Bacteria
Transcription initiation is the first phase, requiring specific protein factors and DNA sequences to begin RNA synthesis.
RNA polymerase cannot initiate transcription alone; in bacteria, a sigma protein must bind to it, forming a holoenzyme.
Sigma recognizes promoters, which are specific DNA sequences where transcription begins.
Bacterial promoters are typically 40–50 base pairs long and contain conserved sequences:
-10 box: TATAAT sequence, located about 10 bases upstream of the transcription start site.
-35 box: TTGACA sequence, about 35 bases upstream.
Promoter orientation determines which DNA strand is used as the template and the direction of RNA polymerase movement.
Events Inside the Holoenzyme
Once the holoenzyme is assembled at the promoter, several key events occur:
RNA polymerase opens the DNA double helix, creating a transcription bubble.
The template strand is threaded through the active site, and incoming NTPs pair with complementary DNA bases, initiating RNA synthesis.
Elongation and Termination in Bacteria
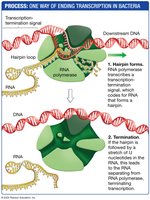
During elongation, RNA polymerase moves along the DNA template, adding nucleotides to the 3' end of the growing RNA molecule. Termination occurs when a specific sequence signals the end of transcription.
Elongation: RNA polymerase synthesizes RNA in the 5' to 3' direction.
Termination: Occurs when RNA polymerase transcribes a transcription-termination signal that codes for RNA forming a hairpin structure. This structure causes the polymerase to dissociate from the DNA and release the RNA transcript.
Transcription in Eukaryotes
Eukaryotic transcription is more complex than in bacteria, involving additional enzymes and regulatory sequences.
Three types of RNA polymerases (I, II, III) transcribe different classes of genes.
Promoters are larger and more diverse, often containing a TATA box.
General transcription factors (not sigma proteins) recognize promoters and help initiate transcription.
Termination involves a poly(A) signal rather than a hairpin; the RNA is cleaved downstream of this signal.
Transcription occurs in the nucleus, while translation occurs in the cytoplasm.
RNA Processing in Eukaryotes
Primary Transcripts and RNA Processing
In eukaryotes, the initial RNA product, called a primary transcript or pre-mRNA, must be processed before it can be translated into protein.
Primary transcripts undergo RNA processing to become mature mRNA.
In bacteria, RNA transcripts are typically functional without further modification.
Discovery of Split Eukaryotic Genes
In 1977, it was discovered that eukaryotic genes contain noncoding sequences called introns interspersed with coding sequences called exons.
Experiments showed that stretches of DNA are not present in mature mRNA, indicating the presence of introns.
Introns are removed during RNA processing; exons remain in the final mRNA.
RNA Splicing
RNA splicing removes introns from the primary transcript and joins exons together. This process is catalyzed by a complex called the spliceosome.
Spliceosomes are made of small nuclear ribonucleoproteins (snRNPs), which contain small nuclear RNAs (snRNAs) and proteins.
Splicing allows for the production of different mRNAs and proteins from a single gene (alternative splicing).
Four steps of splicing:
snRNPs bind to exon–intron and intron–exon boundaries and to an A nucleotide near the end of the intron.
Other snRNPs join to form the spliceosome.
The intron forms a lariat structure (loop with a stem) with A as the branch point.
The lariat is cut out, exons are joined, and the intron is degraded.
Adding Caps and Tails to Transcripts
Two additional modifications are made to pre-mRNA to produce mature mRNA:
5' Cap: A modified guanine nucleotide is added to the 5' end, enabling ribosome binding and protecting the RNA from degradation.
Poly(A) Tail: 100–250 adenine nucleotides are added to the 3' end after cleavage, aiding in translation and stability.
After splicing and addition of the cap and tail, the mature mRNA contains untranslated regions (UTRs) at both ends.
Additional info: The mature mRNA is then exported from the nucleus to the cytoplasm, where it serves as a template for protein synthesis during translation.
