Back
BackBiochemistry of the Genome & Mechanisms of Microbial Genetics: Study Notes
Study Guide - Smart Notes

Biochemistry of the Genome
DNA Nucleotides (Deoxyribonucleotides)
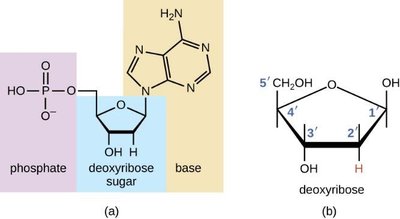
DNA is composed of building blocks called deoxyribonucleotides, each consisting of a deoxyribose sugar, a phosphate group, and a nitrogenous base. The five carbons in deoxyribose are numbered 1' through 5', which is important for understanding the directionality of DNA strands.
Deoxyribose sugar: A five-carbon sugar lacking an oxygen atom at the 2' position compared to ribose.
Phosphate group: Attached to the 5' carbon of the sugar.
Nitrogenous base: Attached to the 1' carbon of the sugar; can be a purine or pyrimidine.
Pyrimidines and Purines
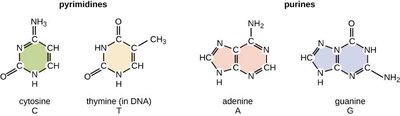
Nitrogenous bases in DNA are classified as either purines or pyrimidines. Purines have a double-ring structure, while pyrimidines have a single-ring structure.
Pyrimidines: Cytosine (C), Thymine (T) (unique to DNA)
Purines: Adenine (A), Guanine (G)
Sugar-Phosphate Backbone
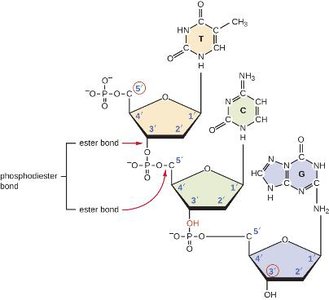
Nucleotides are joined together by phosphodiester bonds between the 5' phosphate of one nucleotide and the 3' hydroxyl of the next, forming the sugar-phosphate backbone of DNA. This backbone provides structural stability and directionality to the DNA strand.
5' end: Free phosphate group
3' end: Free hydroxyl group
The Double Helix
The structure of DNA is a double helix, as proposed by Watson and Crick. The sugar-phosphate backbones are on the outside, while the nitrogenous bases form the rungs of the ladder. The two strands are antiparallel, running in opposite directions (5' to 3' and 3' to 5').
Major and minor grooves: Created by the twisting of the helix, important for protein binding.
Antiparallel orientation: Essential for replication and transcription.
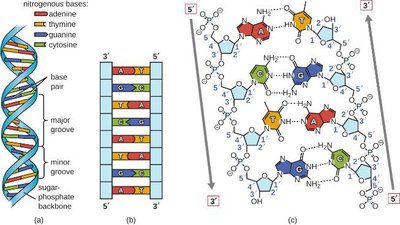
Complementary Base Pairing
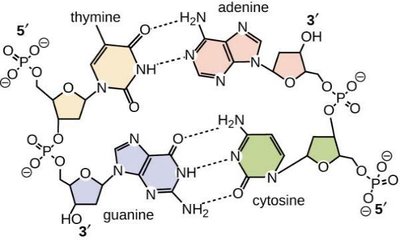
Hydrogen bonds form between specific pairs of nitrogenous bases on opposite DNA strands, stabilizing the double helix.
Adenine (A) pairs with Thymine (T): Two hydrogen bonds
Guanine (G) pairs with Cytosine (C): Three hydrogen bonds
RNA Nucleotides (Ribonucleotides)
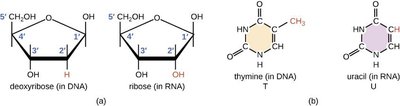
RNA is composed of ribonucleotides, which contain the sugar ribose and the base uracil instead of thymine. RNA is typically single-stranded but can form secondary structures through intramolecular base pairing.
Ribose: Has a hydroxyl group at the 2' position.
Uracil (U): Replaces thymine in RNA.
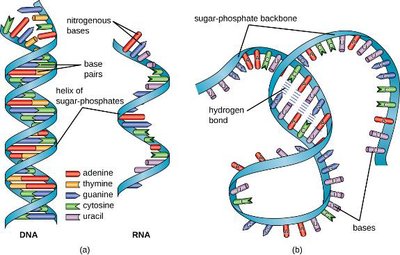
DNA vs. RNA Structure
DNA is double-stranded and forms a stable double helix, while RNA is usually single-stranded and can fold into complex three-dimensional shapes.
DNA: Double helix, contains thymine, deoxyribose sugar.
RNA: Single-stranded, contains uracil, ribose sugar.
Types of RNA
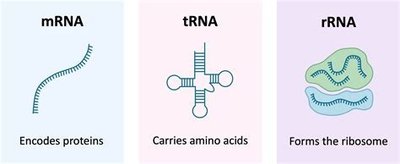
There are three main types of RNA, each with a distinct function in gene expression:
mRNA (messenger RNA): Carries genetic information from DNA to ribosomes for protein synthesis.
tRNA (transfer RNA): Brings amino acids to the ribosome during translation.
rRNA (ribosomal RNA): Structural and catalytic component of ribosomes.
