 Back
BackAmino Acids, Peptides, and Proteins: Structure and Properties
Study Guide - Smart Notes

Amino Acids: Structure and Classification
Principle of Protein Construction
Proteins in all living organisms are constructed from a common set of 20 amino acids. Each amino acid possesses a unique side chain (R group) that imparts distinctive chemical properties, forming the 'alphabet' of protein structure.
Common Structural Features of Amino Acids
Amino acids share a general structure centered around the α carbon, which is a chiral center (except in glycine). The α carbon is tetrahedral and bonded to four substituents:
A carboxyl group (–COO−)
An amino group (–NH3+)
A hydrogen atom
An R group (side chain unique to each amino acid)
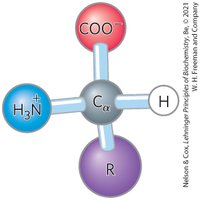
Stereochemistry of Amino Acids
All amino acids (except glycine) exist as two possible stereoisomers (enantiomers): L and D forms. Proteins are composed exclusively of L-amino acids, which are optically active. The D, L system specifies absolute configuration.
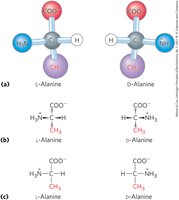
Classification of Amino Acids by R Group
Amino acids are classified into five main groups based on their R group properties:
Nonpolar, aliphatic (hydrophobic)
Aromatic (can absorb UV light)
Polar, uncharged (can form hydrogen bonds)
Positively charged (basic)
Negatively charged (acidic)
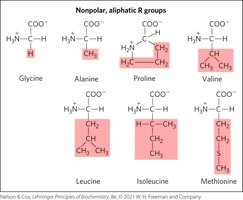
Nonpolar, Aliphatic R Groups
These amino acids are hydrophobic and stabilize protein structure via the hydrophobic effect.
Aromatic R Groups
Aromatic amino acids absorb UV light at 270–280 nm and contribute to hydrophobic interactions.
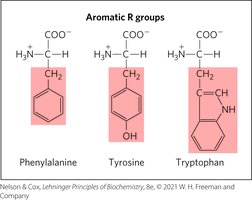
Polar, Uncharged R Groups
Polar amino acids can form hydrogen bonds; cysteine can form disulfide bonds, important for protein stability.
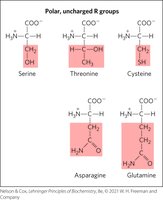
Positively Charged R Groups
These amino acids have significant positive charge at physiological pH (7.0).
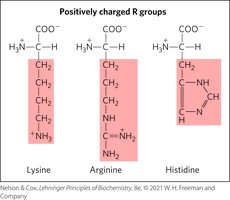
Negatively Charged R Groups
These amino acids have a net negative charge at pH 7.0.
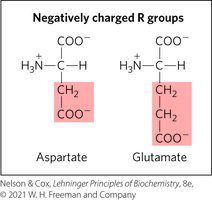
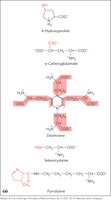
Uncommon Amino Acids and Modifications
Some amino acids are modified after or during protein synthesis, or transiently to alter protein function. Examples include 4-hydroxyproline (collagen), pyrrolysine (methane biosynthesis), and phosphorylation. Free metabolites such as ornithine are intermediates in biosynthetic pathways.
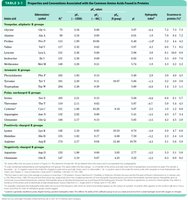
Acid-Base Properties of Amino Acids
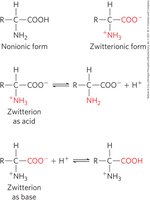
Amino Acids as Acids and Bases
Amino acids contain ionizable groups (amino, carboxyl, and some R groups) that act as weak acids and bases. At neutral pH, amino acids exist as zwitterions, carrying both positive and negative charges.
Titration of Amino Acids
The titration of amino acids reveals transitions between cationic, zwitterionic, and anionic forms. The carboxyl group has an acidic pKa (pK1), and the amino group has a basic pKa (pK2). The isoelectric point (pI) is the pH at which the net charge is zero.
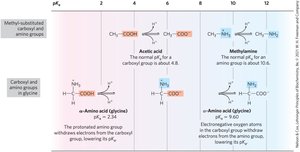
Effect of Chemical Environment on pKa
The α-carboxyl group in amino acids is more acidic than in simple carboxylic acids, and the α-amino group is less basic than in simple amines due to the influence of neighboring groups.
