 Back
BackCellular Inclusions, Surface Structures, and Chemotaxis in Bacteria
Study Guide - Smart Notes

Cellular Inclusions in Bacteria
Overview of Cellular Inclusions
Cellular inclusions are insoluble intracellular structures found in many bacteria, serving as storage sites for nutrients and minerals. These inclusions play a critical role in cell survival, energy management, and orientation within the environment.
Nutrient Storage: Inclusions store essential nutrients such as carbon, phosphate, and sulfur in forms that can be readily accessed when needed.
Energy Storage: Some inclusions act as energy reserves, allowing bacteria to survive periods of nutrient scarcity.
Orientation and Taxis: Certain inclusions help bacteria orient themselves in response to environmental cues.
Cell Survival/Dormancy: Inclusions can aid in dormancy, helping bacteria withstand adverse conditions.

Types of Inclusion Bodies
Bacteria store various nutrients in inclusion bodies, each with specialized functions and chemical compositions.
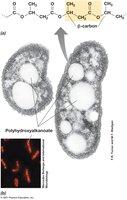
Carbon Storage: Bacteria store carbon in insoluble forms such as polyhydroxyalkanoates (PHAs), cellulose, or glycogen. These compounds can be metabolized to provide energy and carbon skeletons for biosynthesis.
Polyhydroxybutyrate (PHB): A common PHA, PHB is synthesized from acetyl-CoA and serves as a readily accessible carbon and energy source.
Function: Carbon sources stored in inclusions can be oxidized to CO2, producing NADH and ATP via cellular respiration.
Other Nutrient Storage Inclusions
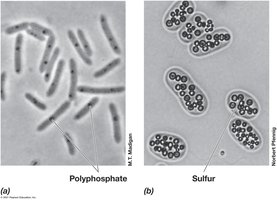
Bacteria also store essential elements such as phosphorus and sulfur in specialized inclusions.
Polyphosphate Granules: Polymers of phosphate used for nucleic acid and phospholipid synthesis.
Sulfur Granules: Mineralized sulfur stored for use as an electron acceptor in metabolic processes.
Trace Metals: Some bacteria store trace metals in mineralized forms for later use.
Surface Structures: Bacterial Flagella
Structure and Function of Flagella
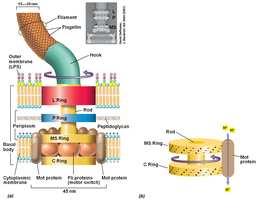
Bacterial flagella are extracellular protein appendages primarily responsible for motility. They function as proton-driven molecular motors, rotating to propel the cell through its environment.
Flagellum Components: The flagellum consists of a filament, hook, and basal body (including central rod and rings).
Proton-Driven Motor: The rotation of the flagellum is powered by the proton motive force, similar to ATP synthase.
Chemotaxis: The direction of flagellar rotation changes in response to chemical signals, allowing bacteria to move toward or away from stimuli.
Flagellar Motor Mechanism
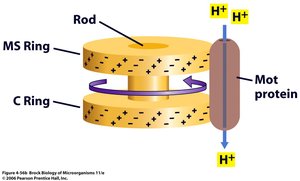
The flagellar motor operates as a rotary engine, with Mot proteins acting as the stator and rings/rod as the rotor. The switch complex (C ring and Fli proteins) controls the direction of rotation.
Proton Turbine Model: Protons flow through Mot proteins, generating torque that rotates the flagellum.
Switching Rotation: The motor can switch between clockwise (CW) and counter-clockwise (CCW) rotation, affecting motility patterns.
Flagellar Arrangements
Bacteria exhibit various flagellar arrangements, which influence their motility strategies.
Monotrichous: Single flagellum at one end.
Lophotrichous: Multiple flagella at one or both ends.
Amphitrichous: Single flagellum at both ends.
Peritrichous: Flagella distributed over the entire cell surface.
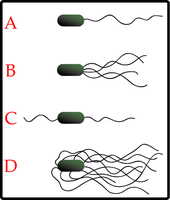
Flagellar Motility Mechanism
Peritrichous flagella form bundles that act as corkscrew propellers. The direction of rotation determines the movement pattern:
Counter-Clockwise (CCW): Flagella bundle, cell moves forward (run).
Clockwise (CW): Flagella splay, cell tumbles and changes direction.