Central Dogma & Gene Expression
The Central Dogma
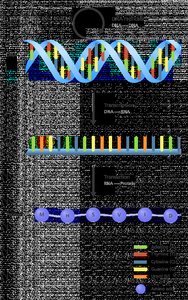
The central dogma of molecular biology describes the flow of genetic information: DNA is transcribed into mRNA, which is then translated into protein. This process is fundamental to all living organisms.
Transcription: Synthesis of mRNA from a DNA template.
Translation: Synthesis of protein from an mRNA template at the ribosome.
Gene Expression and Phenotype
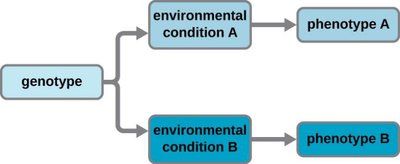
Gene expression determines phenotype by controlling which genes are transcribed and translated under specific environmental conditions. Cells with the same genotype can have different phenotypes due to differential gene expression.
Genotype: The genetic makeup of an organism.
Phenotype: Observable characteristics resulting from gene expression.
DNA Replication
Overview of DNA Replication
DNA replication is the process by which a cell copies its DNA before cell division, ensuring genetic information is passed to daughter cells (vertical gene transfer). The process involves several key enzymes and occurs at the origin of replication.
DNA Helicase: Unwinds the double helix.
DNA Primase: Synthesizes RNA primers.
DNA Polymerase: Adds nucleotides to the 3' end of the new strand.
DNA Ligase: Joins Okazaki fragments on the lagging strand.
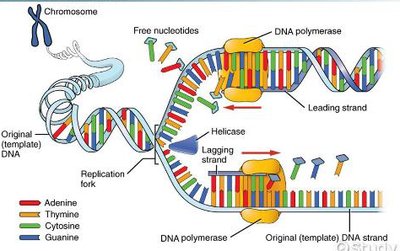
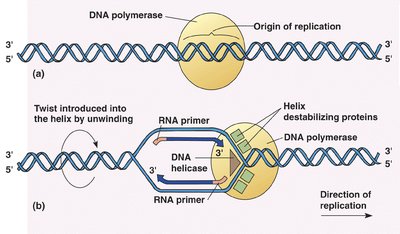
Initiation and Elongation of Replication
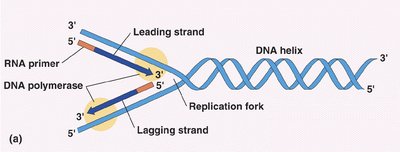
Replication begins at the origin of replication, where the DNA is unwound to form a replication fork. RNA primers are laid down, and new DNA is synthesized in the 5' to 3' direction. The leading strand is synthesized continuously, while the lagging strand is synthesized discontinuously in Okazaki fragments.
Leading strand: Synthesized continuously toward the replication fork.
Lagging strand: Synthesized discontinuously away from the fork.
Transcription and Translation
Transcription
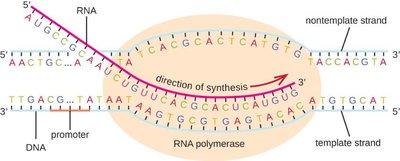
Transcription is the synthesis of RNA from a DNA template. RNA polymerase binds to a promoter sequence and synthesizes mRNA in the 5' to 3' direction until it reaches a termination sequence.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Elongation: RNA polymerase adds ribonucleotides complementary to the DNA template.
Termination: RNA polymerase releases the newly synthesized mRNA.
The Genetic Code
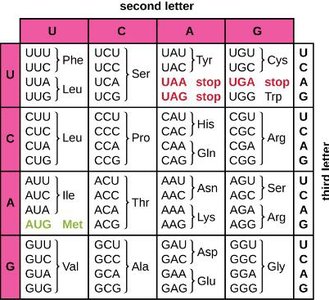
The genetic code is the relationship between mRNA codons and the amino acids they specify. Each codon consists of three nucleotides and codes for a specific amino acid or a stop signal.
Start codon (AUG): Codes for methionine and signals the start of translation.
Stop codons (UAA, UAG, UGA): Signal termination of translation.
Redundancy: Multiple codons can code for the same amino acid.
Wobble position: The third base in a codon is less critical for specifying the amino acid.
tRNA Structure and Function
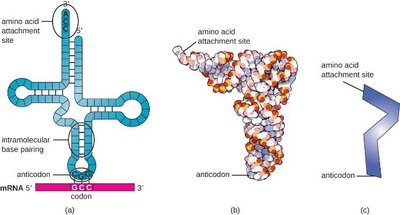
tRNA molecules have a cloverleaf structure with an anticodon that pairs with the mRNA codon and an amino acid attachment site. tRNAs are responsible for translating the genetic code into a sequence of amino acids during protein synthesis.
