 Back
BackEnzymes and Vitamins: Structure, Function, and Regulation
Study Guide - Smart Notes

Enzymes and Vitamins
19.1 Catalysis by Enzymes
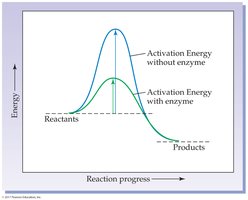
Enzymes are biological catalysts, primarily proteins, that accelerate biochemical reactions without being consumed. They lower the activation energy required for reactions, thus increasing the rate at which equilibrium is reached, but do not alter the equilibrium position itself.
Enzyme: A protein or molecule that catalyzes a biological reaction.
Active Site: The region on the enzyme where the substrate binds and the reaction occurs.
Substrate: The reactant in an enzyme-catalyzed reaction.
Specificity: Enzymes are highly specific, acting on particular substrates or reactions, often with stereochemical selectivity.
Example: Catalase is specific for decomposing hydrogen peroxide, while carboxypeptidase A can remove various C-terminal amino acids from proteins.
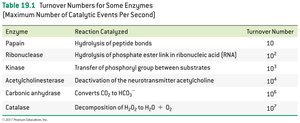
Enzyme activity is measured by the turnover number, which is the maximum number of substrate molecules converted per enzyme molecule per second. Turnover numbers can range from 10 to 10,000,000 per second.
Enzyme | Reaction Catalyzed | Turnover Number |
|---|---|---|
Papain | Hydrolysis of peptide bonds | 10 |
Ribonuclease | Hydrolysis of phosphate ester link in RNA | 102 |
Kinase | Transfer of phosphoryl group | 103 |
Acetylcholinesterase | Deactivation of acetylcholine | 104 |
Carbonic anhydrase | CO2 to HCO3- | 106 |
Catalase | Decomposition of H2O2 | 107 |

19.2 Enzyme Cofactors
Some enzymes require nonprotein components called cofactors for activity. Cofactors may be metal ions or organic molecules called coenzymes. These components provide chemical groups not present in amino acid side chains, enabling a wider range of reactions.
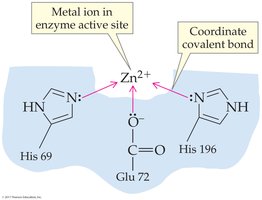
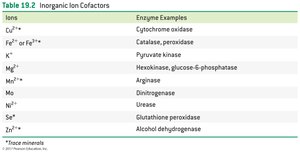
Metal ions (e.g., Zn2+, Fe2+) can form coordinate covalent bonds and act as Lewis acids.
Coenzymes are often derived from vitamins and act as carriers of specific atoms or functional groups.
Ions | Enzyme Examples |
|---|---|
Cu2+ | Cytochrome oxidase |
Fe2+ or Fe3+ | Catalase, peroxidase |
K+ | Pyruvate kinase |
Mg2+ | Hexokinase, glucose-6-phosphatase |
Mn2+ | Arginase |
Mo | Dinitrogenase |
Ni2+ | Urease |
Se | Glutathione peroxidase |
Zn2+ | Alcohol dehydrogenase |
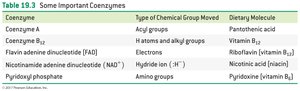
Coenzyme | Type of Chemical Group Moved | Dietary Molecule |
|---|---|---|
Coenzyme A | Acyl groups | Pantothenic acid |
Coenzyme B12 | H atoms and alkyl groups | Vitamin B12 |
FAD | Electrons | Riboflavin (vitamin B2) |
NAD+ | Hydride ion (H-) | Nicotinic acid (niacin) |
Pyridoxyl phosphate | Amino groups | Pyridoxine (vitamin B6) |
19.3 Enzyme Classification
Enzymes are classified into six main classes based on the type of reaction they catalyze. Their names typically end in -ase and often indicate both the substrate and the reaction type.
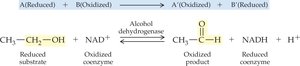
Oxidoreductases: Catalyze oxidation-reduction reactions (e.g., dehydrogenases, oxidases).
