 Back
BackGene Expression: Transcription, RNA Processing, and Translation
Study Guide - Smart Notes

Gene Expression: The Big Picture
Gene expression is the process by which genetic information encoded in DNA is used to direct the synthesis of functional gene products, primarily proteins. This process involves two main stages: transcription (the synthesis of RNA from DNA) and translation (the synthesis of proteins from mRNA). In eukaryotes, gene expression also includes RNA processing steps such as splicing, capping, and polyadenylation.
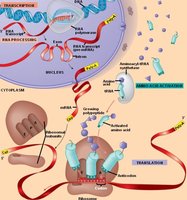
Transcription
Overview of Transcription
Transcription is the process by which RNA is synthesized from a DNA template. This process is catalyzed by the enzyme RNA polymerase. Unlike DNA replication, transcription produces a single-stranded RNA molecule and does not require a primer.
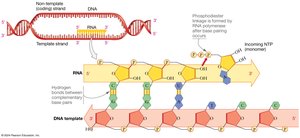
Template strand: The DNA strand that is read by RNA polymerase to synthesize RNA.
Non-template (coding) strand: The DNA strand with the same sequence as the RNA (except T is replaced by U).
Direction: RNA is synthesized in the 5' to 3' direction.
Transcription Initiation: Promoters and Sigma Factor
Transcription begins at specific DNA sequences called promoters. In bacteria, promoters contain conserved regions known as the -10 box and -35 box. The sigma factor is a protein that helps RNA polymerase recognize and bind to the promoter.
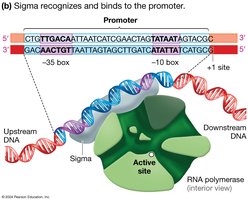
-10 and -35 boxes: Consensus sequences in bacterial promoters that are recognized by sigma factor.
Sigma factor: Directs RNA polymerase to the promoter and initiates transcription.
Steps of Transcription Initiation
Transcription initiation involves several steps:
RNA polymerase holoenzyme binds to the promoter with the help of sigma factor.
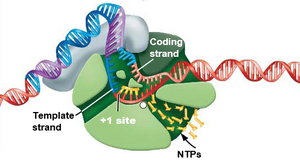
The DNA double helix is unwound near the +1 site (the transcription start site).
RNA synthesis begins as ribonucleotides are added complementary to the DNA template strand.
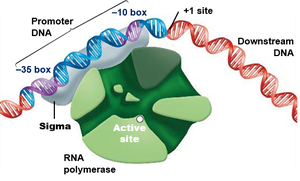
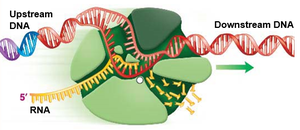
Transcription Elongation
During elongation, RNA polymerase moves along the DNA template, synthesizing RNA in the 5' to 3' direction. The DNA helix reforms behind the polymerase, and the newly synthesized RNA strand detaches from the DNA template.

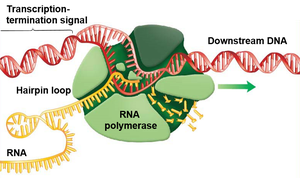
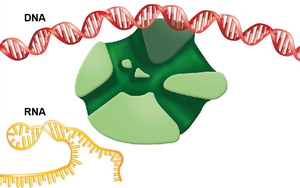
Transcription Termination
Termination occurs when RNA polymerase encounters a termination signal in the DNA. In bacteria, this often involves the formation of a hairpin loop in the RNA, which causes the polymerase to dissociate from the DNA and release the RNA transcript.
RNA Processing in Eukaryotes
Splicing
In eukaryotes, the initial RNA transcript (pre-mRNA) contains both exons (coding regions) and introns (non-coding regions). Splicing removes introns and joins exons to produce a mature mRNA.
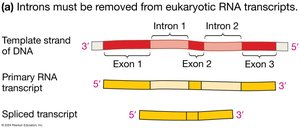
Carried out by: The spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs).
Purpose: To produce a continuous coding sequence for translation.
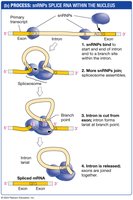
snRNPs and the Spliceosome
snRNPs (small nuclear ribonucleoproteins) recognize splice sites and assemble into the spliceosome. The spliceosome catalyzes the removal of introns and the ligation of exons. The RNA component of snRNPs acts as a ribozyme, catalyzing the splicing reaction.
Capping and Polyadenylation
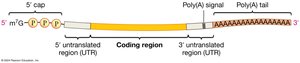
After splicing, eukaryotic mRNAs undergo two additional modifications:
5' cap: Addition of a modified guanine nucleotide to the 5' end of the mRNA.
3' poly-A tail: Addition of a long stretch of adenine nucleotides to the 3' end.
These modifications protect mRNA from degradation, aid in export from the nucleus, and facilitate translation initiation.
Translation
Ribosomes and Polyribosomes
Translation is the process by which ribosomes synthesize proteins using the information in mRNA. Multiple ribosomes can simultaneously translate a single mRNA, forming a structure called a polyribosome or polysome.
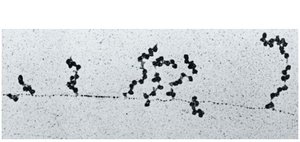
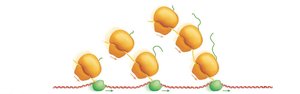
Coupling of Transcription and Translation in Bacteria
In bacteria, transcription and translation are coupled, meaning translation can begin on an mRNA while it is still being synthesized.

