 Back
BackTranslation: The RNA-Directed Synthesis of Polypeptides
Study Guide - Smart Notes

Translation: The RNA-Directed Synthesis of Polypeptides
Overview of Translation
Translation is the cellular process in which genetic information encoded in messenger RNA (mRNA) is used to assemble a specific sequence of amino acids, forming a polypeptide. This process occurs in both prokaryotic and eukaryotic cells and is fundamental to gene expression.
mRNA provides the template, with codons specifying amino acids.
tRNA molecules serve as adaptors, matching amino acids to codons via their anticodons.
Ribosomes are the molecular machines that facilitate the decoding of mRNA and the synthesis of polypeptides.
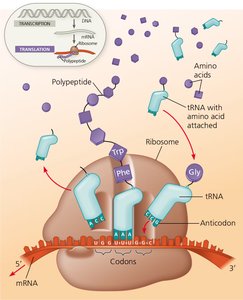
Molecular Components of Translation
The translation process involves several key molecular players:
mRNA: Contains codons, each specifying an amino acid.
tRNA: Transfers specific amino acids to the ribosome; each tRNA has an anticodon complementary to an mRNA codon.
Ribosome: Composed of rRNA and proteins, it coordinates the interaction between mRNA and tRNA and catalyzes peptide bond formation.
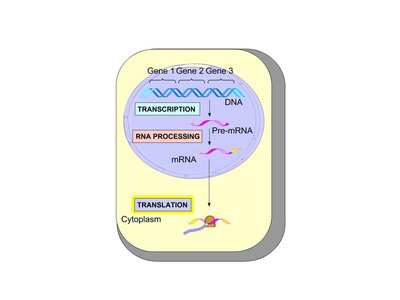
The Structure and Function of Transfer RNA (tRNA)
tRNA Structure
tRNA molecules are single RNA strands (~80 nucleotides) that fold into a characteristic cloverleaf structure due to internal base pairing. The actual three-dimensional shape is L-shaped, with the amino acid attachment site at one end and the anticodon at the other.
Amino Acid Attachment Site: Located at the 3' end, where the specific amino acid is covalently attached.
Anticodon: A triplet of bases that pairs with the complementary mRNA codon.
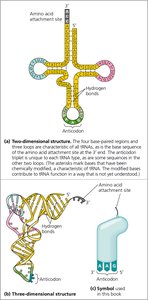
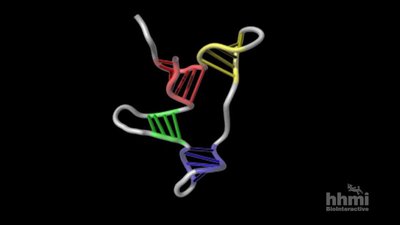
tRNA Function
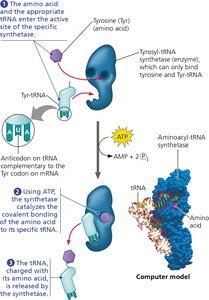
Each tRNA is specific for one amino acid and one or more codons. The correct amino acid is attached to its tRNA by a specific enzyme, ensuring fidelity in translation.
Aminoacyl-tRNA Synthetase: Enzyme that catalyzes the attachment of an amino acid to its corresponding tRNA, using ATP hydrolysis.
There are 20 different synthetases, one for each amino acid.
Wobble Hypothesis
The genetic code is degenerate, meaning some amino acids are specified by more than one codon. The 'wobble' at the third position of the codon allows some tRNAs to pair with more than one codon, increasing efficiency.
The Structure and Function of Ribosomes
Ribosome Structure
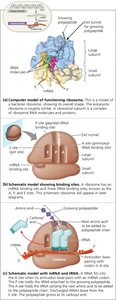
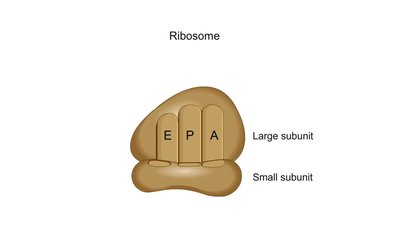
Ribosomes are composed of two subunits (large and small), each made of rRNA and proteins. They have three binding sites for tRNA:
A site (Aminoacyl-tRNA site): Holds the tRNA carrying the next amino acid to be added.
