 Back
BackBacterial Genomes and Replication: Structure, Organization, and Regulation
Study Guide - Smart Notes

DNA: The Genetic Material
Structure and Function of DNA
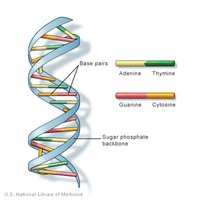
Deoxyribonucleic acid (DNA) is the hereditary molecule in almost all living organisms, encoding the instructions for cellular structure, function, and inheritance. DNA is composed of two antiparallel strands forming a double helix, with a sugar-phosphate backbone and nitrogenous bases paired in the center.
Key Bases: Adenine (A), Thymine (T), Cytosine (C), Guanine (G)
Base Pairing: A pairs with T, C pairs with G via hydrogen bonds
Genetic Code: The sequence of bases encodes genetic information for protein synthesis
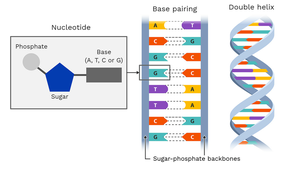
Location and Components of DNA
In eukaryotes, DNA is mainly found in the nucleus, with small amounts in mitochondria. Each DNA molecule is a polymer of nucleotides, each consisting of a deoxyribose sugar, phosphate group, and nitrogenous base.
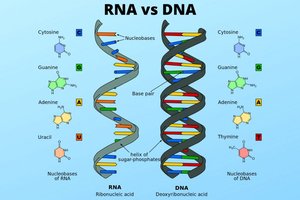
DNA vs. RNA
While DNA serves as the stable, long-term storage of genetic information, ribonucleic acid (RNA) is involved in gene expression and protein synthesis. Messenger RNA (mRNA) carries genetic instructions from DNA to ribosomes.
Genome Organization
Definition and Function
The genome is the complete set of genetic material in an organism, containing all instructions for life, including cell structure, metabolism, growth, reproduction, and environmental response.
Gene: A DNA sequence encoding a functional product, usually a protein
Genome Organization in Prokaryotes
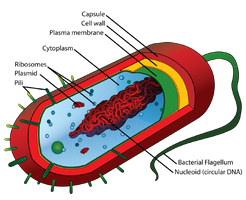
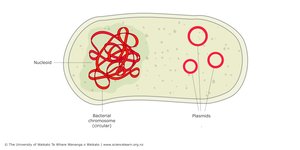
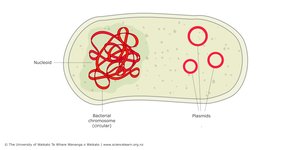
Prokaryotic genomes are typically compact, with one circular chromosome located in the nucleoid region (not membrane-bound). Genes are often organized into operons, and there is little non-coding DNA.
Operons in Prokaryotes
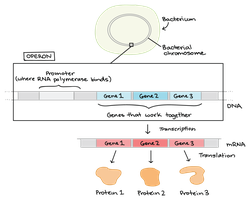
An operon is a cluster of genes under the control of a single promoter, transcribed together as one mRNA. This allows coordinated expression of genes involved in the same pathway.
Promoter: Site where RNA polymerase binds to initiate transcription
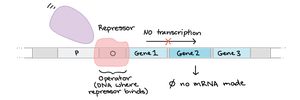
Operator: Regulatory region where proteins bind to turn operon on/off
Structural Genes: Encode proteins/enzymes for a pathway
Regulatory Gene: Encodes a protein (repressor/activator) that regulates the operon
Lac Operon Example
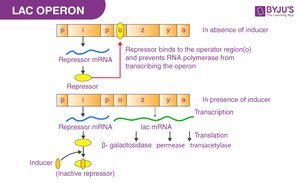
The lac operon in E. coli is an inducible operon that is off by default. When lactose is present, it induces the operon, allowing the cell to produce enzymes for lactose metabolism.
β-galactosidase (lacZ): Breaks lactose into glucose and galactose
Permease (lacY): Transports lactose into the cell
Transacetylase (lacA): Assists lactose metabolism
Plasmids
Plasmids are small, circular DNA molecules separate from the main chromosome. They often carry genes for antibiotic resistance, virulence, or metabolic pathways and can be transferred between bacteria, contributing to rapid evolution.
