 Back
BackTranscription, RNA Processing, and Translation: Mechanisms and Regulation
Study Guide - Smart Notes

Transcription, RNA Processing, and Translation
Overview of Transcription
Transcription is the process by which RNA polymerases synthesize an RNA copy of the genetic instructions stored in DNA. This process is fundamental to gene expression and involves several key steps and molecular players.
RNA Polymerases: Enzymes that use ribonucleoside triphosphates (NTPs) to build an RNA strand complementary to the DNA template strand.
Template and Coding Strands: Only one DNA strand serves as the template for RNA synthesis. The non-template (coding) strand has the same sequence as the RNA (except that RNA contains uracil (U) instead of thymine (T)).
Directionality: RNA polymerases synthesize RNA in the 5' to 3' direction, similar to DNA polymerases, but do not require a primer to begin synthesis.
Polymerase Diversity: Bacteria have a single RNA polymerase, while eukaryotes have at least three distinct types.
Initiation of Transcription in Bacteria
Transcription initiation is tightly regulated and requires specific protein-DNA interactions.
Sigma Factor: In bacteria, the RNA polymerase core enzyme requires a sigma protein to form a holoenzyme, which can recognize promoter sequences where transcription begins.
Promoters: Bacterial promoters are typically 40–50 base pairs long and contain conserved sequences such as the -10 (TATAAT, "TATA box") and -35 (TTGACA) elements upstream of the transcription start site.
Promoter Orientation: Sigma binds in a specific orientation, determining which DNA strand is used as the template and the direction of transcription.
Events Inside the Holoenzyme
Transcription Bubble: RNA polymerase opens the DNA double helix, creating a transcription bubble. The template strand is threaded through the active site, and NTPs are added according to base-pairing rules.
Polymerization: Complementary NTPs pair with the DNA template, and RNA synthesis begins.
Elongation and Termination in Bacteria
During elongation, RNA polymerase moves along the DNA, synthesizing RNA by adding nucleotides to the 3' end. Termination occurs when a specific sequence in the DNA is transcribed, signaling the end of transcription.
Elongation: RNA polymerase reads the DNA template and adds nucleotides to the 3' end of the growing RNA strand.
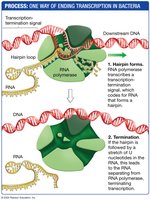
Termination Signal: In bacteria, termination often involves the formation of a hairpin structure in the RNA, which causes the polymerase to dissociate from the DNA and release the RNA transcript.
Transcription in Eukaryotes
Eukaryotic transcription is more complex than in bacteria, involving multiple RNA polymerases and additional regulatory elements.
Multiple RNA Polymerases: Eukaryotes have at least three types of RNA polymerases, each transcribing different classes of genes.
Promoters: Eukaryotic promoters are larger and more diverse, often containing a TATA box.
General Transcription Factors: These proteins, rather than sigma factors, recognize promoters and help recruit RNA polymerase.
Termination: Involves transcription of a poly(A) signal, after which the RNA is cleaved downstream of this site.
Compartmentalization: Transcription occurs in the nucleus, while translation occurs in the cytoplasm.
RNA Processing in Eukaryotes
Primary Transcripts and RNA Processing
In eukaryotes, the initial RNA product (primary transcript or pre-mRNA) must be processed before it can be translated into protein.
Primary Transcript: The direct product of transcription, containing both coding (exons) and noncoding (introns) sequences.
RNA Processing: Includes splicing, capping, and addition of a poly(A) tail.
Discovery of Split Genes
Experiments in 1977 revealed that eukaryotic genes are often interrupted by noncoding sequences (introns) that are not present in mature mRNA.
Introns: Noncoding regions removed during RNA processing.
Exons: Coding regions retained in the mature mRNA.
Experimental Evidence: Hybridization of viral DNA and mRNA revealed loops corresponding to introns.
RNA Splicing
Splicing removes introns from the primary transcript and joins exons together. This process is catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs).
Spliceosome: Made of snRNAs and proteins, responsible for recognizing splice sites and catalyzing intron removal.
Steps of Splicing:
snRNPs bind to exon-intron boundaries and a conserved A nucleotide near the end of the intron.
Additional snRNPs assemble to form the spliceosome.
The intron forms a lariat structure (looped out with A as the branch point).
The lariat is excised, and exons are ligated; the intron is degraded.
Alternative Splicing: Allows a single gene to produce multiple mRNA and protein variants.
Adding Caps and Tails to Transcripts
Pre-mRNAs are further processed by the addition of a 5' cap and a 3' poly(A) tail, which are essential for mRNA stability and translation.
5' Cap: A modified guanine nucleotide added to the 5' end, facilitating ribosome binding and protecting the RNA from degradation.
Poly(A) Tail: A stretch of 100–250 adenine nucleotides added to the 3' end after cleavage, necessary for translation and stability.
Mature mRNA: After splicing, capping, and tail addition, the mature mRNA contains untranslated regions (UTRs) at both ends, which play roles in translation regulation and mRNA stability.
Summary Table: Key Differences in Transcription and RNA Processing
Feature | Bacteria | Eukaryotes |
|---|---|---|
RNA Polymerases | One | At least three |
Promoter Elements | -10 and -35 boxes | TATA box and others |
Initiation Factors | Sigma protein | General transcription factors |
Termination Signal | Hairpin structure in RNA | Poly(A) signal and cleavage |
RNA Processing | Minimal | Splicing, capping, polyadenylation |
Compartmentalization | Cytoplasm | Nucleus (transcription), cytoplasm (translation) |
