 Back
BackBiochemical Identification of Bacteria: Hydrolytic Enzymes, Carbohydrate Utilization, and Multitest Systems
Study Guide - Smart Notes

Hydrolytic Enzymes in Bacterial Identification
DNase Test
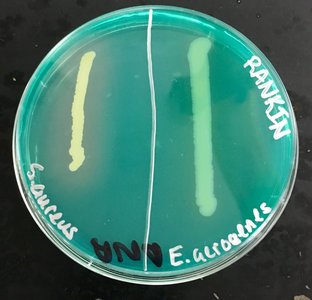
The DNase test is used to determine whether a bacterium produces the enzyme deoxyribonuclease (DNase), which hydrolyzes DNA. DNase agar contains DNA and methyl green, which binds to intact DNA. When DNase is produced, DNA is degraded, releasing methyl green and resulting in a clear zone around the bacterial growth.
Aim: To detect DNase activity for bacterial differentiation.
Media: DNase agar with methyl green and DNA.
Interpretation: A clear halo around growth indicates DNase production (positive); no halo indicates a negative result.
Example: Staphylococcus aureus is typically DNase positive, while Enterobacter aerogenes is negative.
Gelatinase Test
The gelatinase test identifies bacteria that produce gelatinase, an enzyme that hydrolyzes gelatin (a collagen derivative). Gelatinase activity is associated with bacterial virulence.
Aim: To detect gelatinase production.
Media: Nutrient gelatin (gelatin as solidifying agent).
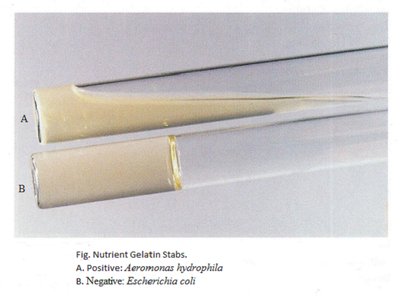
Interpretation: Liquefaction of the medium after incubation and cooling indicates a positive result; solid medium indicates a negative result.
Example: Aeromonas hydrophila is gelatinase positive; Escherichia coli is negative.
Starch Hydrolysis Test
This test detects the ability of bacteria to hydrolyze starch using the enzyme amylase. After incubation, iodine is added to the plate; it reacts with starch to produce a dark color. A clear zone around growth indicates starch hydrolysis.
Aim: To detect amylase production and starch hydrolysis.
Media: Starch agar.
Interpretation: Clear zone after iodine addition indicates positive amylase activity; dark coloration indicates negative result.
Example: Bacillus species are often positive for starch hydrolysis.
Carbohydrate Utilization Tests
Phenol Red Broth Test
Phenol red broth is used to determine if an organism can ferment a specific carbohydrate (e.g., glucose, lactose). The broth contains phenol red (pH indicator) and a Durham tube to detect gas production.
Aim: To detect fermentation of specific sugars and gas production.
Media: Phenol red broth with added carbohydrate and Durham tube.
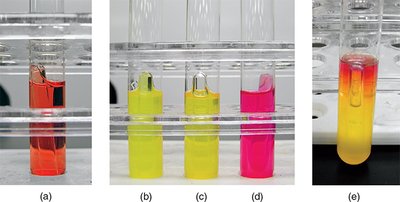
Interpretation:
Yellow color: Acid production (positive fermentation).
Bubble in Durham tube: Gas production.
Pink color: Alkaline reaction (protein catabolism, no fermentation).
Example: Escherichia coli ferments glucose with acid and gas production.

Triple Sugar Iron Agar (TSIA) Test
TSIA differentiates bacteria based on their ability to ferment glucose, lactose, and/or sucrose, and to reduce sulfur. The slant/butt format allows detection of multiple reactions in one tube.
Aim: To differentiate bacteria by sugar fermentation and sulfur reduction.
Media: TSIA slant (contains glucose, lactose, sucrose, iron salt, and phenol red).
Interpretation:
Yellow slant/yellow butt (A/A): Fermentation of glucose and at least one disaccharide.
Red slant/yellow butt (K/A): Glucose fermentation only (reversion to protein catabolism on slant).
Black precipitate: H2S production (sulfur reduction).
No change: Non-fermenter.
Example: Salmonella typhimurium produces H2S (black butt).


Biochemical Pathway Tests
Litmus Milk Test
Litmus milk is a complex medium used to differentiate bacteria based on their ability to metabolize lactose and casein, and to reduce litmus. It detects acid/alkaline reactions, reduction, and digestion.
Aim: To differentiate bacteria by lactose fermentation, casein hydrolysis, and litmus reduction.
Media: Litmus milk (skim milk, litmus indicator).
Interpretation:
Pink: Acid (lactose fermentation).
Blue: Alkaline (casein breakdown).
White: Reduction of litmus.
Clot: Protein denaturation (acid or stormy fermentation).
Clear/whey: Digestion (peptonization).