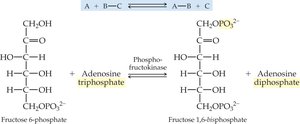
Transferases: Transfer functional groups between molecules (e.g., kinases, transaminases).
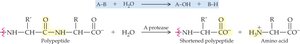
Hydrolases: Catalyze hydrolysis reactions (e.g., proteases, lipases).
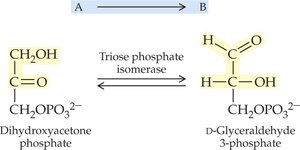
Isomerases: Catalyze isomerization (rearrangement) of molecules.
Lyases: Add or remove groups to form double bonds (e.g., decarboxylases, dehydratases).
Ligases: Join two molecules together, usually with ATP hydrolysis (e.g., synthetases, carboxylases).

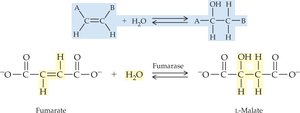
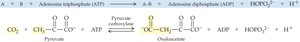
19.4 How Enzymes Work
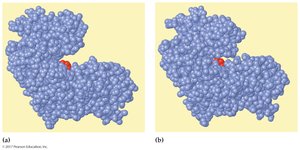
Enzyme specificity is determined by the structure of the active site. Two models describe enzyme-substrate interaction:
Lock-and-Key Model: The substrate fits exactly into the rigid active site.
Induced-Fit Model: The enzyme active site is flexible and molds around the substrate upon binding.
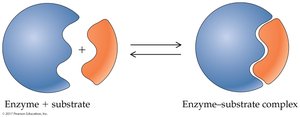
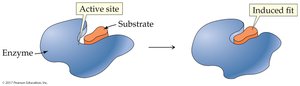
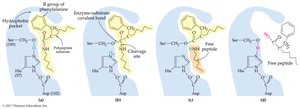
Enzymes catalyze reactions by:
Bringing substrates together (proximity effect)
Orienting substrates correctly (orientation effect)
Providing catalytic groups (catalytic effect)
Inducing strain in substrate bonds (energy effect)
19.5 Factors Affecting Enzyme Activity
Enzyme activity is influenced by substrate concentration, enzyme concentration, temperature, and pH.
Substrate Concentration: Rate increases with substrate concentration until the enzyme is saturated.
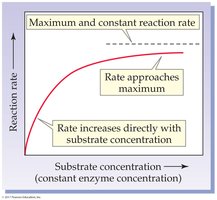
Enzyme Concentration: Rate increases linearly with enzyme concentration if substrate is in excess.
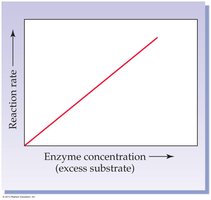
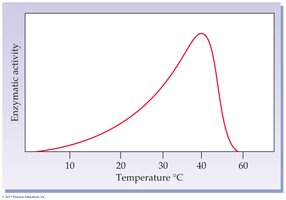
Temperature: Rate increases with temperature up to an optimum, then decreases due to denaturation.
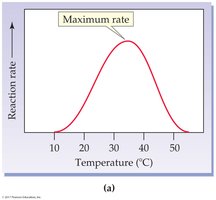
pH: Each enzyme has an optimum pH; extremes can denature the enzyme.
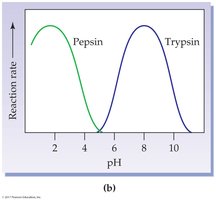
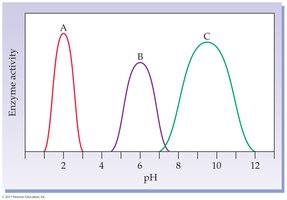
19.6 Enzyme Regulation: Inhibition
Enzyme activity can be regulated by inhibitors, which may be reversible or irreversible.
Reversible Uncompetitive Inhibition: Inhibitor binds only to the enzyme-substrate complex.