P site (Peptidyl-tRNA site): Holds the tRNA with the growing polypeptide chain.
E site (Exit site): Where discharged tRNAs leave the ribosome.
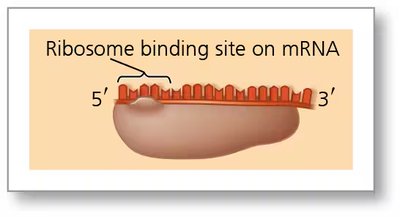
Ribosome Binding Sites on mRNA
Ribosomes recognize specific sequences on mRNA to initiate translation. In bacteria, the ribosome binding site is upstream of the start codon (AUG).
Sequence Logos and Ribosome Binding Sites
Sequence logos graphically represent conserved sequences, such as ribosome binding sites, across multiple genes. The height of each letter indicates the frequency of that base at a given position.
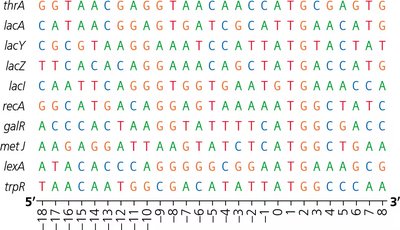
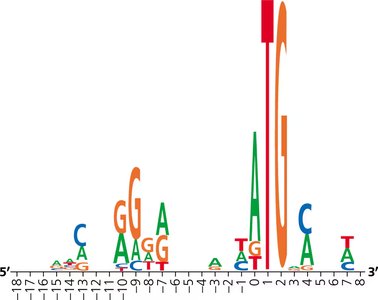
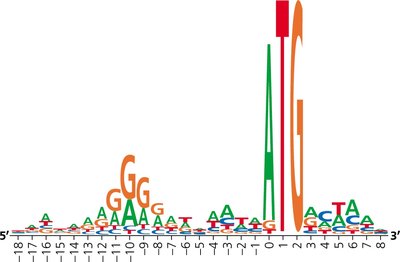
Stages of Translation
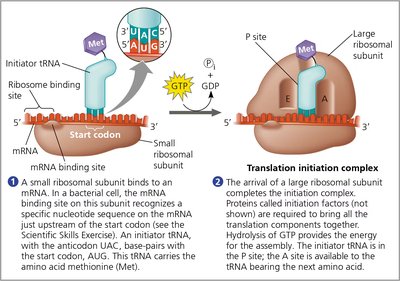
Initiation
Translation initiation brings together mRNA, a tRNA with the first amino acid (methionine), and the ribosomal subunits. Initiation factors and GTP hydrolysis are required for assembly.
In bacteria, the small subunit binds the mRNA and initiator tRNA at the ribosome binding site.
In eukaryotes, the small subunit binds the 5' cap and scans for the start codon.
Elongation
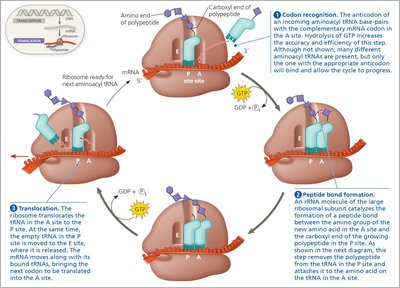
During elongation, amino acids are added one by one to the growing polypeptide chain. Each cycle involves:
Codon Recognition: tRNA anticodon pairs with mRNA codon in the A site.
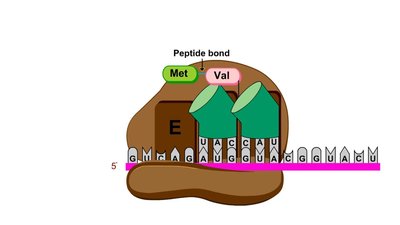
Peptide Bond Formation: Peptide bond forms between the new amino acid and the growing chain.
Translocation: Ribosome moves along mRNA, shifting tRNAs from A to P to E sites.
