 Back
BackAmino Acids and Proteins: Structure, Classification, and Properties
Study Guide - Smart Notes

Amino Acids and Proteins
Introduction to Amino Acids
Amino acids are the fundamental building blocks of proteins, each containing an amino group, a carboxyl group, and a unique side chain (R group) attached to a central α-carbon. At physiological pH, amino acids exist as zwitterions, bearing both positive and negative charges. There are 20 standard α-amino acids that serve as the monomers for proteins in living organisms. Each amino acid is distinguished by the structure of its R group.
Zwitterion: A molecule with both positive and negative charges, but overall neutral.
α-Amino acids: The amino group is attached to the α-carbon, which is also bonded to the carboxyl group and the R group.
Unique feature: The type of R group bonded to the α-carbon makes each amino acid unique.
Example: Glycine is unique because its R group is a hydrogen atom, while proline has a cyclic structure.
Amino Acid Abbreviations and Codes
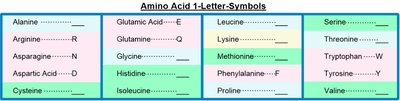
For convenience, amino acids are represented by both three-letter and one-letter codes. These abbreviations are standardized and widely used in biochemistry and molecular biology.
Three-letter codes: Usually the first three letters of the amino acid's name (e.g., Ala for Alanine, Gly for Glycine).
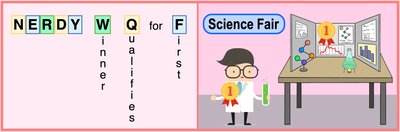
One-letter codes: A single letter, often the first letter of the name, but sometimes chosen for phonetic or frequency reasons (e.g., K for Lysine, W for Tryptophan).
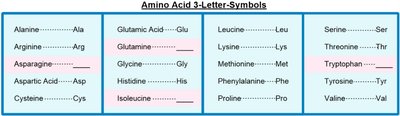
Mnemonic: For three-letter codes, most are straightforward, but some are altered for clarity (e.g., Asn for Asparagine, Gln for Glutamine, Trp for Tryptophan, Ile for Isoleucine).
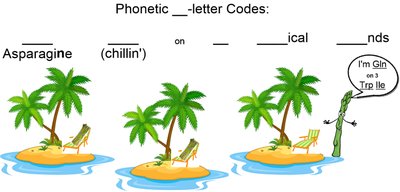
One-letter codes can be phonetic or based on the most common amino acid for a given letter. For example, "N" is used for Asparagine, "Q" for Glutamine, and "W" for Tryptophan.

Classification of Amino Acids
Nonpolar Amino Acids
Nonpolar amino acids have hydrophobic R groups that do not interact with water. These residues are typically found in the interior of proteins, away from aqueous environments.
Hydrophobic: Water-repelling, nonpolar side chains.
Examples: Glycine (Gly), Alanine (Ala), Valine (Val), Leucine (Leu), Isoleucine (Ile), Methionine (Met), Proline (Pro), Phenylalanine (Phe), Tryptophan (Trp).

Polar Amino Acids
Polar amino acids have hydrophilic R groups that can form hydrogen bonds with water, making them likely to be found on the surface of proteins. Cysteine is an exception as it can form disulfide bonds.
Hydrophilic: Water-attracting, polar side chains.
Examples: Serine (Ser), Threonine (Thr), Tyrosine (Tyr), Cysteine (Cys), Asparagine (Asn), Glutamine (Gln).

Charged Amino Acids
Charged amino acids are polar and have side chains that are either positively (basic) or negatively (acidic) charged at physiological pH.
Acidic: Aspartate (Asp, D), Glutamate (Glu, E) – contain carboxylate groups.
Basic: Lysine (Lys, K), Arginine (Arg, R), Histidine (His, H) – contain amino groups.


