 Back
BackClassification and Taxonomy of Microorganisms
Study Guide - Smart Notes

Classification of Microorganisms
Why Classify Organisms?
Taxonomy is the science of classifying living organisms, providing a systematic framework for understanding relationships, identification, and universal naming. With millions of known and unknown organisms, many of which are microorganisms, classification is essential for scientific communication and research.
Relationships: Classification reveals evolutionary and genetic relationships among organisms.
Identity: Provides a common reference point for identifying organisms.
Universal Naming: Scientific names are consistent across languages, unlike common names.
History of Classification
Early Systems and Linnaean Taxonomy
Classification began with simple plant and animal groupings. Carolus Linnaeus formalized taxonomy in 1735, introducing a hierarchical system and binomial nomenclature (genus and species).
Binomial Nomenclature: Each organism has two names: genus (capitalized) and species (lowercase), e.g., Homo sapiens.
Hierarchical System: Organisms are grouped in nested categories.

Evolution of Classification Systems
Classification systems have evolved to reflect new scientific discoveries:
1857: Bacteria and fungi placed in plant kingdom (Carl von Nageli).
1866: Kingdom Protista proposed for bacteria, protozoa, algae, fungi (Ernst Haeckel).
1959: Fungi assigned their own kingdom.
1969: Five kingdom system introduced (Robert Whittaker).
1998: Three domain system proposed (Carl Woese) based on cell types: Bacteria, Archaea, Eukarya.
Classification Hierarchy and Systematics
Taxonomic Hierarchy
Systematics organizes organisms into a hierarchy of groups called taxa, based on evolutionary relationships (phylogeny). The hierarchy is:
Domain
Kingdom
Phylum
Class
Order
Family
Genus
Species
Mnemonic: "Do Kindly Place Candy Out For Good Students"
Three Domain System
Rooted and Unrooted Phylogenetic Trees
The three-domain system divides life into Bacteria, Archaea, and Eukarya, based on ribosomal RNA and cell structure. Phylogenetic trees illustrate evolutionary relationships among these domains.
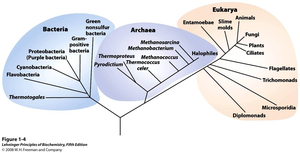
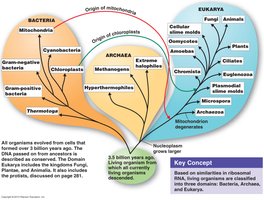
Characteristics of Archaea, Bacteria, and Eukarya
Comparative Features
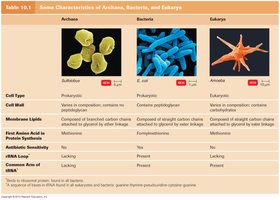
Each domain has distinct cellular characteristics, including cell type, cell wall composition, membrane lipids, and sensitivity to antibiotics.
Feature | Archaea | Bacteria | Eukarya |
|---|---|---|---|
Cell Type | Prokaryotic | Prokaryotic | Eukaryotic |
Cell Wall | Varies; no peptidoglycan | Contains peptidoglycan | Varies; no peptidoglycan |
Membrane Lipids | Branched hydrocarbons, ether linkage | Unbranched hydrocarbons, ester linkage | Unbranched hydrocarbons, ester linkage |
First Amino Acid in Protein Synthesis | Met | Formylmethionine | Met |
Antibiotic Sensitivity | Not sensitive | Sensitive | Not sensitive |
rRNA Loop | Lacking | Present | Lacking |
Common Area of rRNA | Present | Present | Present |
Nomenclature and Taxonomic Hierarchy
Scientific Naming Conventions
Organisms are named using binomial nomenclature. The genus is capitalized, the species is lowercase, and both are italicized or underlined. Abbreviations are used after the first mention if unambiguous (e.g., E. coli).
Strains: Variants within a species are called strains, identified by numbers or letters (e.g., E. coli K-12, E. coli O157:H7).
Bergey’s Manual: Classification scheme for bacteria is detailed in Bergey’s Manual of Systematic Bacteriology.

Classification Methods for Prokaryotes
Phenotypic and Genotypic Methods
Prokaryotes are classified using a variety of methods:
Morphological Characteristics: Cell shape, arrangement, and structure.
Differential Staining: Techniques such as Gram stain, acid-fast stain, and endospore stain.
Biochemical Tests: Metabolic capabilities and enzyme activities.
Serology: Identification based on antigen-antibody reactions (serotypes, serovars, biovars).
Phage Typing: Susceptibility to specific bacteriophages.
Fatty Acid Profiles: Analysis of cellular fatty acids.
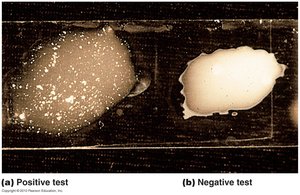
Molecular Methods
DNA Base Composition: G+C content analysis (especially in Gram-positive bacteria).
DNA Fingerprinting: Patterns of DNA fragments for identification.
PCR: Amplification of specific DNA sequences.
rRNA Gene Sequencing: Comparison of ribosomal RNA genes.
DNA Chips: Microarray technology for genetic analysis.

Viruses, Viroids, and Prions
Classification of Viruses
Viruses are not included in the three-domain system because they lack cellular structure and depend on host cells for replication. Viral species are defined by shared characteristics and ecological niche.
Nucleic Acid: Viruses contain either DNA or RNA.
Protein Coat: Surrounds the nucleic acid; may have an envelope.
Replication: Occurs inside host cells using cellular machinery.
Taxonomy: ICTV classifies viruses by nucleic acid type, replication strategy, and morphology.
Naming: Genus names end in -virus, family names in -viridae, order names in -ales. No specific epithet.
Examples: Family Herpesviridae, genus Simplexvirus, human herpesvirus 2.
Viroids and Prions
Viroids and prions are unique infectious agents not classified in traditional systems:
Viroids: Infectious RNA molecules causing plant diseases.
Prions: Infectious protein particles, responsible for neurodegenerative diseases.
Additional info:
Classification is foundational for understanding microbial diversity, evolution, and disease.
Modern taxonomy increasingly relies on molecular and genetic data for accurate classification.
