 Back
BackClassification and Identification of Microorganisms: Chapter 10 Study Guide
Study Guide - Smart Notes

Classification of Microorganisms
Taxonomy and Phylogeny
Taxonomy is the science of classifying organisms to show their degree of similarity, while systematics (phylogeny) is the study of the evolutionary history of organisms. These fields help organize biological diversity and understand relationships among organisms.
Taxon: A category or group in taxonomy, such as genus or species.
Phylogeny: The evolutionary history and relationships among organisms.
Systematics: The study of phylogeny and classification.
Example: Organisms are grouped based on shared characteristics and evolutionary ancestry.
Historical Classification Systems
Classification systems have evolved from two kingdoms (plants and animals) to three domains, reflecting advances in molecular biology and understanding of evolutionary relationships.
Linnaeus: Developed the binomial nomenclature and taxonomic hierarchy.
Whittaker: Proposed the five-kingdom system.
Woese: Introduced the three-domain system based on rRNA sequences.
Three Domains: Bacteria, Archaea, and Eukarya.
The Three Domains: Bacteria, Archaea, and Eukarya
The three-domain system is based on molecular evidence, particularly rRNA sequences, and reflects fundamental differences among major groups of organisms.
Bacteria: Prokaryotic, cell walls contain peptidoglycan.
Archaea: Prokaryotic, cell walls lack peptidoglycan, often found in extreme environments.
Eukarya: Eukaryotic, includes animals, plants, fungi, and protists.
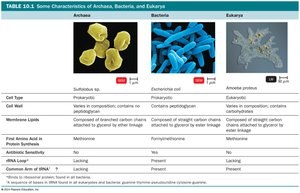
Table Purpose: Comparison of cell type, cell wall, membrane lipids, first amino acid in protein synthesis, antibiotic sensitivity, and presence of common rRNA among the three domains.
Phylogenetic Trees
Phylogenetic trees group organisms according to common properties and evolutionary relationships, using evidence such as fossils and genome sequences.
Common Ancestor: Groups of organisms evolved from a shared ancestor.
rRNA and Genome Sequencing: Used to determine evolutionary relationships among prokaryotes.
Classification of Organisms
Scientific Nomenclature
Scientific names are used to avoid confusion caused by regional and linguistic variations. Binomial nomenclature assigns each organism a genus and species name.
Genus: The first part of the scientific name.
Specific epithet: The second part, indicating the species.
Example: Escherichia coli
Taxonomic Hierarchy
The taxonomic hierarchy is a series of subdivisions developed to classify organisms, from broad to specific categories.
Major Taxa: Domain, Kingdom, Phylum, Class, Order, Family, Genus, Species.
Eukaryotic Species: Groups of closely related organisms that breed among themselves.
Classification of Prokaryotes
Prokaryotes are classified based on genomic similarity, morphology, and biochemical characteristics. Bergey’s Manual is a key resource for prokaryotic classification.
Culture: Bacteria grown in laboratory media.
Clone: Population of cells derived from a single parent cell.
Strain: Genetically different cells within a clone.
Classification of Eukaryotes
Eukaryotes are classified into kingdoms based on cell structure and nutritional modes.
Protista: Mostly unicellular, nutritionally diverse, grouped by rRNA clades.
Fungi: Chemoheterotrophic, cell walls of chitin, develop from spores or hyphal fragments.
Plantae: Multicellular, cellulose cell walls, photosynthetic.
Animalia: Multicellular, no cell walls, ingest organic matter.
Classification of Viruses
Viruses are not classified within the three domains because they are not composed of cells and require a host cell for replication.
Viral Species: Population of viruses with similar characteristics, distinguished by morphology, genome, enzymes, and ecological niche.
Methods of Classifying and Identifying Microorganisms
Classification vs. Identification
Classification involves placing organisms in groups of related species, while identification matches characteristics of an unknown organism to known organisms, often in clinical settings.
