Back
BackCellular Structure and Function in Microbiology: Prokaryotic and Eukaryotic Cells Unit 2
Study Guide - Smart Notes

Cellular Structure and Function
Introduction
This unit explores the fundamental structures and functions of prokaryotic and eukaryotic cells, focusing on their morphologies, membranes, cell walls, surface structures, inclusions, motility mechanisms, and the endosymbiotic theory. Understanding these features is essential for appreciating microbial diversity and physiology.
Cell Size and Morphology
Surface Area to Volume Ratio
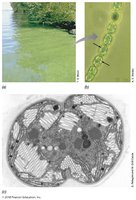
The small size of microbial cells results in a high surface area-to-volume ratio, which facilitates efficient nutrient uptake and waste elimination. This ratio is a key factor in determining the metabolic rate and growth potential of microorganisms.
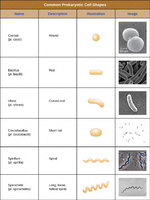
Common Prokaryotic Cell Shapes
Coccus: Spherical shape
Bacillus: Rod-shaped
Vibrio: Curved rod
Coccobacillus: Short rod
Spirillum: Spiral
Spirochete: Long, loose spiral
The Cytoplasmic (Plasma) Membrane
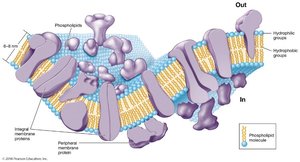
Structure: The Fluid Mosaic Model
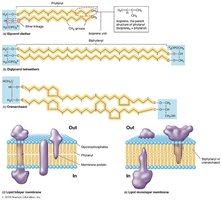
The cytoplasmic membrane is a phospholipid bilayer with embedded proteins, described by the fluid mosaic model. Phospholipids are amphipathic, containing hydrophilic heads and hydrophobic tails. Proteins may be integral (spanning the membrane) or peripheral (attached to the surface).
Bacterial Plasma Membranes
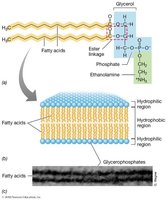
Phospholipids contain ester linkages (like eukaryotes).
Some bacteria have hopanoids (sterol-like molecules) for membrane stability.
Archaeal Plasma Membranes
Phospholipids have ether linkages (not ester).
Contain isoprene units instead of fatty acids.
May form bilayers (phytanyl) or monolayers (biphytanyl).
Lack hopanoids and sterols.
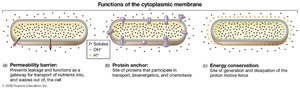
Functions of the Cytoplasmic Membrane
Selective permeability barrier: Controls entry and exit of substances.
Protein anchor: Site for proteins involved in transport, bioenergetics, and chemotaxis.
Energy conservation: Site of generation and use of the proton motive force.
Cell Walls
General Features
Peptidoglycan is unique to Bacteria.
Provides shape, rigidity, and protection against osmotic lysis.
Some Archaea and Eukarya have cell walls, but their composition differs.
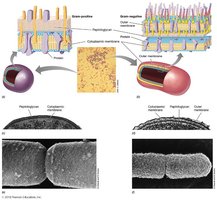
Gram-Positive vs. Gram-Negative Bacteria
Gram-Positive: Thick, multi-layered peptidoglycan; contains teichoic acids.
Gram-Negative: Thin peptidoglycan layer; outer membrane with lipopolysaccharide (LPS).
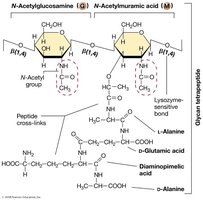
Peptidoglycan Structure
Composed of alternating N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM) sugars.
Cross-linked by short peptides containing unusual amino acids (e.g., D-alanine, D-glutamic acid).
Provides rigidity and resistance to osmotic pressure.
Gram-Positive Cell Wall Specifics
Up to 90% peptidoglycan.
Teichoic acids bind divalent metal ions and are covalently linked to peptidoglycan.
Lipoteichoic acids are anchored in the membrane.
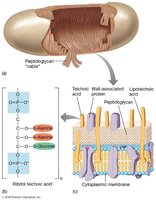
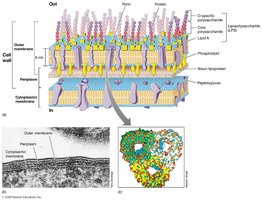
Gram-Negative Cell Wall Specifics
Outer membrane contains LPS (lipopolysaccharide).
