 Back
Backlec 19:Bacterial Gene Transfer, DNA Replication, and Molecular Information Flow
Study Guide - Smart Notes

Bacterial Gene Transfer and Evolution
Overview
Bacteria possess remarkable genetic flexibility, allowing them to adapt rapidly to environmental changes. This adaptability is driven by both mutation and horizontal gene transfer (HGT), which together fuel microbial evolution and the spread of traits such as antibiotic resistance.
DNA Replication
Semiconservative Replication
DNA replication is the process by which a cell copies its DNA, producing two identical DNA molecules from one original molecule. It is semiconservative, meaning each new DNA molecule contains one original (parent) strand and one newly synthesized (daughter) strand.
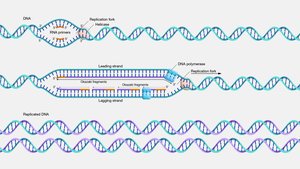
Key Enzymes Involved
Helicase: Unwinds the DNA double helix by breaking hydrogen bonds between base pairs.
Topoisomerase: Relieves supercoiling and twisting stress ahead of the replication fork.
Primase: Synthesizes short RNA primers to provide a starting point for DNA polymerase.
DNA Polymerase III: Extends the new DNA strand by adding nucleotides in the 5′ → 3′ direction.
DNA Polymerase I: Removes RNA primers and replaces them with DNA.
DNA Ligase: Joins Okazaki fragments on the lagging strand to create a continuous DNA strand.
Single-strand binding proteins (SSBs): Stabilize unwound DNA strands and prevent reannealing.
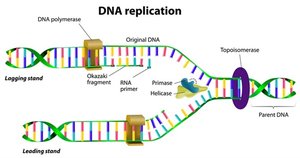
Steps of DNA Replication
Initiation: Replication begins at origins of replication. Initiator proteins open the helix, forming a replication fork.
Elongation:
Leading Strand: Synthesized continuously in the direction of the replication fork.
Lagging Strand: Synthesized discontinuously in short segments called Okazaki fragments, opposite to fork movement.
Termination: RNA primers are removed, replaced with DNA, and DNA ligase seals nicks in the backbone.
Proofreading: DNA polymerases correct errors to ensure high fidelity.
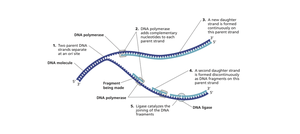
Molecular Information Flow: Transcription and Translation
Transcription
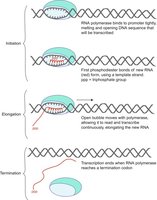
Transcription is the process by which a gene's DNA sequence is copied into messenger RNA (mRNA). In prokaryotes, this occurs in the cytoplasm; in eukaryotes, in the nucleus.
Key Enzyme: RNA polymerase binds to the promoter region, unwinds DNA, and synthesizes RNA using ribonucleotides (A, U, C, G).
Main Stages:
Initiation: RNA polymerase attaches to the promoter.
Elongation: RNA polymerase reads the DNA template and builds the RNA strand.
Termination: RNA polymerase reaches a stop signal and releases the completed mRNA.
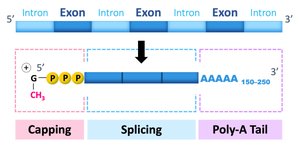
RNA Processing in Eukaryotes
Splicing: Removal of non-coding introns.
5′ Cap: Added to the beginning for stability and ribosome recognition.
Poly-A Tail: Added to the end for stability and export from the nucleus.
Translation
Translation is the process where the information in mRNA is used to synthesize a protein (polypeptide chain). This occurs at ribosomes in the cytoplasm.
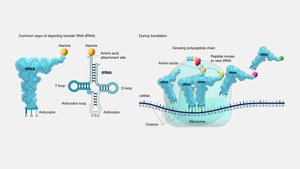
mRNA: Carries genetic instructions in codons (three-nucleotide sequences).
tRNA: Brings the correct amino acid to the ribosome, matching codons with its anticodon.
Ribosome: Reads mRNA and catalyzes peptide bond formation between amino acids.
Initiation: Ribosome binds to mRNA at the start codon (AUG); first tRNA brings methionine.
Elongation: Ribosome moves along mRNA, tRNAs bring amino acids, and the protein chain grows.
