 Back
BackTranscriptional and Translational Regulation in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcriptional Regulation in Prokaryotes
Overview of Gene Regulation
Gene expression in prokaryotes is tightly regulated to ensure that proteins are produced only when needed. This regulation allows bacteria to adapt quickly to changing environments by turning genes on or off in response to environmental signals.
Gene expression refers to the process by which information from a gene is used to synthesize a functional gene product, typically a protein.
Regulation occurs at multiple levels, but transcriptional control is the most common and efficient in prokaryotes.
The Lac Operon: A Model for Negative Regulation
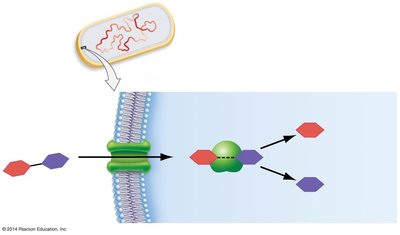
The lac operon in Escherichia coli is a classic example of transcriptional regulation. It controls the metabolism of lactose through the coordinated expression of genes required for lactose uptake and breakdown.
lacZ: Encodes β-galactosidase, which cleaves lactose into glucose and galactose.
lacY: Encodes galactoside permease, a membrane protein that imports lactose into the cell.
lacI: Encodes the repressor protein that regulates the operon.

Negative Control of the Lac Operon
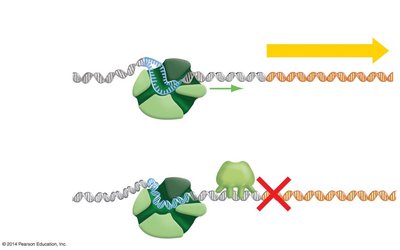
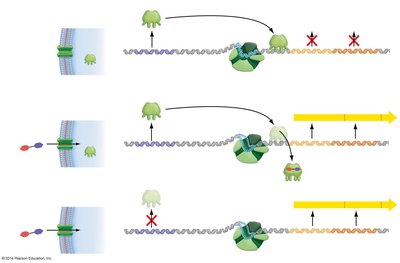
Negative control involves a regulatory protein (repressor) that binds to the operator region of the operon, blocking transcription when lactose is absent.
When lactose is absent, the repressor binds to the operator, preventing RNA polymerase from transcribing the operon.
When lactose is present, it acts as an inducer by binding to the repressor, causing it to release from the operator. This allows transcription to proceed.
If the repressor is nonfunctional (constitutive mutant), transcription occurs regardless of lactose presence.
Positive Control: The Ara Operon
Positive control occurs when a regulatory protein (activator) increases transcription. The ara operon, responsible for arabinose metabolism, is regulated by the AraC protein, which acts as an activator in the presence of arabinose.
When arabinose is present, AraC binds to the initiator region and promotes transcription of the araBAD genes.

