Back
BackCH. 19 p.1 - Gene Expression Regulation in Eukaryotes: Chromatin Remodeling and Transcription Initiation
Study Guide - Smart Notes

Gene Expression Regulation in Eukaryotes
Overview of Gene Regulation in Eukaryotes
Gene expression in eukaryotes is controlled at multiple levels, allowing cells to respond to environmental changes and to differentiate into specialized cell types. This regulation is more complex than in prokaryotes due to the presence of chromatin and compartmentalization within the nucleus.
Necessity of Regulation: Ensures proper cell function, adaptation to environmental changes, and cell specialization in multicellular organisms.
Levels of Control: Includes transcriptional, translational, and post-translational regulation, as well as unique eukaryotic mechanisms such as chromatin remodeling, RNA processing, and mRNA stability.
Cell Specialization: All cells contain the same DNA, but differential gene expression leads to diverse cell types and functions.

Levels of Gene Expression Control
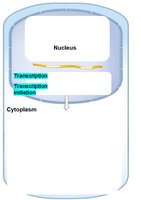
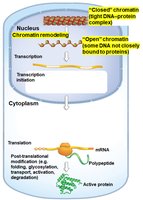
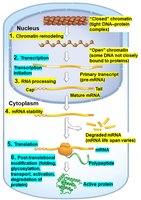
Gene expression can be regulated at several stages from DNA to active protein. The main levels of control are:
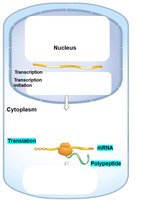
Transcriptional Control: Regulation of whether and how much a gene is transcribed into mRNA.
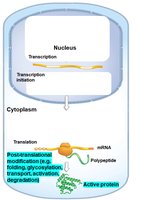
Translational Control: Regulation of whether and how much mRNA is translated into protein.
Post-Translational Control: Regulation of protein folding, modification, transport, activation, and degradation.
Unique Eukaryotic Levels of Control
Chromatin Remodeling: Changes in chromatin structure that affect gene accessibility.
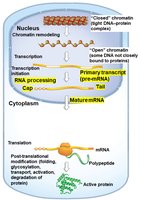
RNA Processing: Modifications such as capping, splicing, and polyadenylation of pre-mRNA.
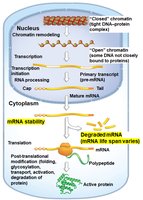
mRNA Stability: Regulation of mRNA degradation rates, affecting how long mRNAs are available for translation.
Chromatin Remodeling (19.2)
Chromatin Structure
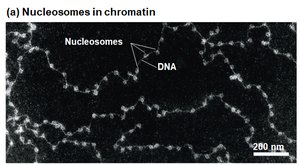
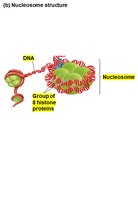
Chromatin is the complex of DNA and proteins (mainly histones) that packages eukaryotic DNA into the nucleus. The basic unit of chromatin is the nucleosome.
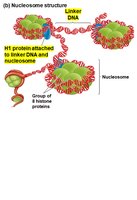
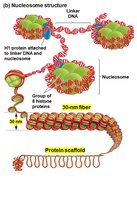
Nucleosome: Consists of DNA wrapped around a core of eight histone proteins.
Linker DNA: The stretch of DNA between nucleosomes, often associated with H1 histone protein.
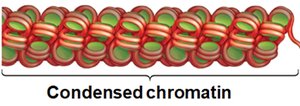
Condensed Chromatin (30 nm fiber): Nucleosomes are further packed into a thicker fiber, which can interact with a protein scaffold for higher-order structure, especially before cell division.
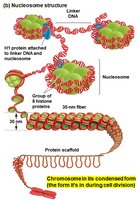
Chromosome Location and Chromatin Dynamics
During interphase, each chromosome occupies a distinct region in the nucleus. Chromatin can unfold to allow access to genes for transcription and refold when transcription is complete.
