 Back
BackChapter 20: Biotechnology, DNA Cloning, and Genomics
Study Guide - Smart Notes

Biotechnology and Its Applications
Introduction to Biotechnology
Biotechnology is the engineering of genes, cells, and organisms for research and practical applications. It has revolutionized biological research and medicine by enabling manipulation and analysis of genetic material.
Recombinant DNA Technology: Mixing and matching DNA sequences from different organisms to produce large amounts of a gene product.
CRISPR-Cas Systems: Editing genomes at specific locations with high precision.
PCR (Polymerase Chain Reaction): Rapidly amplifying specific DNA sequences.
DNA Fingerprinting: Identifying individuals based on unique DNA features.
DNA Sequencing and Genomics: Determining the sequence of DNA and studying entire genomes for insights into evolution, disease, and traits.
Recombinant DNA Technology (DNA Cloning)
Overview of DNA Cloning
DNA cloning involves making multiple identical copies of a gene or DNA segment. Plasmids, restriction enzymes, and DNA ligase are essential tools in this process.
Plasmids: Small, circular DNA molecules used as vectors to carry foreign DNA into host cells.
Restriction Enzymes: Proteins that cut DNA at specific sequences, generating fragments with 'sticky ends' that facilitate joining.
DNA Ligase: Enzyme that seals the sugar-phosphate backbone, joining DNA fragments together.
Antibiotic Resistance Gene: Allows selection of bacteria that have successfully taken up the plasmid.
Steps of DNA Cloning
Cut the gene of interest and plasmid with the same restriction enzyme.
Ligate the gene into the plasmid using DNA ligase, forming a recombinant plasmid.
Transform bacteria by introducing the recombinant plasmid.
Select and screen for bacteria containing the recombinant plasmid (often using antibiotic resistance).
Harvest the gene product (protein or RNA) from the bacteria.