Peptides and Protein Structure
Peptide Bonds and Peptides
Peptides are formed when two or more amino acids are linked by peptide (amide) bonds. The peptide bond forms between the carboxyl group of one amino acid and the amino group of another, releasing water (condensation reaction).
Peptide bond: Covalent bond linking amino acids in a chain.
Polypeptide: A chain of many amino acids.
Residue: Each amino acid unit within a peptide or protein.
Levels of Protein Structure
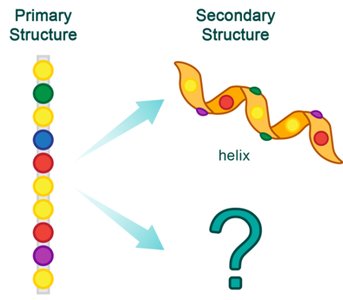
Primary Structure
The primary structure is the linear sequence of amino acids in a protein, held together by peptide bonds. It is always written from the N-terminus (free amino group) to the C-terminus (free carboxyl group).
Backbone: Repeating N–C–C sequence.
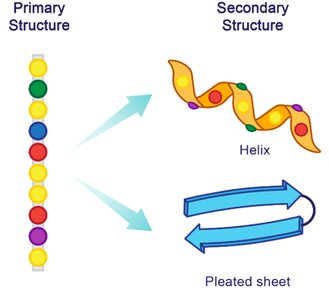
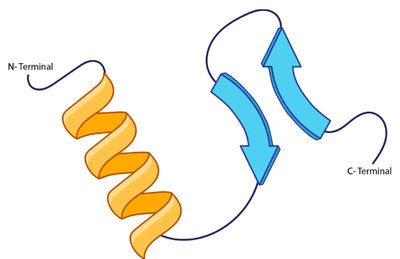
Secondary Structure
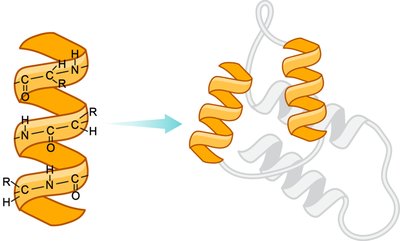
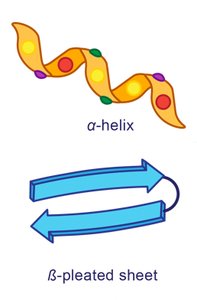
The secondary structure refers to local folding patterns within a polypeptide, stabilized by hydrogen bonds between backbone atoms. The two main types are the α-helix and β-pleated sheet.
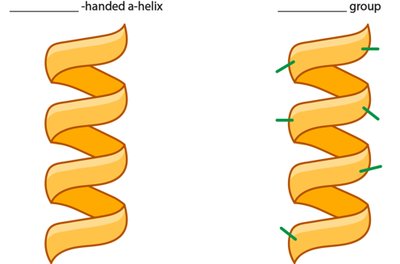
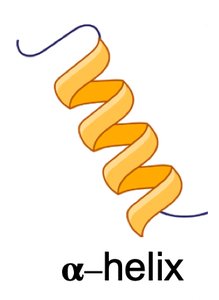
α-Helix: Right-handed spiral stabilized by hydrogen bonds every fourth residue.
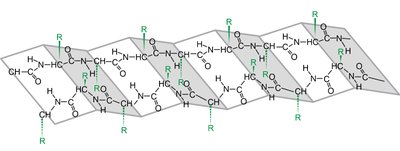
β-Pleated Sheet: Extended strands connected side-by-side by hydrogen bonds, with R groups alternating above and below the plane.
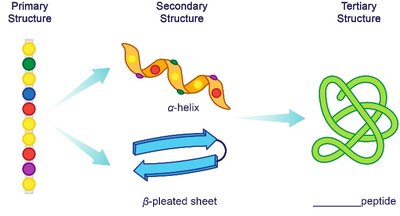
Tertiary Structure
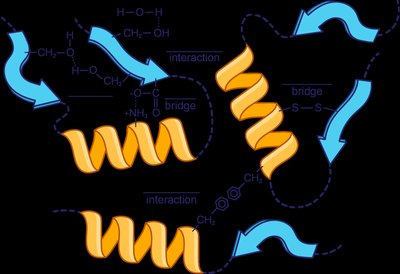
The tertiary structure is the overall three-dimensional shape of a single polypeptide chain, stabilized by interactions between R groups, including hydrophobic interactions, hydrogen bonds, ionic bonds (salt bridges), and disulfide bridges (covalent bonds between cysteine residues).
Hydrophobic interactions: Nonpolar side chains cluster away from water.
Hydrogen bonds: Between polar side chains.
Salt bridges: Ionic bonds between acidic and basic side chains.
Disulfide bridges: Covalent bonds between cysteine residues.
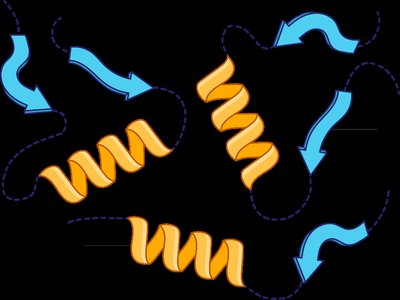
Quaternary Structure
The quaternary structure is the arrangement of multiple polypeptide subunits in a functional protein complex. Subunits are held together by the same types of interactions as in tertiary structure. Multimeric proteins can be dimers, trimers, tetramers, etc.
Subunit: An individual polypeptide chain within a protein complex.
Prosthetic group: A non-protein component essential for protein function.



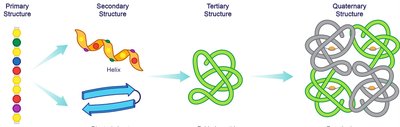
Summary Table: Levels of Protein Structure
Structure Level | Characteristics | Stabilized By |
|---|---|---|
Primary | Sequence of amino acids | Peptide (amide) bonds |
Secondary | Local arrangement (α-helix, β-sheet) | Hydrogen bonds between backbone atoms |
Tertiary | Overall 3D shape of a polypeptide | Hydrophobic interactions, hydrogen bonds, ionic bonds, disulfide bridges |
Quaternary | Association of multiple polypeptides | Same as tertiary structure interactions |



Protein Denaturation
Denaturation is the process by which a protein loses its native structure and, consequently, its biological function. This can occur through heat, changes in pH, heavy metal ions, or mechanical agitation. Denaturation disrupts secondary, tertiary, and quaternary structures but does not break peptide bonds in the primary structure.
Heat: Disrupts nonpolar interactions at temperatures above 50°C.
pH changes: Alter charges on side chains, disrupting ionic and hydrogen bonds.
Heavy metals: Bind to sulfur atoms, disrupting disulfide bridges.
Mechanical agitation: Physical stress unfolds proteins (e.g., whisking egg whites).
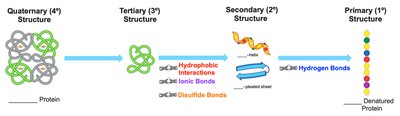
Key Point: The amino acid sequence (primary structure) remains intact during denaturation.