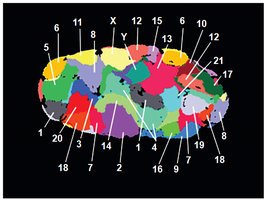
Chromatin Structure and Gene Activity
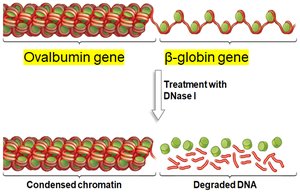
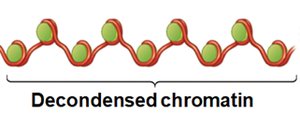
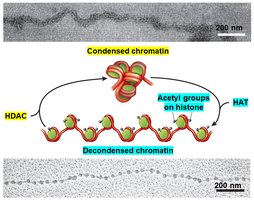
Active genes are found in decondensed chromatin, making their DNA accessible to transcription machinery. Inactive genes remain in condensed chromatin, which is resistant to DNase digestion.
DNase Sensitivity: DNase I can degrade DNA in decondensed regions but not in condensed chromatin.
Example: In chicken blood cells, the β-globin gene (active in blood cells) is DNase-sensitive, while the ovalbumin gene (inactive in blood cells) is DNase-resistant.
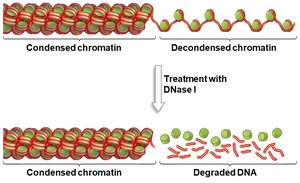
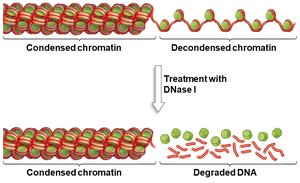
Mechanisms of Chromatin Remodeling
Chromatin structure can be altered by several mechanisms, making genes accessible or inaccessible for transcription.
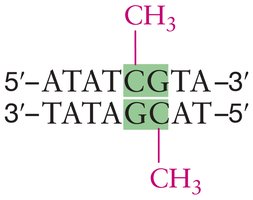
DNA Methylation: Addition of methyl groups to cytosine bases (usually in CpG dinucleotides) leads to chromatin condensation and gene silencing. Demethylation allows chromatin to decondense and genes to be transcribed.
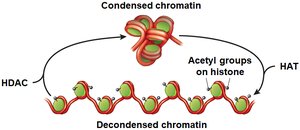
Histone Modification: The 'histone code hypothesis' proposes that combinations of histone modifications (e.g., acetylation, methylation) determine chromatin state. Acetylation by HATs (histone acetyltransferases) decondenses chromatin, while deacetylation by HDACs (histone deacetylases) condenses it.
Chromatin Remodeling Complexes: Protein complexes that reposition or remove nucleosomes, exposing stretches of DNA for transcription.
Initiating Transcription (19.3)
DNA Regulatory Sequences
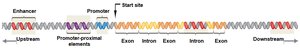
Transcription initiation in eukaryotes involves several types of DNA regulatory sequences and associated proteins:
Core Promoter: The binding site for RNA polymerase II, often containing a TATA box.
Promoter-Proximal Elements: Regulatory sequences located near and upstream of the core promoter; binding sites for activator and repressor proteins.
Enhancers: Distant regulatory sequences that increase transcription when bound by activator proteins. They can be located upstream, downstream, or within introns and function regardless of orientation.
Silencers: Binding sites for repressor proteins that decrease transcription (negative control).
Promoter-Proximal Elements and Gene Regulation
Promoter-proximal elements are crucial for coordinated gene regulation. For example, in yeast, genes involved in galactose metabolism are regulated by a common activator protein binding to the same promoter-proximal element, even though the genes are on different chromosomes.
