 Back
BackMutation, Gene Expression, and the Central Dogma: DNA to Protein
Study Guide - Smart Notes

Mutation: Definition and Types
What is Mutation?
Mutation refers to any change in the DNA sequence or genetic information. Mutations can range from single base changes (point mutations) to large-scale chromosomal alterations. These changes can have a variety of effects, from no observable impact to significant changes in phenotype or fitness.
Germline mutations occur in reproductive cells and are heritable.
Somatic mutations occur in non-reproductive cells and are passed to daughter cells during mitosis, but not to offspring.
Mutations arise during DNA replication, due to DNA damage, or from errors in chromosome segregation during cell division.
Key Point: Mutation is a constant and inevitable process in all living organisms.
Sources of Mutation
Errors in DNA replication: DNA polymerase may insert incorrect bases, skip bases (deletion), or add extra bases (insertion).
DNA damage: Physical (e.g., UV light causing thymine dimers) or chemical agents can alter DNA structure, leading to mutations.
Chromosome segregation errors: Mistakes during mitosis or meiosis can result in aneuploidy (extra or missing chromosomes).
Major Types of Mutation
Missense mutation: A single base change that alters the amino acid sequence of a protein.
Nonsense mutation: A single base change that converts a codon for an amino acid into a stop codon, truncating the protein.
Silent mutation: A single base change that does not alter the amino acid sequence due to redundancy in the genetic code.
Frameshift mutation: Insertions or deletions that shift the reading frame, altering all downstream amino acids.
Point mutations affect a single nucleotide position in the genome.
Gene Expression: From DNA to Protein
Chromatin Structure and Gene Regulation
In eukaryotes, DNA is packaged with histone proteins into chromatin. The interaction between negatively charged DNA and positively charged histones forms nucleosomes, which can further condense into higher-order structures. Chromatin must be decondensed for transcription to occur, often through chemical modification of histones that neutralizes their positive charges and loosens DNA-histone interactions.
mRNA Processing in Eukaryotes
Before mRNA can be translated, it undergoes several processing steps:
5′ capping: Addition of a modified guanine nucleotide to the 5′ end, protecting mRNA and aiding ribosome binding.
Splicing: Removal of non-coding introns and joining of exons by the spliceosome.
Polyadenylation: Addition of a poly-A tail to the 3′ end, enhancing stability and translation efficiency.
Regulation of Protein Activity
Phosphorylation and Dephosphorylation
Protein activity is often regulated by the addition (phosphorylation) or removal (dephosphorylation) of phosphate groups. This modification can activate or deactivate proteins, including enzymes and receptors. Phosphorylation introduces negative charges, causing conformational changes that affect protein function.
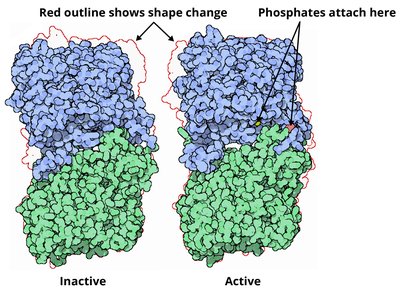
Example: Glycogen phosphorylase is activated by phosphorylation, enabling it to release glucose from glycogen stores.
Receptor Kinases and Cell Signaling
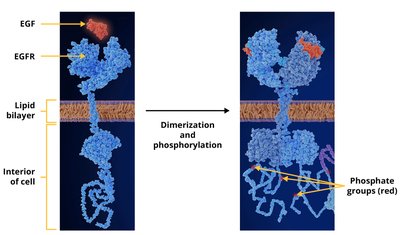
Receptors such as EGFR (Epidermal Growth Factor Receptor) are activated by ligand binding, dimerization, and autophosphorylation. This triggers downstream signaling pathways that regulate cell division and gene expression. Mutations that cause constitutive activation of EGFR are associated with certain cancers, and drugs targeting EGFR can be effective treatments.
Transcription: DNA to RNA
Mechanism of Transcription
Transcription is the process by which RNA polymerase synthesizes RNA from a DNA template. The enzyme binds to the promoter region, unwinds the DNA, and catalyzes the formation of phosphodiester bonds between ribonucleotides. Transcription proceeds in the 5′ to 3′ direction, and the template strand is read in the 3′ to 5′ direction.
Initiation: RNA polymerase and initiation factors bind to the promoter and begin RNA synthesis.
Elongation: RNA polymerase moves along the DNA, extending the RNA chain.
Termination: Specific sequences signal the end of transcription, releasing the RNA transcript.
Translation: RNA to Protein
tRNA and the Genetic Code
Translation converts the nucleotide sequence of mRNA into a sequence of amino acids in a protein. tRNAs serve as adaptors, each carrying a specific amino acid and an anticodon that pairs with the corresponding codon in mRNA. The genetic code is read in triplets (codons), with 64 possible codons specifying 20 amino acids and punctuation signals (start and stop codons).
Ribosome Structure and Function
Ribosomes are composed of large and small subunits, rRNA, and proteins. They have three binding sites for tRNA (A, P, and E sites) and a channel for the emerging polypeptide. Translation proceeds through initiation (assembly of the ribosome on the mRNA), elongation (addition of amino acids), and termination (release of the completed protein).
DNA Replication and Repair
Semiconservative Replication
DNA replication is semiconservative: each new DNA molecule consists of one parental and one newly synthesized strand. DNA polymerases synthesize DNA in the 5′ to 3′ direction, adding nucleotides to the 3′ end. Leading strand synthesis is continuous, while lagging strand synthesis is discontinuous, producing Okazaki fragments.
Replication of Chromosome Ends
Linear chromosomes have ends called telomeres. Replicating these ends poses a challenge, as the lagging strand cannot be fully replicated, leading to gradual shortening. Specialized enzymes like telomerase help maintain telomere length in certain cells.
DNA Repair Mechanisms
Cells possess multiple mechanisms to repair DNA damage and replication errors, including proofreading by DNA polymerases, mismatch repair, and excision repair pathways. High fidelity in DNA replication is essential for maintaining genetic integrity and preventing diseases such as cancer.
Summary Table: Types of Mutations
Type of Mutation | Description | Effect on Protein |
|---|---|---|
Missense | Single base change alters amino acid | May alter protein function |
Nonsense | Single base change creates stop codon | Truncated, usually nonfunctional protein |
Silent | Single base change, no amino acid change | No effect on protein sequence |
Frameshift | Insertion/deletion shifts reading frame | Alters downstream amino acids, usually nonfunctional protein |
Key Vocabulary
Mutation: Any change in the genetic information in DNA.
Missense mutation: Alters the amino acid sequence of a protein.
Nonsense mutation: Introduces a premature stop codon.
Silent mutation: Changes a codon without altering the amino acid.
Frameshift mutation: Alters the reading frame of the gene.
Phosphorylation: Addition of a phosphate group to a protein, regulating its activity.
Kinase: Enzyme that adds phosphate groups to proteins.
tRNA: Transfer RNA, adaptor molecule in translation.
Ribozyme: RNA molecule with catalytic activity.
Codon: Three-base sequence in mRNA specifying an amino acid.
Start/Stop codon: Signals for initiation and termination of translation.