Motility and Chemotaxis
Chemotaxis: Directed Movement
Chemotaxis is the directed movement of bacteria in response to chemical gradients. It involves a network of sensory and response proteins that regulate flagellar rotation.
Model System: Peritrichous Escherichia coli is commonly studied.
Mechanism: In the absence of attractant, bacteria exhibit random run/tumble behavior. In the presence of attractant, movement becomes biased toward the source.
Chemoreceptors: Bacteria sense chemical gradients over time, not space.
Other Stimuli: Chemotaxis can also respond to light, oxygen, osmolarity, etc.
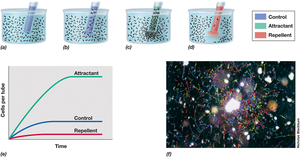
Measuring Chemotaxis: Capillary Tube Assay
The capillary tube assay is used to determine whether a compound is an attractant or repellant for bacteria.
Method: A capillary tube containing the test substance is placed in a bacterial suspension. Bacteria migrate into the tube if the substance is an attractant, avoid it if it is a repellant, or show neutral behavior if indifferent.
Analysis: The number of bacteria entering each tube is counted and compared to controls.
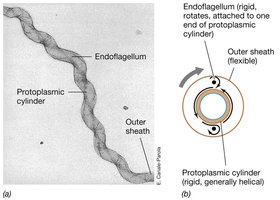
Spirochetes: Flagella and Cell Shape
Spirochetes, such as Treponema pallidum, possess periplasmic flagella that contribute to their unique corkscrew shape and motility.
Cell Shape: Long, helical (corkscrew) morphology.
Motility: Periplasmic flagella enable spirochetes to move in a corkscrew fashion, aiding in tissue penetration and pathogenicity.
Transport Across the Membrane
Importance of Membrane Transport
Transport systems are essential for nutrient uptake, waste removal, and maintaining cellular homeostasis. Bacteria utilize several energy-driven mechanisms to transport solutes against concentration gradients.
Classes of Transport Systems
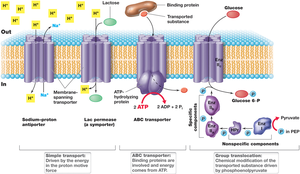
Simple Transport: Utilizes transmembrane proteins and is driven by the proton motive force. Includes symport (solute and proton co-transported in the same direction) and antiport (solute and proton transported in opposite directions).
Group Translocation: The transported substance is chemically modified during transport. Driven by energy-rich organic compounds, not the proton motive force.
ABC Transporter Systems: ATP-binding cassette transporters use ATP hydrolysis to drive the uptake of a wide range of substrates. Substrate-binding proteins outside the cell have high affinity for solutes.
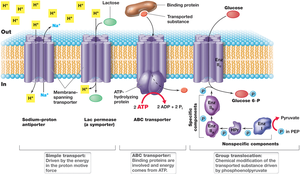
Simple Transport
Simple transport systems use the proton motive force to move solutes across the membrane. Examples include the E. coli lac permease and phosphate/sulfate transporters.
Group Translocation
In group translocation, the transported molecule is chemically modified, often phosphorylated, as it enters the cell. The best-studied system is the phosphotransferase system for glucose uptake.
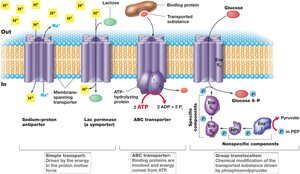
ABC Transporter Systems
ABC transporters are widespread in bacteria, with over 200 systems identified for various organic and inorganic compounds. These systems use ATP hydrolysis to transport solutes against their concentration gradients.
Summary Table: Types of Bacterial Transport Systems
Transport System | Energy Source | Mechanism | Example |
|---|---|---|---|
Simple Transport | Proton Motive Force | Symport/Antiport | Lac permease, phosphate transport |
Group Translocation | Energy-rich organic compound | Chemical modification of solute | Phosphotransferase system (glucose) |
ABC Transporter | ATP hydrolysis | Substrate-binding protein, transmembrane transporter | Various sugars, amino acids, ions |
Key Equations
Proton Motive Force:
Where is the proton motive force, is the membrane potential, is the gas constant, is temperature, is Faraday's constant, and is the pH gradient across the membrane.
ATP Hydrolysis (ABC Transporters):
ATP hydrolysis provides the energy for substrate transport in ABC systems.