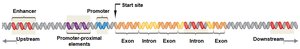
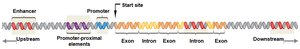
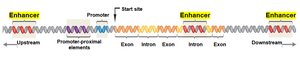
Enhancers and Gene Regulation
Most genes have multiple enhancers, which can be located far from the promoter and still function effectively. Enhancers can be found upstream, downstream, or within introns of the gene they regulate.

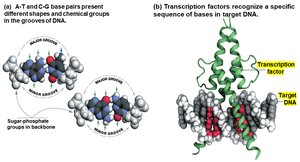
Regulatory Transcription Factors
Regulatory transcription factors include activators and repressors that bind specific DNA sequences to control gene expression. Different cell types express different sets of transcription factors, leading to cell-specific gene expression patterns. The expression of these factors is often regulated by external signals, such as those involved in development or environmental response.
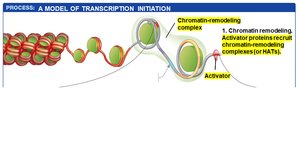
Model for Transcription Initiation
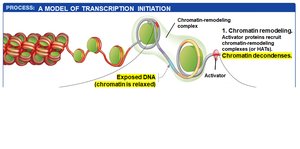
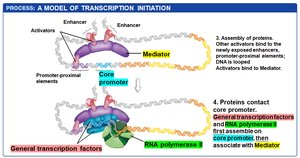
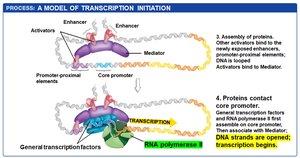
Transcription initiation in eukaryotes is a multi-step process involving the coordinated action of regulatory transcription factors, general transcription factors, mediator proteins, and chromatin remodeling complexes.
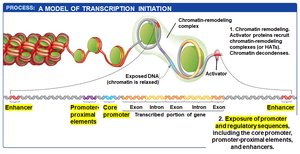
Activator proteins bind to enhancers and promoter-proximal elements, recruiting chromatin-remodeling complexes and HATs to open chromatin structure.
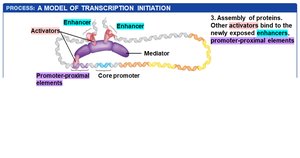
Exposed DNA allows the assembly of additional transcription factors and mediator proteins, which bridge regulatory proteins and RNA polymerase II.
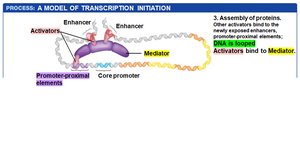
DNA looping brings distant regulatory elements into proximity with the core promoter.
General transcription factors and RNA polymerase II assemble at the core promoter, and transcription begins.
Summary Table: Elements of Transcriptional Regulation
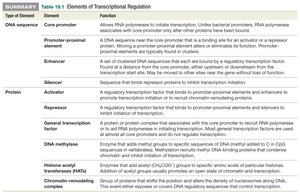
Type of Element | Element | Function |
|---|---|---|
DNA sequence | Core promoter | Allows RNA polymerase to initiate transcription. |
DNA sequence | Promoter-proximal element | Binding site for activator or repressor proteins near the core promoter. |
DNA sequence | Enhancer | Binding site for activator proteins, can be distant from the gene. |
DNA sequence | Silencer | Binding site for repressor proteins, decreases transcription. |
Protein | Activator | Transcription factor that increases transcription by binding regulatory sequences. |
Protein | Repressor | Transcription factor that decreases transcription by binding regulatory sequences. |
Protein | General transcription factor | Associates with the core promoter and RNA polymerase II to initiate transcription. |
Protein | Mediator | Bridges regulatory proteins and RNA polymerase II, integrating regulatory signals. |
Modification | DNA methylation | Condenses chromatin and silences gene expression. |
Modification | Histone acetyl transferase (HAT) | Adds acetyl groups to histones, decondensing chromatin and activating genes. |
Modification | Chromatin remodeling complex | Repositions or removes nucleosomes to expose DNA for transcription. |