Genome Organization in Eukaryotes
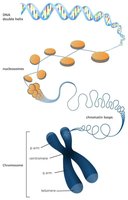
Chromosome Structure and DNA Packaging
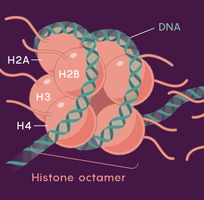
Eukaryotic DNA is organized into multiple linear chromosomes located in the nucleus. DNA wraps around histone proteins to form nucleosomes, which further coil to form chromatin and chromosomes, allowing efficient packaging of large amounts of DNA.
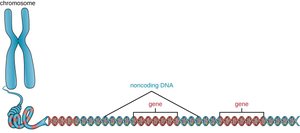
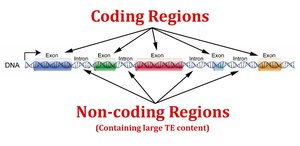
Coding vs. Non-Coding DNA
Eukaryotic genomes contain large amounts of non-coding DNA, including introns, regulatory regions, and repetitive sequences. Only about 1–2% of human DNA codes for proteins.
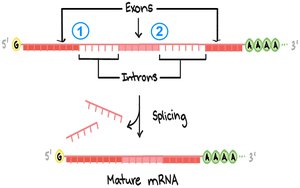
Introns and RNA Splicing
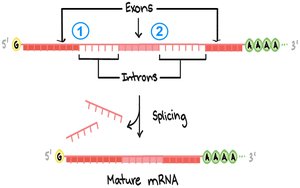
Introns are non-coding sequences within genes, transcribed into RNA but removed during RNA processing. Splicing is the process of removing introns and joining exons to form mature mRNA.
Spliceosome: A complex of snRNAs and proteins that removes introns
Self-Splicing Introns: Some introns can self-remove (Group I and II), acting as ribozymes
Functional Roles of Introns
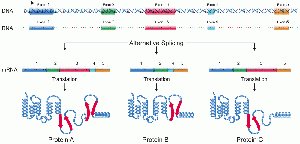
Alternative Splicing: Allows different combinations of exons, producing multiple proteins from one gene
Gene Regulation: Introns may contain regulatory elements
Genetic Stability: Introns buffer against mutations
Non-coding RNAs: Some introns encode microRNAs (miRNAs) and snoRNAs
MicroRNAs (miRNAs) and Gene Silencing
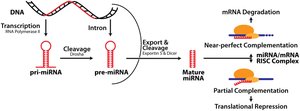
MicroRNAs (miRNAs) are small non-coding RNAs (19–25 nucleotides) that regulate gene expression post-transcriptionally. They bind to target mRNAs via the RNA-induced silencing complex (RISC), leading to translational repression or mRNA degradation.
Target Recognition: miRNA matches a seed region to complementary mRNA
RISC Complex: miRNA binds RISC, which then binds mRNA
Gene Silencing: Blocks translation or degrades mRNA
Plasmids and Secondary Chromosomes
Plasmids
Plasmids are non-essential, mobile DNA elements that provide adaptive advantages, such as antibiotic resistance or metabolic capabilities. They replicate independently and can be transferred between bacteria.
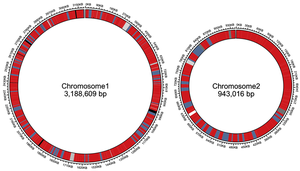
Secondary Chromosomes
Some bacteria possess secondary chromosomes, which are larger than plasmids and essential for survival, containing housekeeping genes. Replication is coordinated with the main chromosome.
Chromosomization
Over evolutionary time, plasmids can acquire essential genes and become secondary chromosomes, a process called chromosomization.
Eukaryotic and Archaeal Chromosomes
Comparative Structure
Archaea and eukaryotes share similarities in chromosome organization, distinct from bacteria. Archaea usually have one circular chromosome with histone-like proteins, while eukaryotes have multiple linear chromosomes with true histones and telomeres.
DNA Packaging and Replication Origins
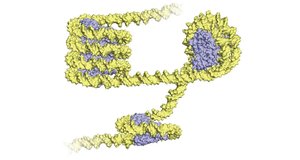
Both archaea and eukaryotes use histone proteins for DNA packaging. Archaea use archaeal histones, while eukaryotes use H2A, H2B, H3, and H4. Archaea often have multiple origins of replication, and their information-processing proteins are more similar to eukaryotes than bacteria.
Feature | Bacteria | Archaea | Eukaryotes |
|---|---|---|---|
Chromosome Structure | Single, circular | Single, circular | Multiple, linear |
DNA Packaging | No histones | Histone-like proteins | Histones (H2A, H2B, H3, H4) |
Replication Origins | One | Multiple | Multiple |
Non-coding DNA | Minimal | Variable | Abundant |