Acts as a barrier to many antibiotics and harmful agents.
Contains porins (protein channels), periplasmic space, and Braun lipoproteins (anchor outer membrane).
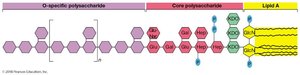
Lipopolysaccharide (LPS) Structure
O-specific polysaccharide: Variable, species-specific.
Core polysaccharide: Contains KDO and heptoses.
Lipid A: Endotoxin, responsible for toxicity in animals.
Archaeal Cell Walls
No peptidoglycan; instead, may have pseudomurein, S-layers (protein/glycoprotein), or polysaccharide walls.
S-layers are paracrystalline and often the outermost layer.
Cell Surface Structures and Inclusions
Fimbriae, Pili, and Hami
Fimbriae: Short, numerous; for attachment and biofilm formation.
Pili: Longer, fewer; for conjugation, attachment, and twitching motility.
Hami: Archaeal structures resembling grappling hooks for attachment.
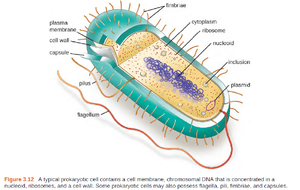
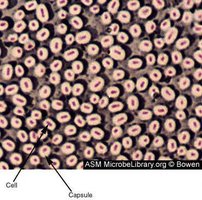
Glycocalyces (Capsules and Slime Layers)
Capsule: Organized, firmly attached; prevents phagocytosis, increases virulence.
Slime layer: Unorganized, loose; aids in biofilm formation.
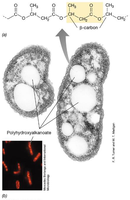
Cell Inclusions and Gas Vesicles
Inclusions: Storage of nutrients (e.g., polyphosphate, PHA, sulfur granules).
Gas vesicles: Protein-bound structures for buoyancy in aquatic microbes.
Endospores
Structure and Function
Highly resistant, dormant cells produced by some Gram-positive bacteria (e.g., Bacillus, Clostridium).
Resistant to heat, desiccation, UV, and chemicals.
Formed by sporulation under adverse conditions; germinate when conditions improve.
Motility in Prokaryotes
Flagellar Motility
Flagella: Helical, rotating appendages for swimming.
Arrangements: monotrichous, amphitrichous, lophotrichous, peritrichous.
Structure: basal body (motor), hook (connector), filament (flagellin).
Movement: "runs" and "tumbles"; powered by proton motive force.
Archaeal Motility (Archaella)
Structurally distinct from bacterial flagella; powered by ATP hydrolysis.
Composed of several different proteins; thinner than bacterial flagella.
Gliding Motility
Found only in Bacteria; requires solid surfaces.
Mechanisms: polysaccharide slime, type IV pili, gliding-specific proteins.
Eukaryotic Cells and the Endosymbiotic Theory
Major Eukaryotic Cell Structures
Nucleus: Contains DNA, surrounded by nuclear envelope.
Organelles: Mitochondria, chloroplasts, endoplasmic reticulum, Golgi apparatus, etc.
Cytoskeleton: Provides structure and facilitates movement.
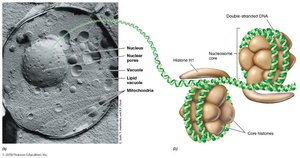
Endosymbiotic Theory
The endosymbiotic theory proposes that mitochondria, hydrogenosomes, and chloroplasts originated from free-living prokaryotes engulfed by ancestral eukaryotic cells. Evidence includes:
Genetic similarity to bacteria (e.g., rRNA sequences, circular DNA).
Binary fission division.
Presence of 70S ribosomes (like bacteria).
Mitochondria
Site of aerobic respiration and ATP production.
Double-membraned; contains cristae and matrix with enzymes for the citric acid cycle.
Hydrogenosomes
Found in anaerobic eukaryotes; produce H2, CO2, and acetate from pyruvate.
Lack TCA cycle enzymes and cristae.
Chloroplasts
Present in phototrophic eukaryotes; site of photosynthesis.
Double-membraned; contain thylakoids and stroma with RubisCO enzyme.