 Back
BackBiotechnology and Recombinant DNA: Principles and Applications
Study Guide - Smart Notes

Biotechnology and Recombinant DNA
Introduction to Biotechnology
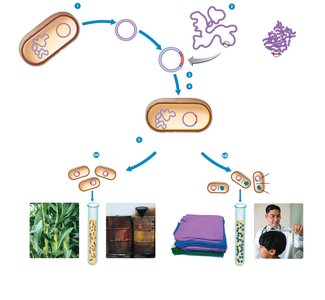
Biotechnology involves the use of microorganisms, cells, or cell components to produce useful products such as foods, antibiotics, vitamins, and enzymes. Recombinant DNA (rDNA) technology refers to the insertion or modification of genes to produce desired proteins, enabling the creation of genetically modified organisms for various applications.
Biotechnology: Utilizes biological systems for industrial and medical purposes.
Recombinant DNA technology: Alters genetic material to express new traits or produce proteins.
Applications: Production of pest-resistant plants, degradative enzymes for waste cleanup, industrial enzymes, and therapeutic proteins.
Example: Human growth hormone produced by recombinant bacteria treats stunted growth.
Selection and Mutation in Biotechnology
Selection and Mutagenesis
Selection involves culturing microbes that naturally produce desired products, while mutation uses mutagens to induce genetic changes that may result in beneficial traits. Site-directed mutagenesis allows for targeted changes in DNA to alter protein function.
Selection: Isolating and culturing microbes with desired traits.
Mutation: Inducing genetic changes to enhance microbial capabilities.
Site-directed mutagenesis: Precisely alters DNA sequences to modify protein products.
Restriction Enzymes and DNA Fragmentation
Restriction Enzymes
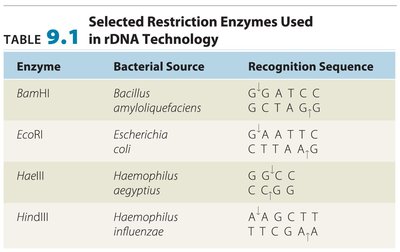
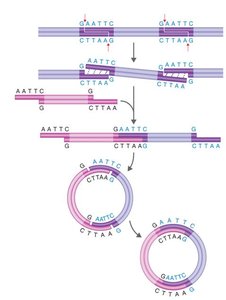
Restriction enzymes are proteins that cut DNA at specific sequences, producing fragments with 'sticky ends' that can be joined to other DNA fragments. This is a fundamental step in creating recombinant DNA.
Recognition sites: Specific DNA sequences where restriction enzymes cut.
Sticky ends: Overhanging sequences that facilitate joining of DNA fragments.
DNA ligase: Enzyme that seals the backbone, forming stable recombinant DNA.
Vectors in Recombinant DNA Technology
Plasmids and Viruses as Vectors
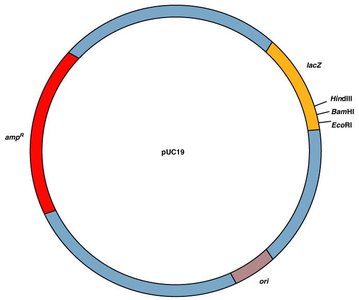
Vectors are DNA molecules used to transfer genes into host cells. Plasmids and viruses are common vectors, often carrying marker genes to identify successful gene uptake.
Marker genes: Enable selection of cells that have incorporated the desired gene (e.g., antibiotic resistance).
Plasmids: Circular DNA molecules used extensively in bacterial transformation.
Viruses: Can deliver genes to eukaryotic cells.
Example: Ampicillin resistance gene used to select bacteria that also produce human insulin.
Polymerase Chain Reaction (PCR)
Amplification of DNA
PCR is a technique developed to amplify specific DNA segments billions of times, enabling direct visualization and analysis. It is used for genetic diagnostics, forensic analysis, and research.
Steps: Denaturation (94°C), annealing (60°C), extension (72°C).
Components: Primers, nucleotides (dNTPs), DNA polymerase.
Applications: Prenatal diagnosis, detection of pathogens, DNA fingerprinting.
DNA Fingerprinting and Southern Blotting
Identification Using DNA Patterns
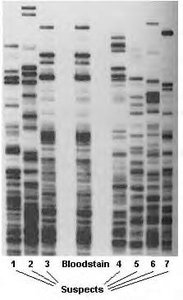
DNA fingerprinting relies on unique patterns of base pair sequences to identify individuals or organisms. Southern blotting is a method to detect specific DNA fragments using labeled probes.
Restriction enzymes: Cut DNA into fragments for analysis.
Gel electrophoresis: Separates DNA fragments by size.
Probes: Hybridize to specific sequences, enabling identification.
Genomic Libraries
Storage and Cloning of Genomic DNA
Genomic libraries are collections of DNA fragments from an organism's genome, stored in plasmids or phages for cloning and analysis.
Plasmid library: DNA fragments cloned into plasmids.
Phage library: DNA fragments cloned into bacteriophage vectors.