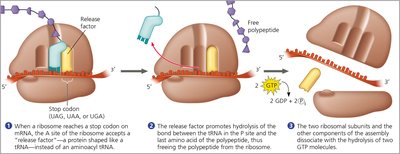
Termination
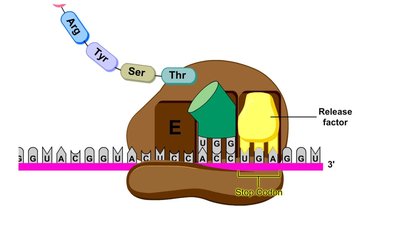
Termination occurs when a stop codon (UAG, UAA, or UGA) is reached. A release factor binds to the stop codon, causing the addition of a water molecule, which releases the completed polypeptide and disassembles the translation complex.
Protein Folding and Post-Translational Modifications
Folding and Modifications
Newly synthesized polypeptides fold into their functional three-dimensional shapes, sometimes with the help of chaperone proteins. Additional modifications may include cleavage, addition of chemical groups (e.g., sugars, phosphates), or assembly into multi-subunit complexes.
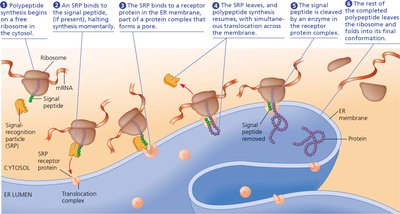
Targeting Polypeptides to Specific Locations
Proteins destined for secretion or for specific organelles contain signal peptides that direct their cellular localization. The signal-recognition particle (SRP) guides ribosomes to the endoplasmic reticulum (ER) for co-translational import of proteins into the ER.
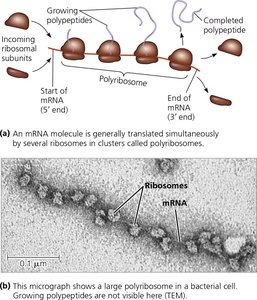
Polyribosomes and Coupled Transcription-Translation
Polyribosomes
Multiple ribosomes can simultaneously translate a single mRNA, forming a polyribosome (polysome), which increases the efficiency of protein synthesis.
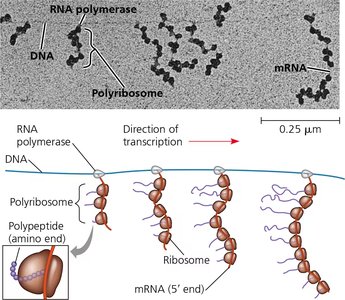
Coupled Transcription and Translation in Bacteria
In bacteria, transcription and translation are coupled, allowing ribosomes to begin translating mRNA while it is still being synthesized. This is possible due to the lack of a nuclear envelope.
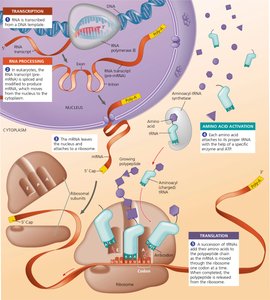
Transcription and Translation in Eukaryotes
In eukaryotes, transcription occurs in the nucleus, and mRNA is processed before being exported to the cytoplasm for translation. This compartmentalization allows for additional regulation of gene expression.
Summary Table: Key Steps and Components of Translation
Step | Main Components | Key Events |
|---|---|---|
Initiation | mRNA, initiator tRNA, ribosomal subunits, initiation factors, GTP | Assembly of initiation complex at start codon |
Elongation | Elongation factors, aminoacyl-tRNAs, GTP | Codon recognition, peptide bond formation, translocation |
Termination | Release factor, stop codon, GTP | Release of polypeptide, disassembly of complex |
Key Terms and Concepts
Codon: A sequence of three nucleotides in mRNA that specifies an amino acid.
Anticodon: A sequence of three nucleotides in tRNA complementary to an mRNA codon.
Aminoacyl-tRNA Synthetase: Enzyme that attaches the correct amino acid to its tRNA.
Polyribosome (Polysome): A group of ribosomes translating the same mRNA simultaneously.
Wobble: Flexible base pairing at the third codon position, allowing one tRNA to recognize multiple codons.
Signal Peptide: Short peptide sequence that directs the transport of a protein to specific cellular locations.
Equations and Reactions
Aminoacylation of tRNA:
Peptide Bond Formation:
Additional info: The notes above integrate foundational concepts from molecular biology, including the central dogma, and highlight differences between prokaryotic and eukaryotic translation. The included images directly reinforce the structure and function of translation machinery, the process of translation, and the regulation of protein targeting.
