 Back
BackStudy guide 24
Study Guide - Smart Notes

Transcriptional Regulation in Prokaryotes
Overview of Gene Regulation in Bacteria
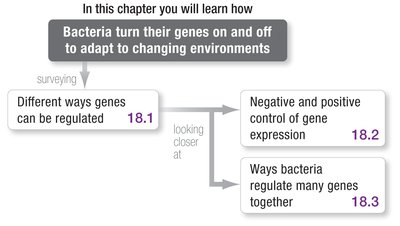
Gene regulation in bacteria allows cells to adapt to changing environments by turning genes on or off as needed. This process is essential for efficient resource use and survival.
Gene regulation refers to the mechanisms that increase or decrease the expression of specific genes.
Bacteria use regulatory proteins and DNA sequences to control gene expression in response to environmental signals.
Negative and Positive Control of Gene Expression
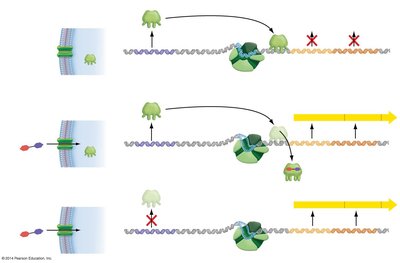
Gene expression in bacteria can be regulated by negative or positive control mechanisms:
Negative control: A repressor protein binds to DNA and blocks transcription.
Positive control: An activator protein increases the likelihood of transcription.
Example: The lac operon in Escherichia coli is a classic model of negative regulation, where the LacI repressor binds to the operator to prevent transcription in the absence of lactose.
Transcriptional Regulation in Eukaryotes
Overview of Gene Regulation in Eukaryotes
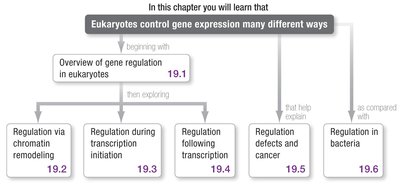
Eukaryotic gene expression is regulated at multiple levels, including chromatin remodeling, transcription initiation, RNA processing, and post-translational modifications. This complexity allows for precise spatial and temporal control of gene activity.
Regulation via chromatin remodeling (making DNA more or less accessible for transcription)
Regulation during transcription initiation (assembly of transcription machinery at promoters)
Regulation following transcription (RNA processing, stability, and translation)
Chromatin Structure and Remodeling
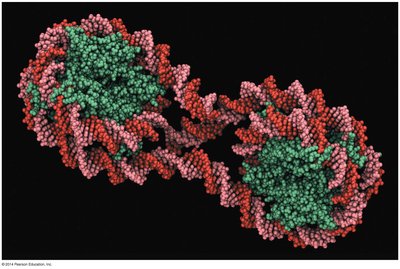
In eukaryotes, DNA is packaged with histone proteins into nucleosomes, forming chromatin. The structure of chromatin can be modified to regulate gene accessibility:
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around a core of eight histone proteins.
Chromatin remodeling: Chemical modifications (e.g., acetylation, methylation) of histones or DNA alter chromatin structure, influencing gene expression.
Levels of Chromatin Organization
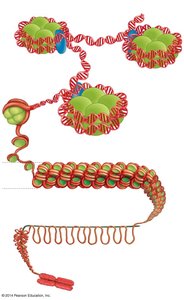
Chromatin exists in various levels of compaction, affecting gene accessibility:
DNA wraps around histones to form nucleosomes.
Nucleosomes coil to form a 30-nm fiber, which further loops and folds to form higher-order structures.
Highly condensed chromatin (heterochromatin) is generally transcriptionally inactive, while less condensed chromatin (euchromatin) is accessible for transcription.
Epigenetic Regulation
Epigenetic modifications, such as DNA methylation and histone modification, can be inherited and affect gene expression without altering the DNA sequence. Environmental factors, such as nutrition, can influence these modifications and impact offspring phenotype.
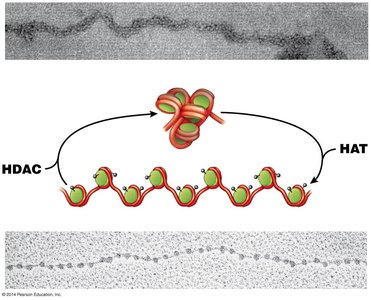
Histone acetylation (by HAT enzymes) generally promotes open chromatin and active transcription.
Histone deacetylation (by HDAC enzymes) leads to condensed chromatin and gene silencing.
Transcription Initiation in Eukaryotes
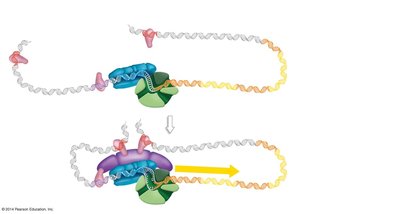
Transcription initiation involves the assembly of multiple proteins at the promoter region of a gene:
Enhancers: DNA sequences that increase transcription when bound by activator proteins.
Promoter-proximal elements: Regulatory sequences near the promoter that help recruit transcription factors.
Basal transcription factors: Proteins required for the assembly of the transcription initiation complex.
Mediator complex: Bridges regulatory proteins and RNA polymerase II, facilitating transcription initiation.
Summary Table: Comparison of Prokaryotic and Eukaryotic Gene Regulation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
DNA Packaging | Minimal (no nucleosomes) | Extensive (nucleosomes, chromatin) |
Regulatory Elements | Operators, promoters | Promoters, enhancers, silencers |
Regulatory Proteins | Repressors, activators | Transcription factors, coactivators, repressors |
Gene Organization | Operons (polycistronic mRNA) | Mostly monocistronic mRNA |
Epigenetic Regulation | Rare | Common (DNA/histone modifications) |
