 Back
BackBacterial Cell Structure and the Cell Envelope
Study Guide - Smart Notes

Bacterial Cell Structure
Overview of Bacterial Cell Components
Bacterial cells possess a variety of structures that contribute to their survival, adaptation, and pathogenicity. Understanding these components is essential for microbiology students as it forms the foundation for topics such as microbial physiology, genetics, and pathogenesis.
Plasma membrane: Selectively permeable barrier, site of metabolic processes, and environmental sensing.
Gas vacuole: Provides buoyancy in aquatic environments.
Ribosomes: Sites of protein synthesis.
Inclusions: Storage of nutrients and sites of specific chemical reactions.
Nucleoid: Localization of genetic material (DNA).
Periplasmic space: Contains enzymes and proteins for nutrient processing (prominent in Gram-negative bacteria).
Cell wall: Maintains shape and protects from osmotic stress.
Capsules and slime layers: Aid in adherence and resistance to phagocytosis.
Fimbriae and pili: Attachment, gene transfer, and motility.
Flagella: Motility structures.
Endospore: Survival under harsh conditions.
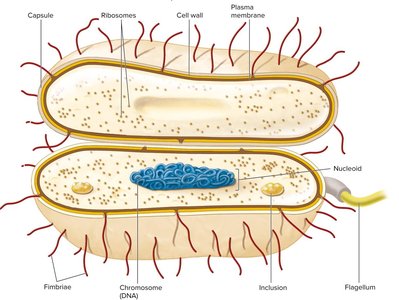
The Bacterial Cell Envelope
Layers of the Cell Envelope
The cell envelope is a complex, multi-layered structure that protects the cell and mediates interactions with the environment. It typically consists of:
Capsule/Slime layer (Glycocalyx): Outermost layer, composed of polysaccharides, aids in attachment and protection.
Cell wall: Provides rigidity and shape, composed mainly of peptidoglycan.
Plasma membrane: Phospholipid bilayer with embedded proteins, essential for transport and metabolism.
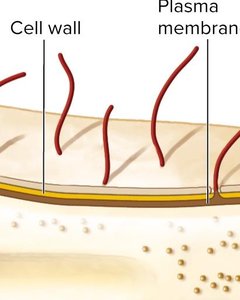
Capsules and Slime Layers
Capsules: Well-organized, not easily removed, usually polysaccharide-based. Provide resistance to phagocytosis, desiccation, and exclusion of viruses/detergents. Example: Bacillus anthracis (proteinaceous capsule), Streptococcus pneumoniae (capsule confers resistance to phagocytosis).
Slime layers: Diffuse, unorganized, easily removed. Facilitate motility and surface attachment.
External Structures: Pili, Fimbriae, and Flagella
Pili and Fimbriae
These are filamentous structures extending from the cell surface, involved in attachment, motility, and genetic exchange.
Fimbriae: Short, bristle-like fibers, numerous (up to 1000/cell), found in both Gram-positive and Gram-negative bacteria. Mediate attachment to surfaces and DNA uptake.
Pili: Longer, less numerous, found mainly in Gram-negative bacteria. Specialized pili (sex pili) are involved in conjugation (gene transfer).
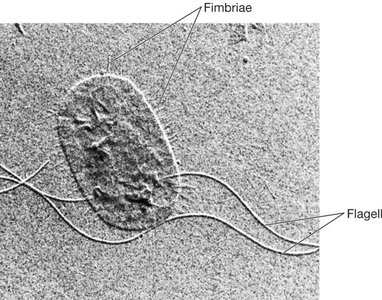
Flagella
Flagella are long, whip-like structures responsible for bacterial motility. They are composed of three main parts: filament, hook, and basal body. The arrangement of flagella varies among species:
Monotrichous: Single flagellum at one end.
Amphitrichous: One flagellum at each end.
Lophotrichous: Cluster of flagella at one or both ends.
Peritrichous: Flagella distributed over the entire cell surface.
