 Back
BackTranscriptional Control of Gene Expression in Eukaryotes
Study Guide - Smart Notes

Transcriptional Control of Gene Expression
Overview of Eukaryotic Transcription
Transcriptional control is the primary mechanism by which eukaryotic cells regulate gene expression. Eukaryotes possess three distinct nuclear RNA polymerases, each responsible for synthesizing different classes of RNA, which are essential for various cellular functions.
RNA polymerase I: Synthesizes pre-rRNA, which is processed into 28S, 18S, and 5.8S rRNAs, the main components of ribosomes.
RNA polymerase II: Synthesizes mRNAs (protein-coding), small nuclear RNAs (snRNAs) involved in splicing, and regulatory RNAs such as miRNAs and siRNAs.
RNA polymerase III: Synthesizes tRNAs, 5S rRNA, and other small stable RNAs.
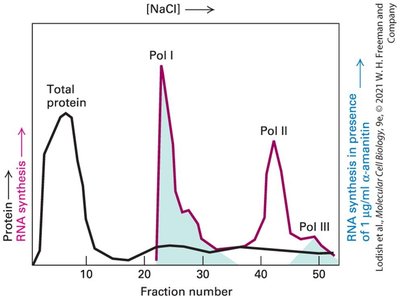
Example: The majority of protein-coding genes are transcribed by RNA polymerase II, while tRNAs are transcribed by RNA polymerase III.
RNA Polymerase II Promoters and General Transcription Factors
Transcription initiation by RNA polymerase II requires specific DNA sequences called promoters and the assembly of general transcription factors. There are three main types of promoter sequences in eukaryotic DNA:
TATA box: Common in highly transcribed genes.
Initiator (Inr) elements: Found in some genes, often overlapping the transcription start site.
CpG islands: Characteristic of about 70% of vertebrate protein-coding gene promoters, typically associated with genes transcribed at low rates.
The sequential binding of general transcription factors (e.g., TFIID, TFIIB, TFIIE, TFIIF, TFIIH) is required to recruit RNA polymerase II and initiate transcription.
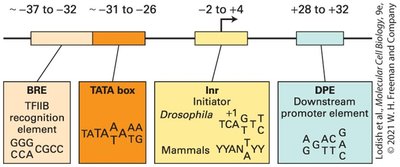
Example: The TATA box is located approximately 25-30 base pairs upstream of the transcription start site in many genes.
Regulatory Sequences in Protein-Coding Genes and the Proteins Through Which They Function
Gene expression in eukaryotes is regulated by multiple protein-binding DNA regions, which can be located at varying distances from the transcription start site. These include:
Promoters: Direct RNA polymerase II binding and determine transcription initiation site and frequency.
Promoter-proximal elements: Located near the promoter, often cell-type-specific.
Enhancers: Can be located far from the gene, increase transcription in a cell-type-specific manner.
Transcription activators and repressors are modular proteins with distinct DNA-binding and activation/repression domains.
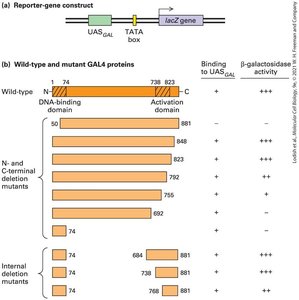
Example: The GAL4 protein in yeast contains a DNA-binding domain and an activation domain, each responsible for specific functions.
Molecular Mechanisms of Transcription Repression and Activation
Eukaryotic transcription activators and repressors regulate gene expression by interacting with multisubunit co-activators or co-repressors. These complexes can modulate chromatin structure or interact directly with RNA polymerase II and general transcription factors. The assembly of the preinitiation complex is highly cooperative and often requires several activators, which are produced in a cell-type-specific manner.
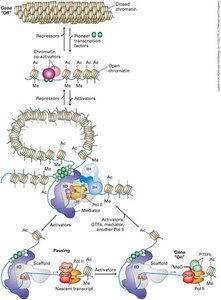
Example: The Mediator complex acts as a co-activator, bridging transcription factors and RNA polymerase II.
Regulation of Transcription-Factor Activity
Transcription factor activity is regulated by intracellular signaling pathways, often initiated by cell-surface receptor activation. Lipid-soluble hormones can diffuse into cells and bind to intracellular receptors, which then regulate gene expression by binding to specific DNA response elements. DNase I hypersensitive sites (DHSs) mark regions of open chromatin accessible to transcription factors, and their positions can change during development. Additionally, the elongation phase of RNA polymerase II is subject to regulation.
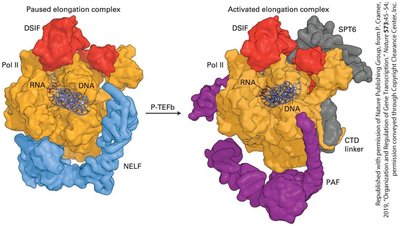
Example: The HIV Tat protein recruits P-TEFb to phosphorylate RNA polymerase II, promoting elongation of viral transcripts.
Epigenetic Regulation of Transcription
Epigenetic mechanisms maintain transcriptional states through cell division. These include DNA methylation and histone modifications. Polycomb group complexes maintain gene repression, while Trithorax group complexes maintain gene activation. In female mammals, X-chromosome inactivation is mediated by the long noncoding RNA Xist.
Modification | Sites of Modification | Effect on Transcription |
|---|---|---|
Acetylated lysine | H3 (K9, K14, K18, K23, K27); H4 (K5, K8, K12, K16); H2A (K5, K8, K13); H2B (K5, K12, K15, K20) | Activation |
Hypoacetylated lysine | Various | Repression |
Phosphorylated serine/threonine | H3 (T3, S10, T11, S28); H2A (S1, T120); H2B (S14) | Activation |
Methylated arginine | H3 (R2, R17, R26); H4 (R3) | Activation |
Methylated lysine | H3 (K4) Me3 (promoter); H3 (K4) Me1 (enhancer); H3 (K36, K79) (transcribed region); H3 (K9, K27) Me2/3; H4 (K20) Me2/3 | Activation (K4, K36, K79); Repression (K9, K27, K20) |
Ubiquitinylated lysine | H2B (K120 in mammals, K123 in S. cerevisiae); H2A (K119 in mammals) | Activation (H2B); Repression (H2A) |
Example: Acetylation of histone H3 at lysine 9 (H3K9ac) is associated with active gene transcription.
Other Eukaryotic Transcription Systems
RNA polymerase I and III have transcription initiation mechanisms similar to RNA polymerase II but require distinct sets of general transcription factors. RNA polymerase I transcribes the 45S pre-rRNA, which is processed into 18S, 5.8S, and 28S rRNAs. RNA polymerase III transcribes tRNAs, 5S rRNA, and other small RNAs.
Example: The 5S rRNA gene is transcribed by RNA polymerase III, while the 18S rRNA gene is transcribed by RNA polymerase I.
Summary Table: Classes of RNA Transcribed by Eukaryotic RNA Polymerases
Polymerase | RNA Transcribed | RNA Function |
|---|---|---|
RNA polymerase I | Pre-rRNA (28S, 18S, 5.8S rRNAs) | Ribosome components, protein synthesis |
RNA polymerase II | mRNA, snRNAs, siRNAs, miRNAs | Protein coding, RNA splicing, translation control, chromatin repression |
RNA polymerase III | tRNAs, 5S rRNA, snRNA U6, 7S RNA, other small RNAs | Protein synthesis, ribosome component, RNA splicing, signal recognition, various |