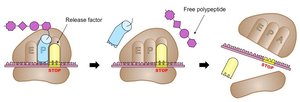
Termination: Ribosome reaches a stop codon (UAA, UAG, UGA); protein is released.
Protein Folding, Modification, and Degradation
Protein Folding
Newly synthesized proteins must fold into specific three-dimensional shapes to function. Molecular chaperones (e.g., Hsp70, Hsp60) assist in proper folding and prevent aggregation. Misfolded proteins can cause diseases such as Alzheimer's and cystic fibrosis.
Protein Modification
Phosphorylation: Addition of phosphate groups to regulate activity.
Glycosylation: Addition of sugars for stability and recognition.
Lipidation: Attachment of lipids for membrane association.
Cleavage: Proteolytic processing to activate proteins.
Protein Degradation
Ubiquitin–Proteasome System: Tags proteins with ubiquitin for degradation by the proteasome.
Autophagy: Degrades larger protein aggregates or organelles via lysosomes.
Microbial Mutation and DNA Repair
Types of Mutations
Spontaneous mutations: Occur naturally during DNA replication or due to chemical changes.
Induced mutations: Caused by mutagens such as UV light, X-rays, or chemicals.
Point mutations: Single nucleotide changes (silent, missense, nonsense).
Frameshift mutations: Insertions or deletions that shift the reading frame.
Chromosomal rearrangements: Large-scale changes like duplications or inversions.
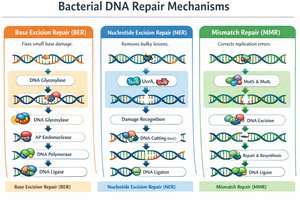
DNA Repair Mechanisms
DNA repair systems are essential for maintaining genome stability and cell survival. Without repair, mutations accumulate, potentially leading to cell death.
Bacterial Gene Transfer and Horizontal Gene Transfer (HGT)
Overview
Bacteria can acquire new genes from other bacteria or the environment, a process known as horizontal gene transfer (HGT). This is distinct from vertical inheritance (parent to offspring) and is a major driver of bacterial evolution.
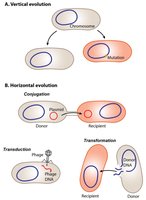
Main Types of Gene Transfer
Transformation: Uptake of free DNA from the environment.
Transduction: Transfer of DNA by bacteriophages (viruses that infect bacteria).
Conjugation: Direct transfer of DNA (usually plasmids) between bacteria via a pilus.
Evolutionary Importance
HGT enables rapid adaptation by spreading traits such as antibiotic resistance, new metabolic pathways, and virulence factors.
Mobile Genetic Elements (MGEs)
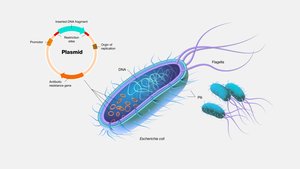
Plasmids
Plasmids are small, circular DNA molecules that replicate independently of the bacterial chromosome. They often carry genes for antibiotic resistance or special metabolic functions and are commonly transferred by conjugation.
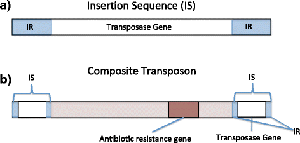
Transposable Elements ("Jumping Genes")
Insertion Sequences (IS): Simple elements containing only the gene for transposase, which catalyzes movement.
Transposons (Tn): Larger elements that include transposase and additional genes, often for antibiotic resistance.
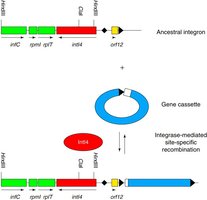
Integrons
Integrons are genetic elements that can capture and express multiple gene cassettes, frequently conferring multidrug resistance.
Bacteriophages (Phages)
Bacteriophages are viruses that infect bacteria. Their DNA can integrate into the bacterial chromosome as a prophage. Lysogenic conversion can provide new traits, such as toxin production, to the host bacterium.
Gene Transfer Mechanism | Description | Key Features |
|---|---|---|
Transformation | Uptake of free DNA from environment | Requires competence; DNA often from dead cells |
Transduction | DNA transfer via bacteriophage | Generalized or specialized; can transfer toxin genes |
Conjugation | Direct DNA transfer between cells | Requires cell-to-cell contact; often plasmid-mediated |
