 Back
BackClassification and Identification of Microorganisms
Study Guide - Smart Notes

Classification of Microorganisms
Introduction to Taxonomy
Taxonomy is the science of classification, particularly of living organisms. It provides a systematic framework for naming, identifying, and organizing organisms based on shared characteristics and evolutionary relationships. The goals of taxonomy include classifying organisms, establishing relationships, and providing a universal language for scientific communication.
Definition: Taxonomy is derived from the Greek for "orderly arrangement."
Purpose: To differentiate organisms, establish relationships, and provide reference for identification.
Application: Used in clinical settings to identify pathogens and guide treatment.
Systematics and Phylogeny
Systematics, or phylogeny, is the study of the evolutionary history and relationships among organisms. It uses genetic and molecular data to construct phylogenetic trees that reflect evolutionary pathways.
Tools: DNA/RNA sequencing, protein sequence comparisons.
Goal: To understand evolutionary connections and ancestry.
Example: Bacillus subtilis is more closely related to Listeria monocytogenes than to Staphylococcus aureus.
Taxonomy vs. Phylogeny
Feature | Taxonomy | Phylogeny |
|---|---|---|
Focus | Classification and naming | Evolutionary relationships |
Purpose | Organize and identify organisms | Understand ancestry and lineage |
Based On | Shared traits (morphological, biochemical, genetic) | Genetic and evolutionary data |
End Product | Hierarchical classification (Domain → Species) | Phylogenetic tree |
Example Output | Escherichia coli (classified organism) | Tree showing E. coli relatedness to Salmonella |
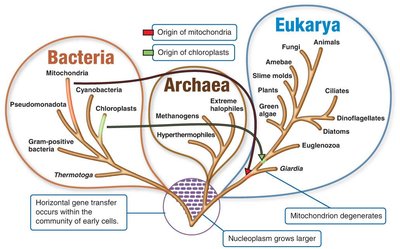
The Three Domains of Life
The three-domain system, developed by Carl Woese in 1978, is based on rRNA nucleotide sequences and divides all life into:
Bacteria: Includes typical bacteria, cyanobacteria, and others.
Archaea: Includes methanogens, extreme halophiles, and hyperthermophiles.
Eukarya: Includes animals, plants, fungi, and protists.
Scientific Nomenclature
Scientific names provide a standardized system for naming organisms, reducing confusion caused by regional or linguistic differences. Binomial nomenclature assigns each organism a two-part name: the genus and the specific epithet (species).
Example: Escherichia coli
Universality: Used worldwide for consistency and accuracy.
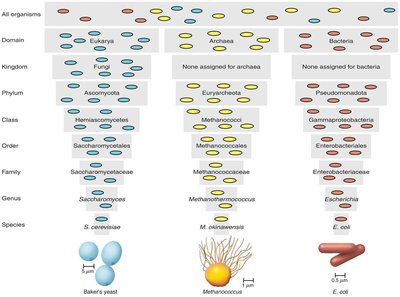
The Taxonomic Hierarchy
The taxonomic hierarchy organizes organisms into nested categories, from the most general to the most specific. For prokaryotes, the hierarchy is outlined in Bergey’s Manual of Systematics of Archaea and Bacteria.
Domain
Kingdom (not used for Bacteria and Archaea)
Phylum
Class
Order
Family
Genus
Species
Strain (for prokaryotes) or subspecies
Classification of Viruses
Unique Nature of Viruses
Viruses are not classified within the three domains because they are not composed of cells and require a host cell for replication. Viral species are defined as populations of viruses with similar characteristics, distinguishable by morphology, genome type, enzymes, and ecological niche.
Not cellular: Viruses lack cellular structure.
Classification: Based on multiple laboratory and genetic methods.
Methods of Classifying and Identifying Microorganisms
Overview of Identification Methods
Microorganisms are identified using a variety of laboratory techniques, which are essential for diagnosis and treatment in clinical settings. Bergey’s Manual of Determinative Bacteriology is a key reference for bacterial identification.
Criteria: Morphology, cell wall composition, differential staining, oxygen requirements, biochemical testing.
Clinical application: Specimens are collected, transported, and analyzed to guide treatment.