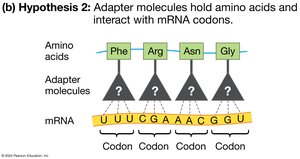
Codons and Adapter Molecules
mRNA codons specify amino acids, but mRNA cannot directly interact with amino acids. tRNA molecules act as adapters, matching codons with their corresponding amino acids during translation.
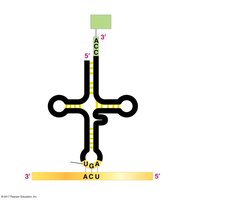
tRNA Structure and Function
tRNAs have a characteristic cloverleaf structure with two important regions:
Amino acid attachment site: The 3' end of the tRNA, where the amino acid is covalently attached.
Anticodon: A three-nucleotide sequence that base-pairs with the complementary mRNA codon.
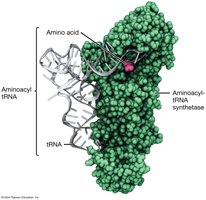
Aminoacyl-tRNA Synthetases (ARSs)
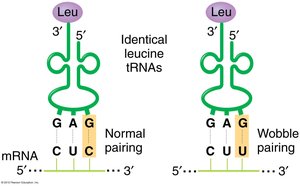
ARSs are enzymes that attach the correct amino acid to its corresponding tRNA. There are fewer ARSs and tRNAs than codons due to the wobble hypothesis, which allows some tRNAs to recognize multiple codons.
Ribosome Structure and Function
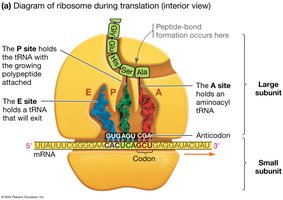
Ribosomes are composed of protein and ribosomal RNA (rRNA) and have two subunits (large and small). They contain three sites for tRNA binding:
A site (Aminoacyl): Holds the tRNA carrying the next amino acid to be added.
P site (Peptidyl): Holds the tRNA with the growing polypeptide chain.
E site (Exit): Holds the tRNA that is about to exit the ribosome.
Translation: Initiation, Elongation, and Termination
Initiation
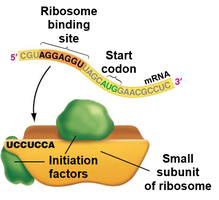
Translation initiation involves the assembly of the ribosome on the mRNA at the start codon, with the initiator tRNA in the P site.
Elongation
During elongation, amino acids are added one by one to the growing polypeptide chain as the ribosome moves along the mRNA.
Key steps:
tRNA entry into the A site
Peptide bond formation
Translocation of the ribosome
Termination
Termination occurs when a stop codon is reached. Release factors bind to the stop codon, causing the ribosome to release the completed polypeptide and dissociate from the mRNA.
Comparison of Gene Expression in Prokaryotes and Eukaryotes
Gene expression differs between prokaryotes and eukaryotes in several key ways:
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Transcription and Translation | Coupled (occur simultaneously) | Separated by nuclear envelope |
RNA Processing | Minimal (no splicing, capping, or polyadenylation) | Extensive (splicing, capping, polyadenylation) |
Location | Cytoplasm | Transcription in nucleus, translation in cytoplasm |
Summary
Gene expression involves transcription, RNA processing (in eukaryotes), and translation.
Transcription is initiated at promoters, involves RNA polymerase, and ends at termination signals.
RNA processing includes splicing, capping, and polyadenylation.
Translation is carried out by ribosomes, with tRNAs serving as adapters between codons and amino acids.
There are important differences in gene expression between prokaryotes and eukaryotes.