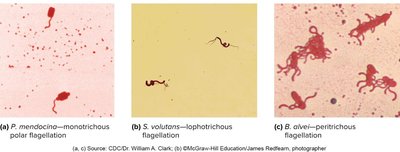

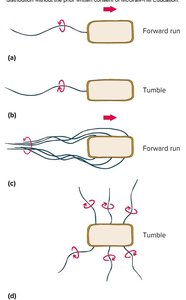
Flagella rotate like propellers, powered by the proton motive force (PMF). Counterclockwise rotation results in a forward run, while clockwise rotation causes tumbling and reorientation.
Bacterial Cell Wall
Structure and Function
The cell wall is a rigid structure that surrounds the plasma membrane, providing shape and protection from osmotic lysis. The main component is peptidoglycan (murein), a polymer of sugars and amino acids forming a mesh-like layer.
Peptidoglycan: Composed of alternating N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM) residues, crosslinked by peptide chains for strength.
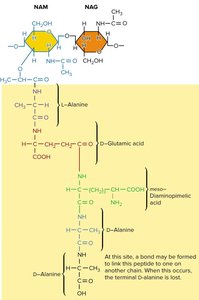
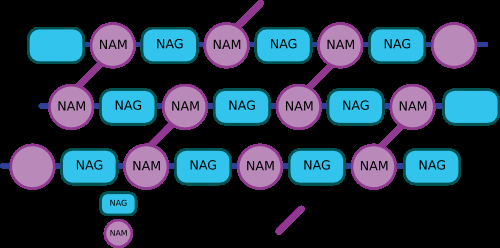
Gram-Positive vs. Gram-Negative Cell Walls
Bacteria are classified based on their cell wall structure and Gram stain reaction:
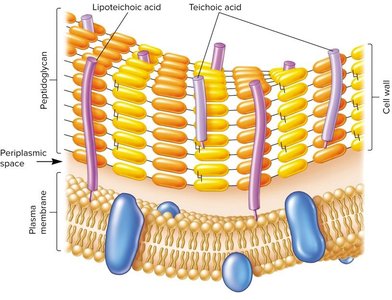
Gram-positive: Thick peptidoglycan layer, stains purple, contains teichoic acids and lipoteichoic acids.
Gram-negative: Thin peptidoglycan layer, stains pink/red, has an outer membrane containing lipopolysaccharide (LPS), no teichoic acids.
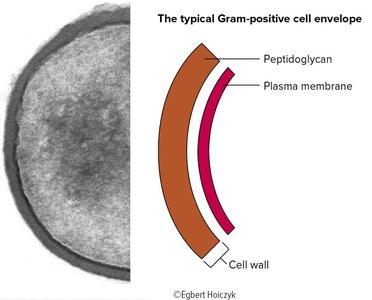
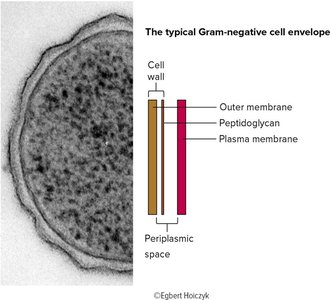
Teichoic Acids and Lipoteichoic Acids (Gram-Positive)
Teichoic acids: Polymers of glycerol or ribitol phosphate, contribute to cell wall maintenance, protection, and negative charge.
Lipoteichoic acids (LTA): Anchored in the plasma membrane, can stimulate immune responses.
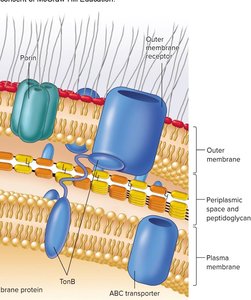
Outer Membrane and LPS (Gram-Negative)
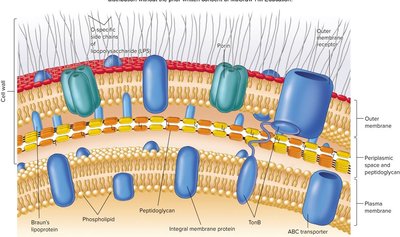
Outer membrane: Asymmetric lipid bilayer with LPS in the outer leaflet, provides a permeability barrier.
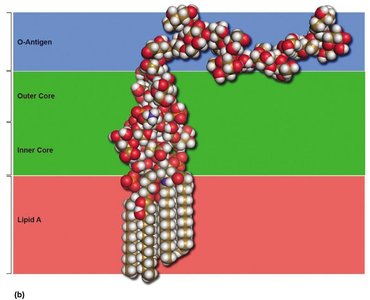
Lipopolysaccharide (LPS): Consists of O antigen (elicits immune response), core polysaccharide, and Lipid A (endotoxin).