Conventional Identification Methods
Morphology: Useful for eukaryotes; limited for prokaryotes. Presence of endospores or flagella can be informative.
Differential staining: Gram staining and acid-fast staining are common but not useful for all bacteria or archaea.
Biochemical tests: Detect the presence of specific bacterial enzymes and metabolic pathways.
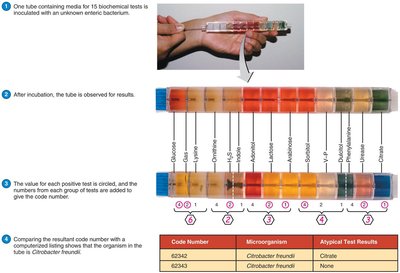
Biochemical Tests and Rapid Identification
Biochemical tests are widely used to differentiate bacteria based on enzymatic activities. Rapid identification methods, such as the EnteroPluri Test, allow simultaneous testing of multiple biochemical reactions, providing results within 4 to 24 hours. These methods are sometimes called numerical identification systems.
Limitation: Mutations and plasmid acquisition can alter test results, requiring multiple tests for accuracy.
Numerical identification: Each test result is assigned a value, and the sum is compared to a database.
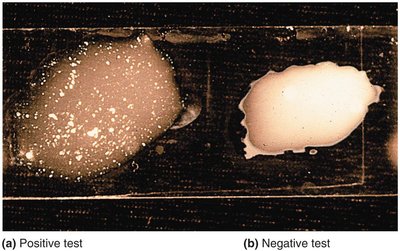
Serology
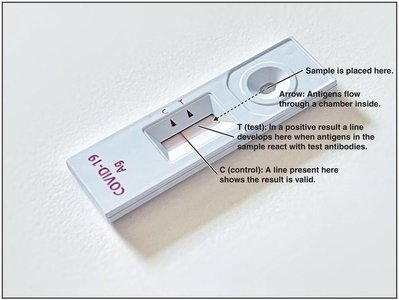
Serology is the study of serum and immune responses. Microorganisms are antigenic and stimulate antibody production. Serological tests use known antibodies (antisera) to detect specific antigens in unknown samples, allowing differentiation between species and strains.
Slide agglutination test: Bacteria clump when mixed with specific antibodies, indicating a positive reaction.
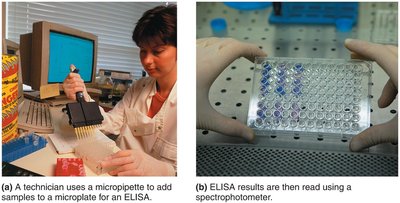
ELISA (Enzyme-Linked Immunosorbent Assay): Used for detecting antigens (direct ELISA) or antibodies (indirect ELISA) in patient samples.


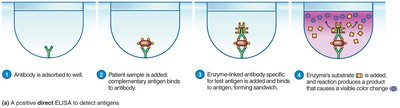
Putting Classification Methods Together
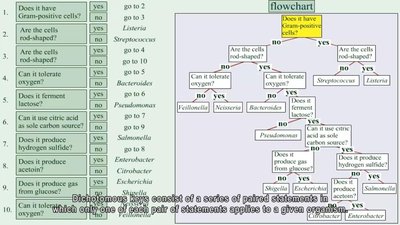
Dichotomous Keys
Dichotomous keys are identification tools based on a series of paired statements or questions, each with two possible answers. By following the sequence of choices, users can identify unknown organisms efficiently.
Structure: Each step narrows down the possibilities until a final identification is reached.
Application: Widely used in microbiology for bacterial identification.
Summary Table: Main Methods for Microbial Identification
Method | Principle | Application |
|---|---|---|
Morphology | Shape, arrangement, staining | Initial screening, eukaryotes |
Differential Staining | Cell wall properties | Gram/acid-fast bacteria |
Biochemical Tests | Enzyme/metabolic activity | Bacterial species/strain ID |
Serology | Antigen-antibody reactions | Species/strain differentiation |
Rapid Identification | Simultaneous biochemical tests | Clinical diagnostics |
Dichotomous Keys | Stepwise paired choices | Systematic identification |
Additional info: Modern microbial taxonomy increasingly relies on molecular methods, such as 16S rRNA sequencing, for precise classification and identification, especially for unculturable organisms.