Bergey’s Manual: Provides identification schemes for bacteria and archaea.
Criteria: Morphology, cell wall composition, differential staining, oxygen requirements, biochemical testing.
Conventional Identification Methods
Traditional methods include morphological observation, differential staining, and biochemical tests to identify microorganisms.
Morphology: Useful for eukaryotes; presence of endospores or flagella can be informative.
Differential Staining: Gram staining, acid-fast staining; not useful for bacteria without cell walls or archaea.
Biochemical Tests: Determine presence of bacterial enzymes; rapid methods can provide results in 4–24 hours.
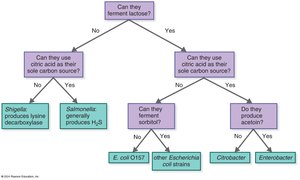
Table Purpose: Dichotomous key for identifying enteric bacteria based on metabolic characteristics.
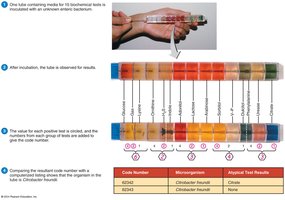
Image Purpose: Shows a rapid identification method using multiple biochemical tests for bacteria.
Serological Methods
Serology studies immune responses in serum. Microorganisms are antigenic and stimulate antibody formation. Serological tests use known antibodies to identify unknown bacteria.
Slide Agglutination Test: Bacteria agglutinate when mixed with specific antibodies.
ELISA: Enzyme-linked immunosorbent assay; direct ELISA detects antigens, indirect ELISA detects antibodies.
Western Blot: Used for confirmation of specific proteins or antibodies.
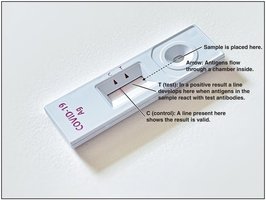

Image Purpose: Example of a direct ELISA test for COVID-19.
Molecular Identification Methods
Molecular approaches provide more accurate classification and identification, using DNA sequencing, DNA fingerprinting, and PCR.
Whole Genome Sequencing: Determines base composition (G+C content) and relatedness.
Nucleic Acid Hybridization: Measures ability of DNA strands from different organisms to hybridize; >70% hybridization indicates same species.
Nucleic Acid Amplification Tests (NAATs): Use PCR to amplify DNA for identification.
Southern Blotting: Uses DNA probes to identify microorganisms.
DNA Chips (Microarrays): Detect pathogens by hybridization and fluorescence.
Ribotyping: Uses rRNA sequencing.
FISH: Fluorescent in situ hybridization stains targeted microorganisms.
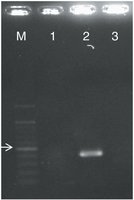
Image Purpose: Gel electrophoresis result showing identification of a coronavirus strain using PCR.
Dichotomous Keys and Cladograms
Dichotomous keys are identification tools based on successive questions with two possible answers. Cladograms are maps showing evolutionary relationships based on molecular data.
Dichotomous Key: Used for stepwise identification of organisms.
Cladogram: Visual representation of evolutionary relationships.
Summary Table: Characteristics of Archaea, Bacteria, and Eukarya
Characteristic | Archaea | Bacteria | Eukarya |
|---|---|---|---|
Cell Type | Prokaryotic | Prokaryotic | Eukaryotic |
Cell Wall | Varies in composition; contains no peptidoglycan | Contains peptidoglycan | Varies; contains carbohydrates |
Membrane Lipids | Composed of branched carbon chains attached to glycerol by ether linkage | Composed of straight carbon chains attached to glycerol by ester linkage | Composed of straight carbon chains attached to glycerol by ester linkage |
First Amino Acid in Protein Synthesis | Methionine | Formylmethionine | Methionine |
Antibiotic Sensitivity | No | Yes | No |
Common rRNA | Present | Present | Present |
Key Equations
DNA Base Composition:
Hybridization Percentage:
