 Back
BackGene Regulation in Bacteria: Sigma Factors, Regulatory RNAs, Second Messengers, and Quorum Sensing
Study Guide - Smart Notes

Gene Regulation in Bacteria
Overview
Gene regulation in bacteria is a complex process that allows cells to adapt to changing environments by controlling the expression of specific genes. This regulation can occur at multiple levels, including transcriptional, post-transcriptional, and through DNA rearrangements. Key mechanisms include the use of sigma factors, regulatory RNAs, second messengers, and quorum sensing.
Sigma Factors and Regulons
Sigma Factors
Sigma factors are specialized bacterial proteins that bind to RNA polymerase and direct it to specific promoter sequences, initiating transcription of particular sets of genes. Alternative sigma factors allow bacteria to rapidly change gene expression in response to environmental stresses such as starvation, oxidative stress, or changes in pH.
Regulon: A set of genes and operons scattered throughout the chromosome but coordinately controlled by a single regulatory protein (repressor or activator).
Bacteria can activate multiple regulons simultaneously by regulating the synthesis or activity of a specific sigma factor.
Some sigma factors are regulated by anti-sigma factors, which inhibit their activity by direct binding rather than degradation. Anti-anti-sigma factors can neutralize anti-sigma factors, allowing sigma factors to function.
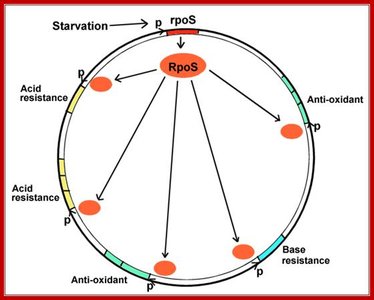
Example: The RpoS sigma factor in Escherichia coli controls genes involved in acid resistance, antioxidant defense, and base resistance during starvation.
Regulation by RNA
Riboswitches
Riboswitches are regulatory RNA sequences, typically found in the 5' untranslated region (UTR) of mRNAs, that change their structure upon binding specific metabolites. This structural change can control gene expression by affecting transcription or translation.
Riboswitches can form alternative stem-loop structures in response to ligand binding.
They can terminate transcription early or block the ribosome-binding site to prevent translation.
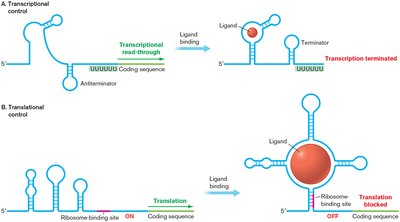
Example: A riboswitch may bind a metabolite, causing the formation of a transcription terminator or blocking translation initiation.
Untranslated Regulatory RNAs (sRNAs and asRNAs)
Small regulatory RNAs (sRNAs) and cis-antisense RNAs (asRNAs) are non-coding RNAs that regulate gene expression post-transcriptionally. sRNAs are typically 100–200 nucleotides and are encoded by intergenic regions, while asRNAs are longer and transcribed from the opposite strand of protein-coding genes.
sRNAs can inhibit or activate translation, promote or prevent mRNA degradation, or alter regulatory protein activity.
asRNAs usually affect only the gene from which they are transcribed, influencing transcription, translation, or mRNA stability.
These RNAs are energetically efficient and allow rapid response to environmental changes.
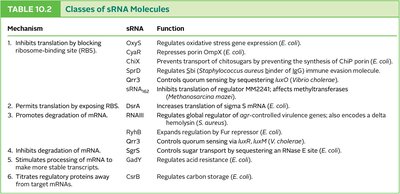
Table: Classes of sRNA molecules, their mechanisms, and functions in various bacteria.
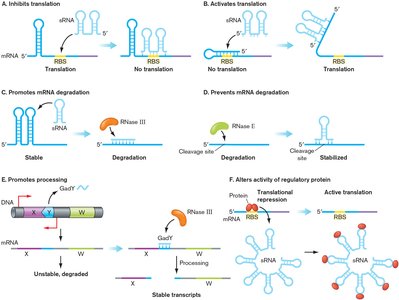
Example: The OxyS sRNA in E. coli regulates oxidative stress gene expression by blocking ribosome binding.
Second Messengers in Gene Regulation
cAMP and the Lactose Operon
Second messengers are small molecules that transmit signals within the cell, often in response to environmental changes. Cyclic AMP (cAMP) is a key second messenger in bacteria, especially in the regulation of the lactose operon (lac operon).
cAMP is synthesized from ATP by adenylate cyclase, especially when the cell is starved for carbon sources.
cAMP binds to the cAMP receptor protein (CRP), forming a complex that activates transcription of the lac operon.
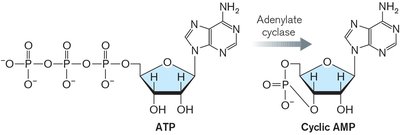
Equation:
The Lactose Operon (lac Operon)
The lac operon is a classic example of gene regulation in bacteria. It encodes proteins for lactose uptake and metabolism and is regulated by both a repressor (LacI) and an activator (cAMP-CRP complex).
In the presence of lactose, allolactose (an isomer of lactose) binds to LacI, reducing its affinity for the operator and allowing transcription.
Maximum expression requires both the removal of LacI and the presence of cAMP-CRP, which binds upstream of the promoter and enhances RNA polymerase binding.
Catabolite repression occurs when glucose is present, lowering cAMP levels and thus reducing lac operon expression even if lactose is available.
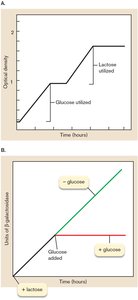
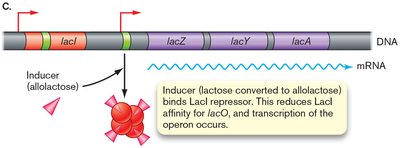
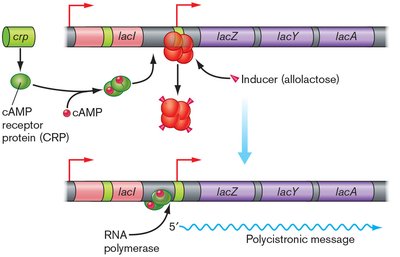
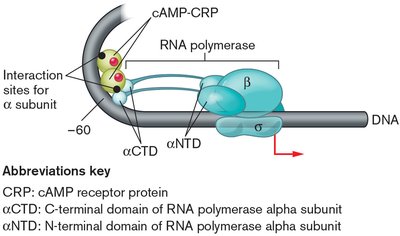