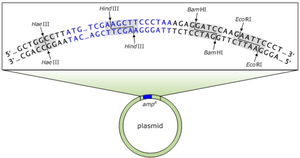
Cutting and Pasting DNA
Restriction enzymes generate complementary, single-stranded fragments called sticky ends, which can easily join together to form recombinant DNA.
Selection and Screening
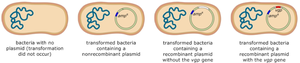
After transformation, bacterial cells may contain no plasmid, a nonrecombinant plasmid, a recombinant plasmid without the gene of interest, or a recombinant plasmid with the gene of interest. Selection is typically achieved using antibiotic resistance markers.
CRISPR-Cas Systems
The CRISPR Locus in Bacteria
CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) loci are segments of prokaryotic DNA containing short repetitions of base sequences. These loci function as a bacterial defense system against viruses.
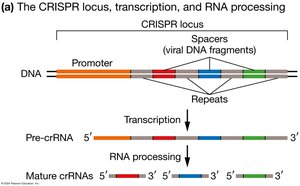
CRISPR-Cas Bacterial Defense System
The CRISPR-Cas system uses crRNA and tracrRNA to guide the Cas protein to viral DNA, which is then cleaved, providing immunity against future infections by the same virus.
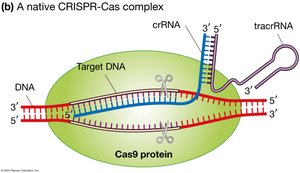
CRISPR-Cas9 for Gene Editing
In biotechnology, the CRISPR-Cas9 system is adapted for gene editing by replacing crRNA and tracrRNA with a single guide RNA (sgRNA) that targets any desired DNA sequence. Cas9 then introduces a double-strand break at the target site.
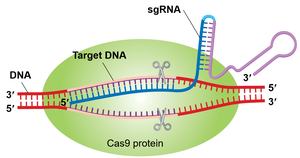
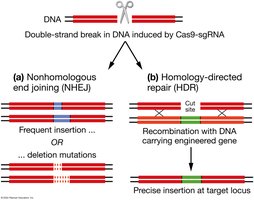
Genome Editing Repair Mechanisms
After Cas9-induced double-strand breaks, cells repair DNA via:
Nonhomologous End Joining (NHEJ): Error-prone, may introduce insertions or deletions.
Homology-Directed Repair (HDR): Uses a homologous template for precise repair, allowing insertion of engineered genes.
Applications of CRISPR
Development of crops with improved traits (e.g., browning-resistant mushrooms, drought-resistant soybeans).
Gene therapy for human diseases (e.g., sickle-cell disease, cystic fibrosis, neurodegenerative diseases).
Polymerase Chain Reaction (PCR)
Principle of PCR
PCR is a DNA synthesis reaction in which the products serve as templates for subsequent reactions, resulting in exponential amplification of a targeted DNA region.
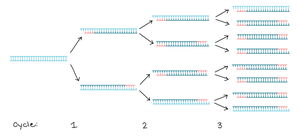
Components of a PCR Reaction
Template DNA: The DNA segment to be amplified.
Primers: Short DNA sequences that flank the target region and provide starting points for DNA synthesis.
DNA Polymerase: Enzyme that synthesizes new DNA strands (commonly Taq polymerase).
dNTPs: Deoxynucleotide triphosphates, the building blocks for new DNA strands.
Buffer: Maintains optimal conditions for the reaction.
Steps of PCR
Denaturation: Heating to separate double-stranded DNA (C).
Annealing: Cooling to allow primers to bind to target sequences (C).
Extension: DNA polymerase extends primers to synthesize new DNA (C).
DNA Fingerprinting
Principle of DNA Fingerprinting
DNA fingerprinting identifies individuals based on unique features of their genomes, most commonly using Short Tandem Repeats (STRs). The number of repeats at each STR locus varies between individuals, creating a unique DNA profile.
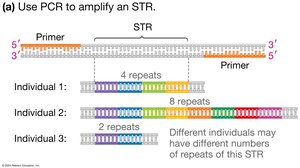
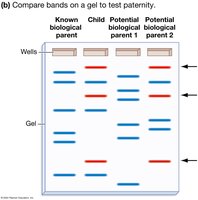
STR Analysis and Gel Electrophoresis
STR regions are amplified by PCR and separated by size using gel electrophoresis. Each band represents a DNA fragment of a specific length, allowing comparison between individuals for applications such as paternity testing and forensics.
DNA Sequencing (Sanger Sequencing)
Principle of Sanger Sequencing
Sanger sequencing uses dideoxynucleotides (ddNTPs), which lack a 3' hydroxyl group, causing DNA synthesis to terminate when incorporated. This generates DNA fragments of varying lengths that can be separated and analyzed to determine the DNA sequence.
ddNTPs: Chain-terminating nucleotides used to stop DNA synthesis at specific bases.
Visualization: DNA fragments are separated by size and detected, often using fluorescent labels.

Genome-Wide Association Studies (GWAS) and Genomics
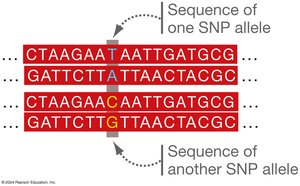
GWAS and SNPs
GWAS compare genomes to identify genetic markers, such as Single Nucleotide Polymorphisms (SNPs), associated with diseases or traits. SNPs are single base-pair variations between individuals.
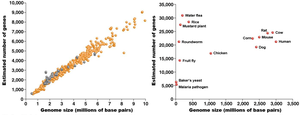
Insights from Sequencing Many Genomes
Genome sequencing has revealed that genome size does not always correlate with gene number. Comparative genomics helps identify evolutionary relationships and functional elements in genomes.
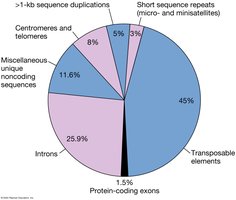
Composition of the Human Genome
The human genome is composed of various elements, including protein-coding exons, introns, transposable elements, and noncoding sequences. Protein-coding regions make up only a small fraction of the genome.
Additional info: These notes provide a comprehensive overview of key biotechnology concepts, including DNA cloning, CRISPR-Cas systems, PCR, DNA fingerprinting, Sanger sequencing, GWAS, and genome composition, as relevant to a college-level biology course.