Applications: Gene discovery, functional studies.
cDNA and Expression of Eukaryotic Genes in Bacteria
Complementary DNA (cDNA)
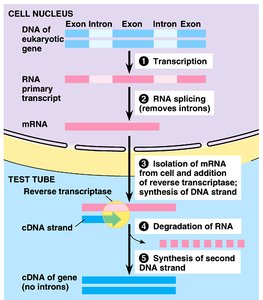
cDNA is synthesized from mRNA using reverse transcriptase, allowing expression of eukaryotic genes (without introns) in bacterial systems. This is crucial for producing eukaryotic proteins in prokaryotes.
Introns: Non-coding regions removed during mRNA processing.
Reverse transcriptase: Enzyme that synthesizes cDNA from mRNA.
Applications: Measuring gene expression, cloning eukaryotic genes.
Blue-White Screening Method
Selection of Recombinant Bacteria
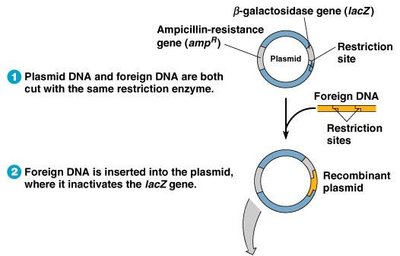
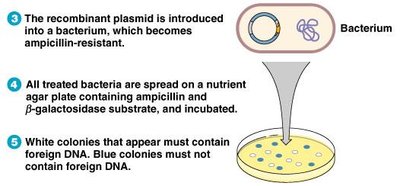
The blue-white screening method uses plasmids with ampicillin resistance and β-galactosidase genes. Insertion of foreign DNA disrupts β-galactosidase, allowing identification of recombinant colonies.
White colonies: Contain foreign DNA, cannot metabolize X-gal.
Blue colonies: Do not contain foreign DNA, metabolize X-gal.
Selection: Only bacteria resistant to ampicillin and unable to produce β-galactosidase are selected.
Radioactive Probes and Colony Hybridization
Identification of Specific Genes
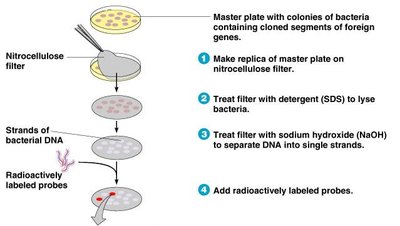
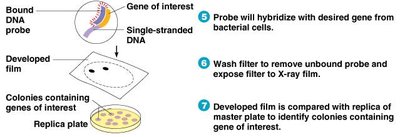
Radioactive or fluorescent probes are used to identify colonies containing specific genes. Probes hybridize to complementary DNA sequences, and visualization techniques reveal positive colonies.
Probes: Short, single-stranded DNA matching the gene of interest.
Hybridization: Formation of double-stranded DNA at the target site.
Visualization: Radioactive or fluorescent labeling enables detection.
Gene Expression Analysis
Microarrays and Measuring mRNA Levels
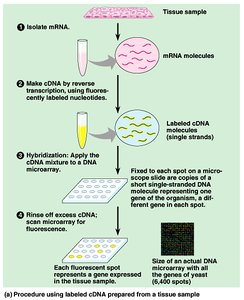
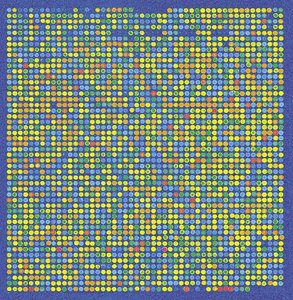
Gene expression can be analyzed using microarrays, which allow simultaneous measurement of thousands of genes. Labeled cDNA hybridizes to DNA spots on a chip, and fluorescence indicates expression levels.
Microarrays: Chips with DNA from different genes attached at each spot.
Fluorescence: Indicates relative mRNA abundance for each gene.
Applications: Comparative gene expression studies, disease profiling.
Therapeutic Applications and Forensic Microbiology
Medical and Forensic Uses
Recombinant DNA technology enables production of human enzymes, subunit vaccines, and gene therapy. PCR and real-time PCR are used in forensic microbiology for organism identification and quantification.
DNA vaccines: Use nonpathogenic viruses carrying pathogen genes.
Gene therapy: Replaces defective or missing genes.
Forensic PCR: Detects specific organisms or genetic markers.
CRISPR-Cas9 Genome Editing
Mechanism and Applications
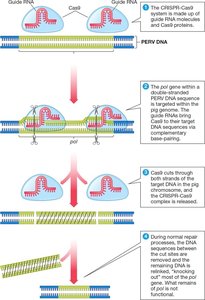
The CRISPR-Cas9 system is a bacterial defense mechanism adapted for genome editing. Cas9 proteins, guided by RNA, cut DNA at specific sites, enabling targeted gene modification. This technology is used for gene deletion, disease modeling, and organ compatibility enhancement.
Guide RNA: Directs Cas9 to the target DNA sequence.
Cas9 protein: Cuts both DNA strands at the target site.
Applications: Removal of PERVs from pig genomes for safer organ transplantation.
Enzyme | Bacterial Source | Recognition Sequence |
|---|---|---|
BamHI | Bacillus amyloliquefaciens | G\GATCC GCTAG\G |
EcoRI | Escherichia coli | G\AATTC CTTAA\G |
HaeIII | Haemophilus aegyptius | GG\CC CC\GG |
HindIII | Haemophilus influenzae | A\AGCTT TTCGA\A |