Information from Titration Curves
Titration curves provide quantitative measures of the pKa of each ionizing group, regions of buffering power, and the relationship between net charge and pH. The isoelectric point (pI) is a key property.
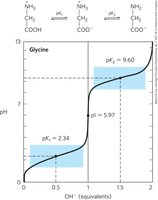
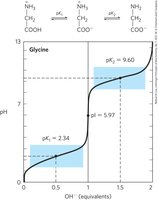
Amino Acids as Buffers
Amino acids act as buffers, preventing changes in pH near their pKa values. Glycine, for example, has two buffer regions centered around its α-carboxyl (pK1 = 2.34) and α-amino (pK2 = 9.6) groups.
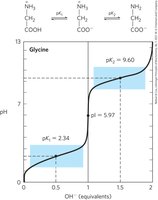
Isoelectric Point (pI)
For amino acids without ionizable side chains, the isoelectric point (pI) is the pH at which the net charge is zero. At pH > pI, the amino acid has a net negative charge; at pH < pI, it has a net positive charge.
Acid-Base Properties of Amino Acids with Ionizable Side Chains
Ionizable side chains have their own pKa values, act as buffers, influence the pI, and can be titrated, resulting in titration curves with three ionization steps.
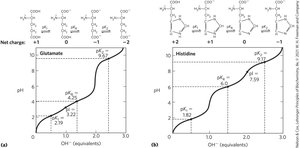
Peptides and Proteins: Structure and Nomenclature
Formation of Peptide Bonds
Amino acids are joined in linear sequences by peptide bonds, which are covalent amide linkages formed through condensation reactions and broken by hydrolysis. The sequence of amino acids forms the primary structure of proteins.
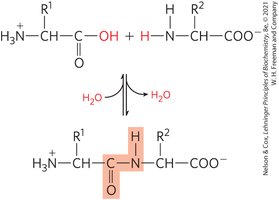
Types of Peptides
Dipeptide: 2 amino acids, 1 peptide bond
Tripeptide: 3 amino acids, 2 peptide bonds
Oligopeptide: a few amino acids
Polypeptide: many amino acids, molecular weight < 10 kDa
Protein: thousands of amino acids, molecular weight > 10 kDa
Peptide Terminals and Naming
Peptides are numbered and named starting from the amino-terminal residue (N-terminal) to the carboxyl-terminal residue (C-terminal).
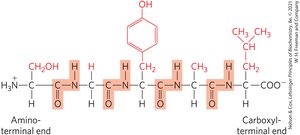
Peptides can be named using:
Full amino acid names (e.g., serylglycyltyrosylalanylleucine)
Three-letter code abbreviations (e.g., Ser–Gly–Tyr–Ala–Leu)
One-letter code abbreviation (e.g., SGYAL)
Ionization Behavior of Peptides
Peptides contain ionizable groups: one free α-amino group, one free α-carboxyl group, and some ionizable R groups.
Biologically Active Peptides and Protein Subunits
Peptides and polypeptides vary greatly in length and composition. Multisubunit proteins consist of two or more polypeptides associated noncovalently. Oligomeric proteins have at least two identical subunits called protomers.
Amino Acid Composition and Estimation
The amino acid composition of proteins is highly variable. The number of residues in a protein can be estimated by:
The average molecular weight of an amino acid is ~128, but a molecule of water (18) is removed during peptide bond formation, so the effective average is 110.
Conjugated Proteins
Some proteins contain permanently associated chemical groups (prosthetic groups) such as lipids (lipoproteins), sugars (glycoproteins), or metals (metalloproteins).
Protein Structure: Levels and Function
Levels of Protein Structure
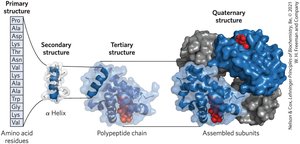
Proteins exhibit four levels of structural organization:
Primary structure: Covalent bonds linking amino acid residues in a polypeptide chain
Secondary structure: Recurring structural patterns (e.g., α helix, β sheet)
Tertiary structure: Three-dimensional folding of a single polypeptide
Quaternary structure: Association of two or more polypeptide subunits
Function Depends on Amino Acid Sequence
The amino acid sequence determines the three-dimensional structure, which in turn determines protein function. Most human proteins are polymorphic, meaning they have sequence variants. Edman degradation is a classic method for sequencing amino acids in proteins.
Additional info: These notes expand on the original content by providing definitions, examples, and context for each topic, ensuring completeness and academic quality for general chemistry students studying biochemistry.
