 Back
BackBacterial Gene Expression: Transcription and Translation
Study Guide - Smart Notes

Bacterial Gene Expression
Overview of the Central Dogma
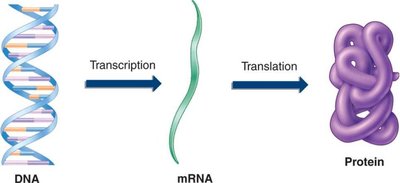
The central dogma of molecular biology describes the flow of genetic information within a biological system. It explains how genetic information stored in DNA is transcribed into messenger RNA (mRNA) and then translated into proteins, which perform essential cellular functions.
Transcription: Synthesis of single-stranded mRNA from a DNA template. (1 strand of DNA is transcribed)
Translation: Conversion of the mRNA sequence into an amino acid sequence, forming a polypeptide (protein).
Types of RNA
Functional Roles of RNA Molecules
Different types of RNA are encoded by distinct genes and play specialized roles in gene expression:
mRNA (Messenger RNA): Encodes the amino acid sequence of a polypeptide; product of transcription.
tRNA (Transfer RNA): Transports amino acids to ribosomes during translation.
rRNA (Ribosomal RNA): Forms complexes with proteins to create ribosomes, the site of mRNA translation.
Summary: Only mRNA is directly involved in transcription, while tRNA and rRNA are essential for translation.
Gene Structure and Organization
Basic Features of Protein-Coding Genes
The basic unit of genetic information.
Gene - Linear sequence of nucleotides with a fixed start point and end point.
Gene - also defined as the nucleic acid sequence that codes for a polypeptide,
e.g. tRNA or rRNA.
Codons are found in mRNA and code for single amino acids.
A gene is a linear sequence of nucleotides with defined start and end points. In bacteria, genes are typically continuous sequences (no introns) that code for polypeptides, tRNA, or rRNA. Codons within mRNA specify individual amino acids.
Promoter: DNA region where RNA polymerase binds to initiate transcription; not transcribed.
Is the recognition/binding site for RNA polymerase.
• Functions to orient polymerase.
• Shine-Dalgarno sequence important for initiation of translation.
• Begins with DNA sequence 3’-TAC-5’, that twill be transcribed into start codon AUG.
Coding region ends with a stop codon, immediately followed by the trailer sequence which contains a terminator sequence used to stop transcription. Coding sequence is continuous (no introns that are found in eukaryotic cells).
Shine-Dalgarno Sequence: Important for translation initiation in bacteria.
Start Codon: Usually AUG, marks the beginning of the coding region.
Stop Codon: Signals the end of translation (UAA, UAG, UGA).
Trailer Sequence: Follows the stop codon and contains signals for transcription termination.
DNA sense strand: also known as the coding strand
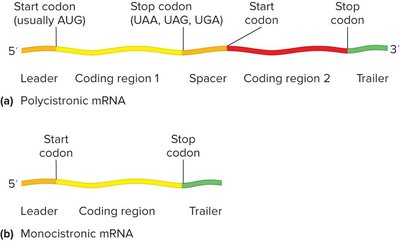
Operons and Polycistronic mRNA
Bacterial genes involved in related processes are often organized into operons—clusters of genes transcribed from a single promoter, resulting in polycistronic mRNA (encodes multiple proteins). Monocistronic mRNA encodes a single protein. Polycistronic mRNA often found in bacteria and archaea.
Transcription in Bacteria
RNA Polymerase Structure and Function
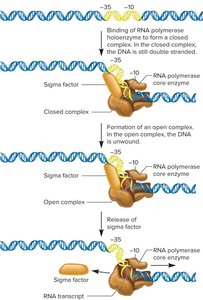
Bacterial RNA polymerase is a multi-subunit enzyme responsible for synthesizing RNA from a DNA template. The holoenzyme consists of the core enzyme (five subunits: two α, β, β', ω) and a sigma (σ) factor. The sigma factor is essential for promoter recognition but not for RNA synthesis (no catalytic activity).