Anticodon: Triplet of bases complementary to the mRNA codon.
Amino acid attachment site: Binds the specific amino acid corresponding to the anticodon.
Translation
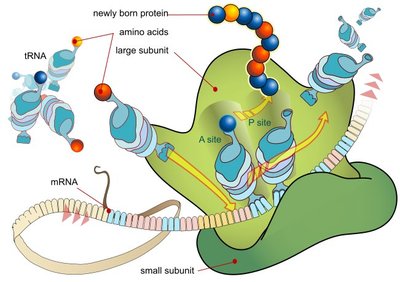
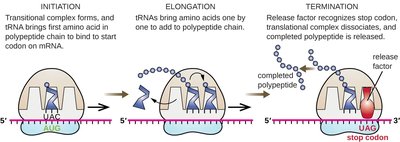
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. It occurs in three stages: initiation, elongation, and termination.
Initiation: The ribosome assembles around the start codon on the mRNA, and the first tRNA binds.
Elongation: tRNAs bring amino acids to the ribosome, where they are joined to form a polypeptide chain.
Termination: The process ends when a stop codon is reached, releasing the completed protein.
Prokaryotes vs. Eukaryotes: Gene Expression
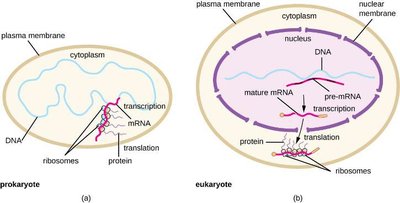
In prokaryotes, transcription and translation occur simultaneously in the cytoplasm, allowing rapid responses to environmental changes. In eukaryotes, transcription occurs in the nucleus and translation in the cytoplasm, with additional RNA processing steps.
Prokaryotes: Coupled transcription and translation.
Eukaryotes: Spatial and temporal separation of transcription and translation; mRNA processing required.
Mutations
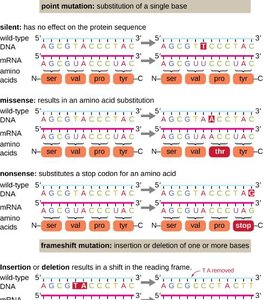
Types of Mutations
Mutations are changes in the DNA sequence that can arise spontaneously or be induced. They can affect protein function in various ways.
Point mutation: Substitution of a single base.
Frameshift mutation: Insertion or deletion of bases, altering the reading frame.
Silent mutation: No effect on protein sequence.
Missense mutation: Changes one amino acid in the protein.
Nonsense mutation: Introduces a stop codon, truncating the protein.
Clinical Focus: E. coli and Horizontal Gene Transfer
Pathogenic E. coli Strains
Escherichia coli is a Gram-negative rod commonly found in the colon. While most strains are harmless, some are pathogenic and cause diarrheal diseases, often associated with contaminated food or water.
ETEC (Enterotoxigenic E. coli): Causes traveler's diarrhea; produces heat-stable and heat-labile toxins.
EIEC (Enteroinvasive E. coli): Invades intestinal cells, causing inflammation.
EPEC (Enteropathogenic E. coli): Causes severe diarrhea, especially in infants.
EHEC (Enterohemorrhagic E. coli): Produces Shiga toxin; can cause bloody diarrhea and hemolytic uremic syndrome.
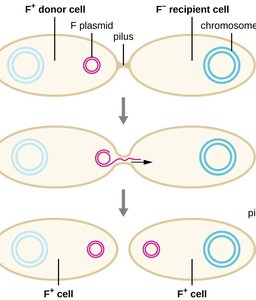
Horizontal Gene Transfer in Bacteria
Bacteria can acquire new genetic material through horizontal gene transfer, which contributes to the spread of virulence factors and antibiotic resistance.
Transformation: Uptake of free DNA from the environment.
Transduction: Transfer of DNA by bacteriophages.
Conjugation: Direct transfer of DNA between cells via a pilus.
Microbial Infections: Skin and Soft Tissue
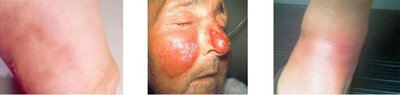
Streptococcus pyogenes Skin Conditions
Streptococcus pyogenes is a Gram-positive coccus that can cause various skin infections, including cellulitis, erysipelas, and erythema nodosum. Its virulence factors enable it to invade tissues and evade the immune system.
Virulence factors: Hyaluronidase, streptokinase, streptolysins, capsule, M proteins.
Necrotizing Fasciitis
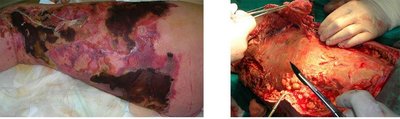
Necrotizing fasciitis is a severe soft tissue infection, often caused by S. pyogenes (Group A Strep), but also by other bacteria. It requires prompt surgical and antibiotic treatment due to rapid tissue destruction.
Symptoms: Rapidly spreading tissue necrosis, severe pain, systemic illness.
Treatment: Debridement or amputation, intravenous antibiotics.