IMViC Tests
The IMViC series (Indole, Methyl Red, Voges-Proskauer, Citrate) is used to differentiate gram-negative enteric bacteria.
SIM Test: Detects sulfur reduction, indole production, and motility.
MRVP Test: Methyl Red detects mixed acid fermentation; Voges-Proskauer detects acetoin production.
Citrate Test: Determines if citrate can be used as a sole carbon source (bromthymol blue indicator).

Urease Test
The urease test identifies bacteria capable of hydrolyzing urea to ammonia and carbon dioxide using the enzyme urease. Ammonia production raises the pH, changing the color of phenol red indicator.
Aim: To detect rapid or slow urease production.
Media: Urease broth (urea, phenol red, phosphate buffer).
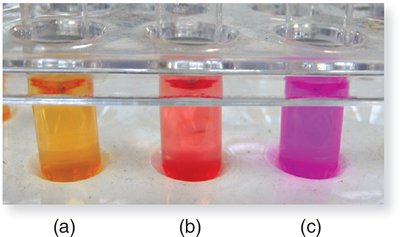
Interpretation:
Bright pink: Rapid urease producer (positive).
Dull pink: Weak positive.
Orange/red: Negative.
Example: Proteus species are rapid urease producers.

Nitrate Reductase Test
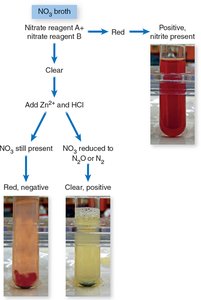
This test determines the ability of bacteria to reduce nitrate (NO3-) to nitrite (NO2-) or further to nitrogen gas. Reagents detect nitrite; zinc and HCl are used to confirm negative results.
Aim: To detect nitrate and nitrite reductase activity.
Media: Nitrate broth with potassium nitrate.
Interpretation:
Red after reagents: Positive for nitrite.
Red after zinc/HCl: Negative (nitrate not reduced by bacteria).
No color after zinc/HCl: Positive for complete reduction to N2 or N2O.
Oxygen Metabolism: Catalase and Oxidase Tests
Catalase Test
The catalase test detects the presence of catalase, which breaks down hydrogen peroxide into water and oxygen. Bubbles indicate a positive result.
Aim: To detect catalase enzyme in bacteria.
Interpretation: Bubble formation after adding H2O2 indicates a positive result.


Oxidase Test
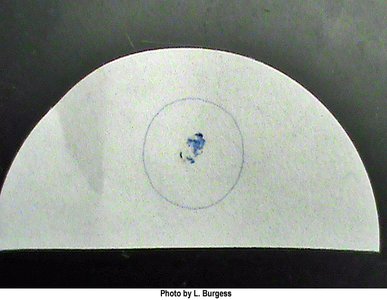
The oxidase test identifies bacteria that produce cytochrome c oxidase, an enzyme of the electron transport chain. The reagent turns blue when oxidized.
Aim: To detect cytochrome c oxidase.
Interpretation: Blue color indicates a positive result; no color change is negative.

Multiple Test Systems: EnteroPluri and API 20E
EnteroPluri-Test
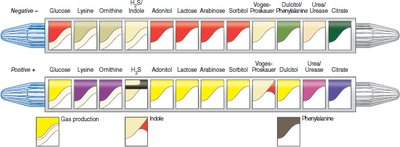
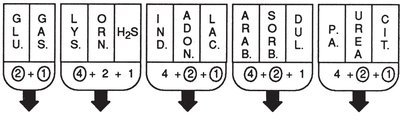
The EnteroPluri-Test is a multitest system for identifying gram-negative, oxidase-negative bacteria. It contains 12 media sectors for 15 biochemical reactions. Results are scored and combined into a code for identification.
Aim: Rapid identification of Enterobacteriaceae and related bacteria.
Interpretation: Color changes in each sector are scored; the sum forms a code matched to a database for identification.
API 20E Test System
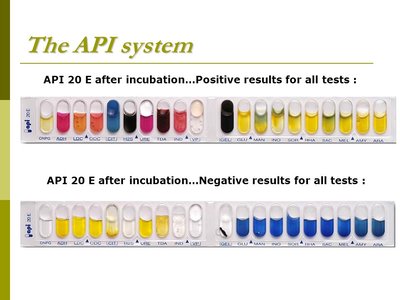
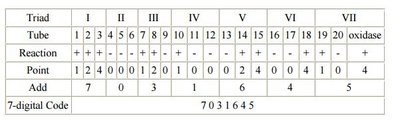
The API 20E system is a panel of 20 miniaturized biochemical tests for identifying Enterobacteriaceae. Each compartment contains dehydrated media for a specific test. After inoculation and incubation, results are interpreted using a color chart and scored to generate a 7-digit code for organism identification.
Aim: Identification of Enterobacteriaceae and other gram-negative rods.
Interpretation: Positive and negative results are scored in triplets (1, 2, 4), summed, and combined into a code.
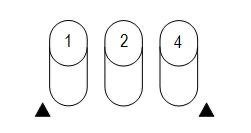
Additional info: These biochemical tests are essential for the identification and differentiation of clinically relevant bacteria, especially in medical microbiology and diagnostic laboratories. Understanding the principles, media, and interpretation of each test is crucial for accurate microbial identification.