Transcription Initiation
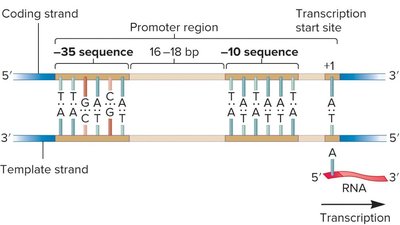
Transcription begins when the RNA polymerase holoenzyme binds to the promoter region. The sigma factor recognizes two key promoter regions: the -35 and -10 sequences (the latter is also called the Pribnow box, with the consensus sequence TATAAT). Once bound, the DNA unwinds at the -10 region to form an open complex, and the sigma factor dissociates after initiation. Only a short segment of DNA is transcribed.
Transcription Elongation
During elongation, RNA polymerase moves along the DNA, unwinding it and synthesizing RNA in the 5' to 3' direction. A transcription bubble forms, within which a temporary RNA:DNA hybrid exists.
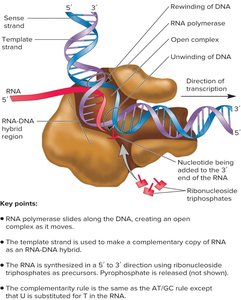
Transcription Termination
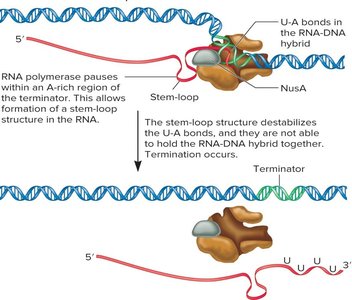
Termination occurs when RNA polymerase dissociates from the DNA template. There are two main mechanisms:
Factor-independent (intrinsic) termination: Involves the formation of a stem-loop structure in the RNA, destabilizing the RNA-DNA hybrid and causing release.
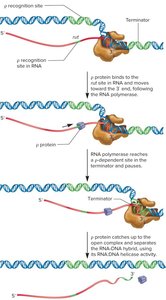
Rho-dependent termination: Requires the rho (ρ) protein, which binds to the RNA and moves toward the polymerase, causing dissociation when it reaches the transcription complex.
The Genetic Code
Structure and Properties of the Genetic Code
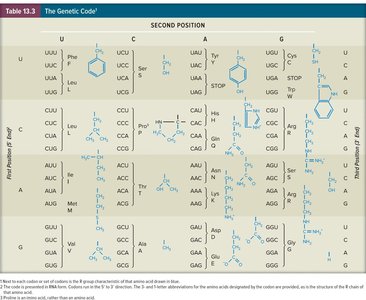
The genetic code is composed of codons—three-nucleotide sequences on mRNA that specify amino acids. The code is nearly universal and degenerate, meaning multiple codons can encode the same amino acid.
Start Codon: AUG (methionine) marks the initiation of translation.
Stop Codons: UAA, UAG, UGA signal translation termination and do not encode amino acids.
Sense Codons: The 61 codons that specify amino acids.
Degeneracy: Up to six codons can code for a single amino acid.
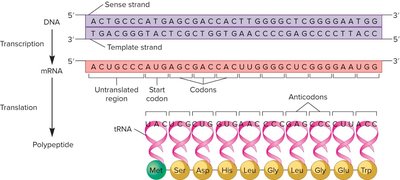

Wobble Hypothesis
The wobble hypothesis explains that the third position of the codon is less critical for base pairing, allowing a single tRNA to recognize multiple codons. This reduces the number of tRNAs needed and decreases the impact of some mutations.
Translation in Bacteria
Overview of Translation
Translation is the process by which the nucleotide sequence of mRNA directs the synthesis of a polypeptide. In bacteria, transcription and translation are often coupled, and multiple ribosomes can translate a single mRNA simultaneously (polyribosome or polysome).