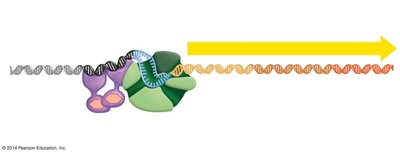
Types of Transcriptional Control
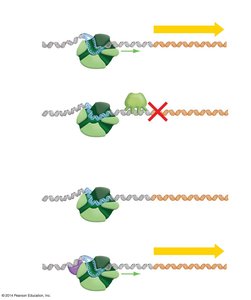
Negative control: Regulatory protein (repressor) shuts down transcription.
Positive control: Regulatory protein (activator) triggers transcription.
Translation in Prokaryotes
Initiation of Translation
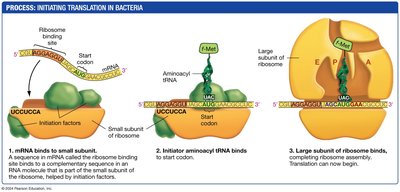
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. In bacteria, translation initiation involves the assembly of the ribosome on the mRNA, guided by specific sequences and initiation factors.
The small ribosomal subunit binds to the mRNA at the ribosome binding site.
The initiator tRNA carrying formylmethionine (fMet) binds to the start codon.
The large ribosomal subunit joins, completing the initiation complex.
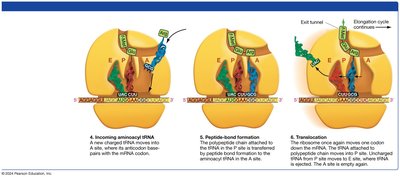
Elongation Phase of Translation
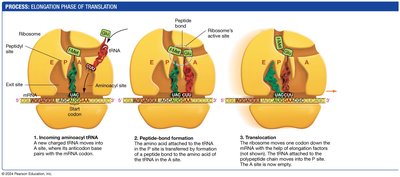
During elongation, amino acids are sequentially added to the growing polypeptide chain.
Aminoacyl-tRNA enters the A site of the ribosome, matching its anticodon to the mRNA codon.
A peptide bond forms between the amino acid in the A site and the growing chain in the P site.
The ribosome translocates, moving the tRNA from the A site to the P site, and the empty tRNA exits from the E site.
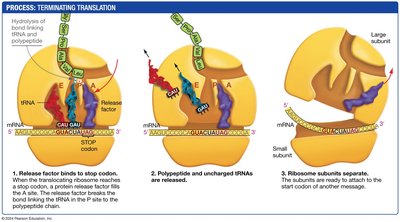
Termination of Translation
Termination occurs when a stop codon is encountered. A release factor binds to the stop codon, causing the release of the newly synthesized polypeptide and the disassembly of the ribosome.
Release factors recognize stop codons and promote hydrolysis of the bond linking the polypeptide to the tRNA.
The ribosomal subunits dissociate, ready to initiate translation on another mRNA.
Transcriptional Regulation in Eukaryotes
Overview of Gene Regulation in Eukaryotes
Eukaryotic gene expression is regulated at multiple levels, including chromatin remodeling, transcription initiation, RNA processing, mRNA stability, translation, and post-translational modification. This allows for complex and precise control of gene activity.
Chromatin structure affects accessibility of DNA to transcription machinery.
Regulatory sequences such as enhancers and promoter-proximal elements modulate transcription initiation.
Alternative splicing and mRNA stability further diversify gene expression.
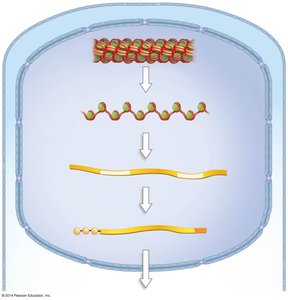
Chromatin Structure and Remodeling
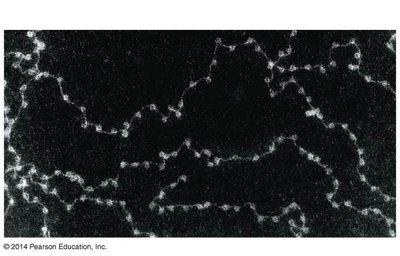
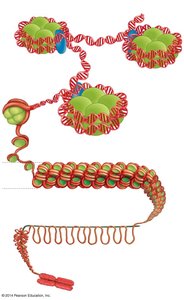
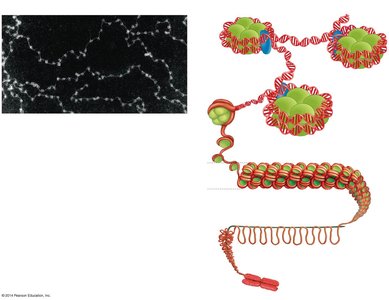
DNA in eukaryotic cells is packaged into chromatin, which consists of DNA wrapped around histone proteins to form nucleosomes. Chromatin can be remodeled to either condense (silence genes) or decondense (activate genes).
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around a histone octamer.
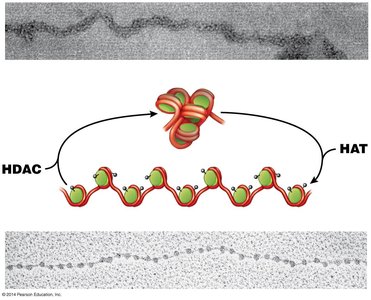
Chromatin remodeling complexes and chemical modifications (e.g., acetylation) regulate gene accessibility.
Epigenetic Effects and Environmental Influence
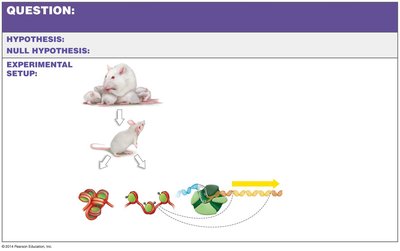
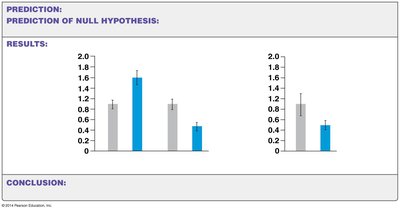
Epigenetic modifications, such as histone modification and DNA methylation, can be influenced by environmental factors like maternal diet, affecting gene expression in offspring.
Protein deprivation in mothers can lead to abnormal histone modifications in offspring, altering gene expression patterns.

Regulatory Elements in Eukaryotic Transcription
Eukaryotic genes are regulated by multiple DNA elements:
Promoter: The site where RNA polymerase binds to initiate transcription.
Promoter-proximal elements: Regulatory sequences close to the promoter that bind specific transcription factors.
Enhancers: Distant regulatory sequences that increase transcription when bound by activator proteins.
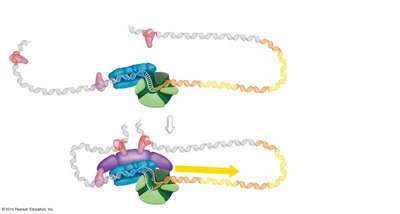
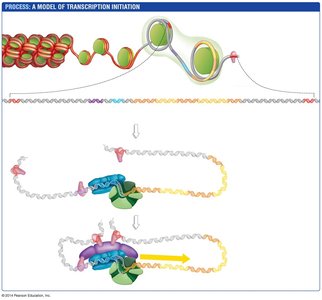
Transcription Initiation in Eukaryotes
Transcription initiation involves the assembly of a complex of proteins at the promoter, including basal transcription factors, activators, and the mediator complex. DNA looping brings enhancers into contact with the promoter region, facilitating transcription initiation.
Chromatin remodeling exposes regulatory sequences.
Assembly of transcription factors and mediator complex enables RNA polymerase II to begin transcription.
Alternative Splicing
Alternative splicing allows a single gene to produce multiple mRNA variants, leading to the synthesis of different protein isoforms from the same gene. This increases the diversity of proteins in eukaryotic cells.
Exons can be included or excluded from the final mRNA, depending on the splicing pattern.
This process is regulated by splicing factors and is crucial for tissue-specific gene expression.
Summary Table: Comparison of Prokaryotic and Eukaryotic Gene Regulation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Chromatin Structure | Absent | Present, regulates accessibility |
Regulatory Elements | Operators, promoters | Promoters, enhancers, silencers |
Transcription Factors | Few, often repressors or activators | Many, complex combinations |
RNA Processing | Minimal | Extensive (capping, splicing, polyadenylation) |
Location of Transcription & Translation | Cytoplasm (coupled) | Nucleus (transcription), cytoplasm (translation) |