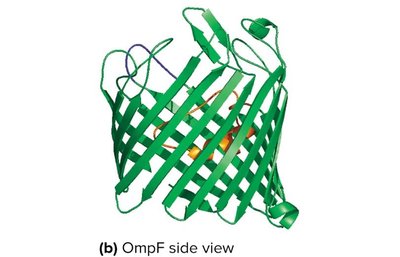
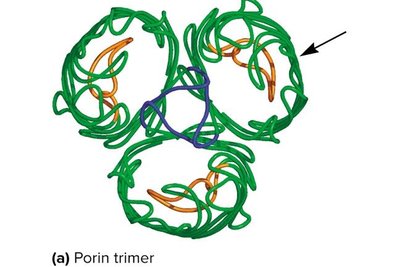
Porins: Channel proteins that allow passage of small molecules.
Gram Stain Mechanism
The Gram stain differentiates bacteria based on cell wall structure:
Crystal violet stains all cells.
Iodine forms a complex with the dye.
Ethanol decolorizes Gram-negative cells (thin peptidoglycan), but not Gram-positive (thick peptidoglycan).
Safranin counterstains Gram-negative cells pink/red.
Plasma Membrane Structure and Function
Composition and Organization
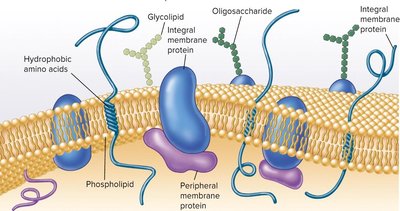
The plasma membrane is a phospholipid bilayer with embedded proteins, forming a dynamic and selectively permeable barrier. It is the site of crucial metabolic processes and environmental sensing.
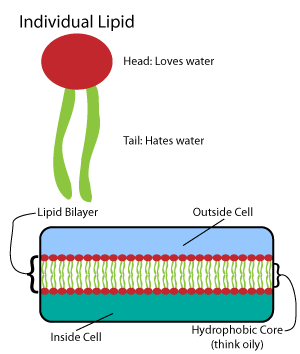
Amphipathic lipids: Have hydrophilic (polar) heads and hydrophobic (non-polar) tails.
Membrane proteins: Integral (embedded) and peripheral (loosely attached), involved in transport, signaling, and enzymatic activity.
Membrane Protein Functions
Solute transport
Secretion
Aquaporins (water channels)
Receptors and signaling
Enzymatic activity (e.g., ATP synthase)
Quorum sensing (regulation of gene expression in response to cell density)
Nutrient Uptake Mechanisms
Types of Transport
Bacteria utilize several mechanisms to transport nutrients across the plasma membrane:
Passive diffusion: Movement down a concentration gradient (e.g., H2O, O2, CO2).
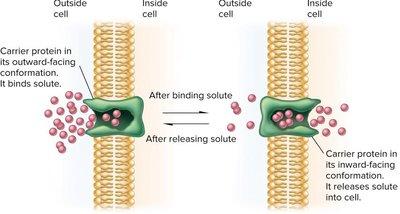
Facilitated diffusion: Uses carrier or channel proteins, no energy required, substrate-specific.
Active transport: Moves substances against a concentration gradient, requires energy (ATP or proton motive force).
Group translocation: Chemical modification of the transported molecule (e.g., PTS system for sugars).
Summary Table: Comparison of Gram-Positive and Gram-Negative Cell Walls
Feature | Gram-Positive | Gram-Negative |
|---|---|---|
Peptidoglycan Layer | Thick | Thin |
Teichoic Acids | Present | Absent |
Outer Membrane | Absent | Present |
Lipopolysaccharide (LPS) | Absent | Present |
Periplasmic Space | Small or absent | Large |
Gram Stain Color | Purple | Pink/Red |
Additional info: The cell envelope and its associated structures are critical for bacterial survival, pathogenicity, and antibiotic resistance. Understanding these features is foundational for advanced studies in microbiology, including antimicrobial drug action and immune evasion mechanisms.