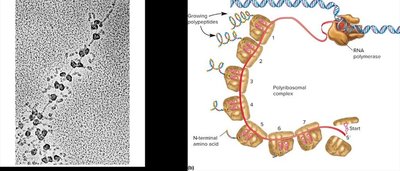
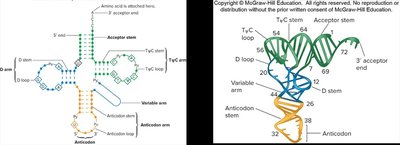
Transfer RNA (tRNA) Structure and Function
tRNA molecules have a characteristic cloverleaf secondary structure and a complex tertiary structure due to internal base pairing. Each tRNA has an anticodon complementary to the mRNA codon and an acceptor stem for amino acid attachment.
Amino Acid Activation
Amino acids are attached to their corresponding tRNAs by specific enzymes called aminoacyl-tRNA synthetases. There are at least 20 synthetases, one for each amino acid.
Bacterial Ribosome Structure
The bacterial ribosome is the site of protein synthesis and consists of two subunits:
30S subunit: Contains 16S rRNA and 21 proteins.
50S subunit: Contains 23S and 5S rRNA and 34 proteins.
70S ribosome: The functional complex in bacteria.
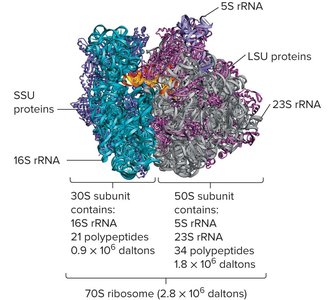
Role of Ribosomal RNA in Translation
rRNA molecules contribute to ribosome structure and function:
16S rRNA: Binds to the Shine-Dalgarno sequence on mRNA for initiation.
23S rRNA: Acts as a ribozyme, catalyzing peptide bond formation.
Initiation of Protein Synthesis
Initiation involves the assembly of the ribosome, mRNA, and initiator tRNA (N-formyl methionine-tRNA in bacteria). The Shine-Dalgarno sequence aligns the mRNA with the 16S rRNA. Initiation factors (IF-1, IF-2) and GTP are required for complex formation.
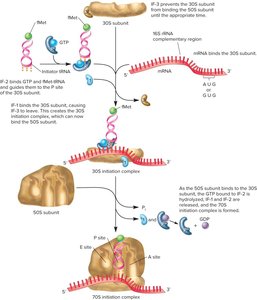
Elongation of the Polypeptide Chain
Elongation involves the sequential addition of amino acids to the growing polypeptide chain. The ribosome has three tRNA binding sites:
P (Peptidyl) site: Holds the tRNA with the growing polypeptide.
A (Aminoacyl) site: Binds incoming aminoacyl-tRNA.
E (Exit) site: Releases empty tRNA.
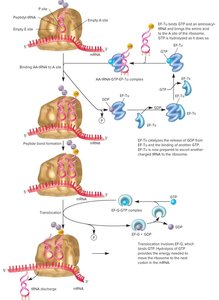
Translocation
During translocation, the ribosome moves one codon along the mRNA, shifting the peptidyl-tRNA from the A site to the P site and releasing the empty tRNA from the E site. This process requires GTP hydrolysis.
Termination of Protein Synthesis
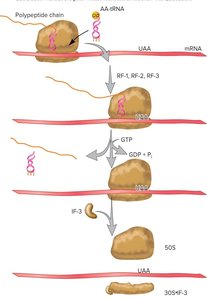
Termination occurs when a stop codon (UAA, UAG, UGA) is encountered. Release factors (RFs) recognize the stop codon, promote the release of the polypeptide, and dissociate the ribosome. GTP hydrolysis is required for this process.
Additional info: The notes above integrate and expand upon the provided slides, ensuring a comprehensive, self-contained summary suitable for microbiology students studying bacterial gene expression, transcription, and translation.
