 Back
BackEnterobacteriaceae and Pseudomonas: Identification and Differentiation in Clinical Microbiology
Study Guide - Smart Notes

Enterobacteriaceae and Pseudomonas
Background and General Characteristics
The Enterobacteriaceae (enterics) and Pseudomonas are important groups of Gram-negative rods relevant to clinical microbiology. Enterics are normal inhabitants of the intestines in humans and animals, while Pseudomonas species are primarily free-living in soil and water. Both groups include significant human pathogens.
Enterobacteriaceae: Gram-negative rods, facultative anaerobes, ferment glucose, may or may not ferment lactose.
Pseudomonas: Gram-negative rods, oxidase positive, primarily non-fermenters, notable for environmental resilience and opportunistic infections.
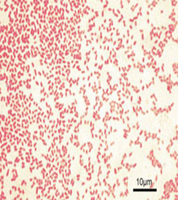
Normal Flora vs. Pathogenic Enterics
Some enterics are part of the normal intestinal flora, while others are pathogenic. The ability to ferment lactose is a key differentiator:
Lactose fermenters (Normal Flora): Escherichia coli
Lactose non-fermenters (Pathogens): Salmonella spp., Shigella spp., Vibrio cholerae
Pathogenic Enterics
Salmonella: Causes enteric fever (typhoid), gastroenteritis, and septicemia. Transmission via contaminated water, food (especially eggs), or reptiles (e.g., turtles).
Shigella: Infects humans and primates, causing symptoms from mild diarrhea to severe bacillary dysentery.
Vibrio: Vibrio cholerae causes cholera, characterized by severe watery diarrhea ("rice water" stools). Vibrio parahaemolyticus is associated with seafood-borne gastroenteritis.
Isolation and Differentiation of Enteric Bacteria
Selective and Differential Media
Isolation and identification of enteric bacteria rely on selective and differential media that exploit differences in lactose fermentation and Gram reaction.
EMB (Eosin Methylene Blue) Agar
Selective: Inhibits Gram-positive bacteria via eosin and methylene blue dyes.
Differential: Distinguishes lactose fermenters (colored colonies) from non-fermenters (colorless colonies).
Results: E. coli produces a green metallic sheen; Enterobacter aerogenes forms pink colonies with dark centers.
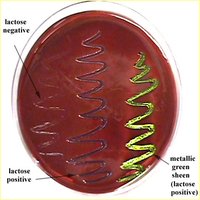
MacConkey Agar
Selective: Bile salts and crystal violet inhibit Gram-positive bacteria.
Differential: Lactose fermenters form red/purple colonies; non-fermenters are colorless.
Indicator: Neutral red dye stains lactose fermenters.
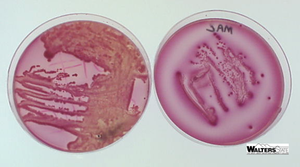
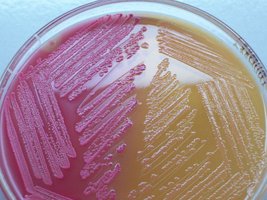
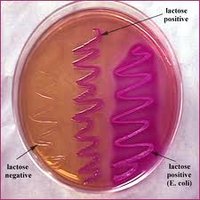
Colony Morphology on Selective Media
Lactose fermenters: Pink/red colonies on MacConkey; green sheen or dark centers on EMB.
Lactose non-fermenters: Colorless or pale colonies on both media.
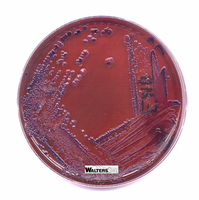
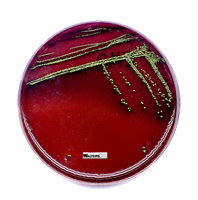
Biochemical Identification of Enteric Bacteria
Triple Sugar Iron (TSI) Agar Test
The TSI test differentiates enteric bacteria based on their ability to ferment glucose, lactose, and sucrose, produce gas, and generate hydrogen sulfide (H2S).
Indicator: Phenol red (acid production), iron compounds (H2S detection).
Interpretation:
Yellow butt/red slant: Glucose fermentation only
Yellow butt and slant: Lactose and/or sucrose fermentation
Red butt and slant: No sugar fermentation
Blackening: H2S production
Cracks/lifting: Gas production




IMViC Tests
The IMViC series (Indole, Methyl Red, Voges-Proskauer, Citrate) is used to further differentiate enteric bacteria, especially E. coli and Enterobacter species.
Indole Test: Detects indole production from tryptophan using Kovac’s reagent (red color = positive).
Methyl Red Test: Detects mixed acid fermentation (red at pH 4 = positive).
Voges-Proskauer Test: Detects butanediol fermentation (red color after reagents = positive).
Citrate Utilization: Detects ability to use citrate as sole carbon source (blue color = positive).

Pseudomonas
Background and Identification
Pseudomonas aeruginosa is a Gram-negative, oxidase-positive rod, notable for its environmental resilience and role as an opportunistic pathogen. It does not ferment glucose and is distinguished from enterics by its oxidase positivity and pigment production.
Growth Media: Pseudosel agar (PSA) and Mueller Hinton agar are used for isolation.
Pigment Production: PSA stimulates production of pyocyanin (blue-green pigment) and fluorescein (fluorescent under UV).
Colony Odor: Fruity, grape-like smell.
Key Biochemical Tests for Pseudomonas
Oxidase Test: Positive (turns black within 20–30 minutes).
Glucose Fermentation: Negative (no acid production in glucose media).
Summary Table: Key Biochemical Tests for Enterics and Pseudomonas
Test | Enterobacteriaceae | Pseudomonas aeruginosa |
|---|---|---|
Gram Reaction | Negative rods | Negative rods |
Oxidase | Negative | Positive |
Glucose Fermentation | Positive | Negative |
Lactose Fermentation | Variable (used for differentiation) | Negative |
Pigment Production | None | Pyocyanin, fluorescein |
Additional info: The Enterotube system is a rapid identification method for enterics, allowing multiple biochemical tests from a single colony. The IMViC series is especially useful for distinguishing E. coli (Indole+, Methyl Red+, VP-, Citrate-) from Enterobacter (Indole-, Methyl Red-, VP+, Citrate+).