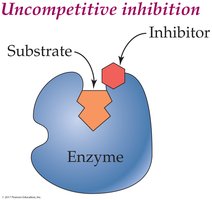
Reversible Competitive Inhibition: Inhibitor competes with substrate for the active site.
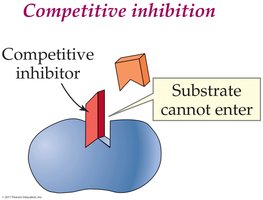
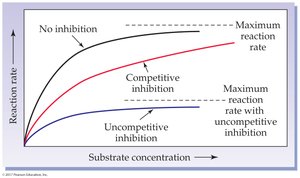
Irreversible Inhibition: Inhibitor forms covalent bonds with the enzyme, permanently inactivating it (e.g., heavy metals like mercury and lead).
19.7 Enzyme Regulation: Allosteric Control and Feedback Inhibition
Enzyme activity can also be regulated by allosteric and feedback mechanisms.
Allosteric Control: Binding of a regulator at a site other than the active site changes enzyme activity. Can be positive (activator) or negative (inhibitor).
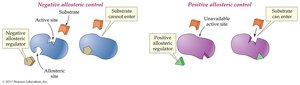
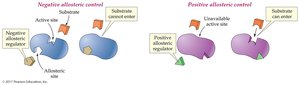
Feedback Control: The end product of a pathway inhibits an enzyme early in the pathway, preventing overproduction.
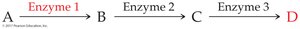
19.8 Enzyme Regulation: Covalent Modification and Genetic Control
Enzyme activity can be regulated by covalent modification or by controlling enzyme synthesis.
Zymogens: Inactive enzyme precursors activated by cleavage (e.g., trypsinogen to trypsin).
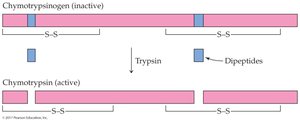
Phosphorylation/Dephosphorylation: Addition or removal of phosphate groups by kinases and phosphatases regulates enzyme activity.
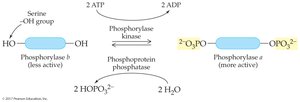
Genetic Control: Regulation of enzyme synthesis at the gene level (e.g., lactase production in infants vs. adults).
19.9 Vitamins, Antioxidants, and Minerals
Vitamins are essential organic molecules required in trace amounts, often as coenzyme precursors. Deficiencies or excesses can lead to health problems.
Water-Soluble Vitamins: Include vitamin C and B-complex vitamins; function mainly as coenzymes.
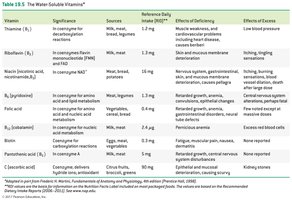
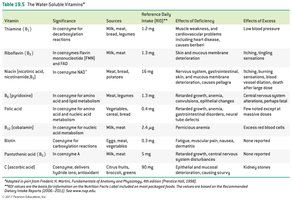
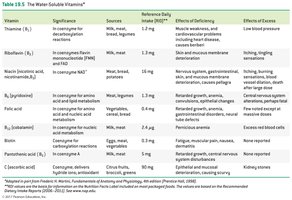
Fat-Soluble Vitamins: Vitamins A, D, E, and K; stored in body fat and not typically coenzymes.
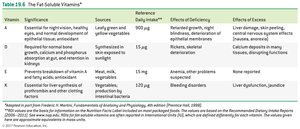
Antioxidants: Substances like vitamin C, vitamin E, and selenium that prevent oxidation and protect cells from free radicals.
Minerals: Inorganic elements required for enzyme function (as cofactors), structural roles, or as electrolytes. Macrominerals are needed in larger amounts, while microminerals (trace elements) are required in smaller quantities.
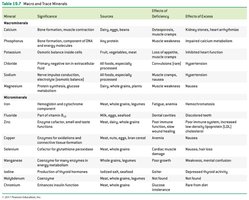
Example: Selenium is a cofactor for glutathione peroxidase, an enzyme that protects cells from oxidative damage.