Example: When both glucose and lactose are present, E. coli will use glucose first (diauxic growth), repressing the lac operon until glucose is depleted.
Quorum Sensing and Cell-Cell Communication
Quorum Sensing
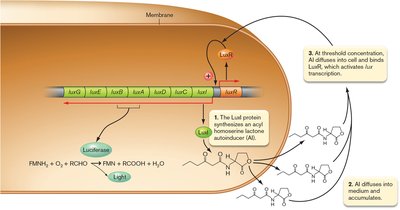
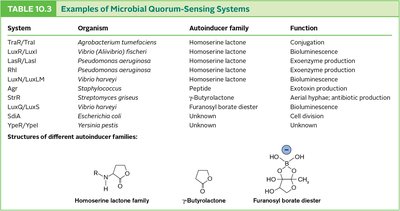
Quorum sensing is a mechanism by which bacteria coordinate gene expression based on cell density, using small signaling molecules called autoinducers. When a threshold concentration of autoinducer is reached, it triggers changes in gene expression across the population.
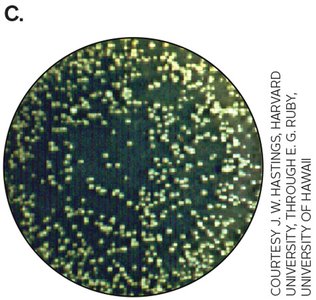
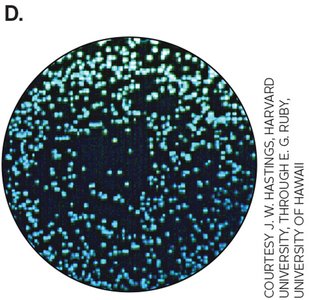
First discovered in Aliivibrio fischeri, which produces light only at high cell densities due to activation of the lux operon by autoinducer-LuxR complex.
Quorum sensing regulates diverse processes such as bioluminescence, virulence, biofilm formation, and competence for transformation.

Example: Pseudomonas aeruginosa uses quorum sensing to regulate biofilm formation and virulence factor production in cystic fibrosis patients.
Quorum Sensing and Natural Transformation
In some Gram-positive bacteria, quorum sensing regulates the development of competence, the ability to take up extracellular DNA. This process is controlled by the accumulation of a signaling peptide (CSP) that activates a phosphorylation cascade, leading to the expression of genes required for DNA uptake and integration.
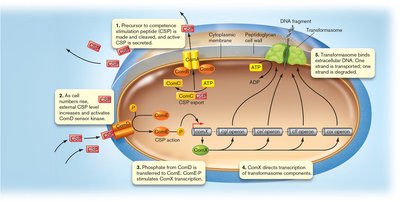
Example: In Streptococcus, CSP triggers the expression of ComX, an alternative sigma factor, which activates genes for DNA uptake.
DNA Rearrangements That Alter Gene Expression
Phase Variation
Some bacteria use DNA rearrangements to periodically change the expression of surface proteins, a process known as phase variation. This allows pathogens to evade the host immune system by altering their antigenic profile.
Gene inversion: A segment of DNA containing a promoter is flipped, changing which gene is expressed. Example: Salmonella enterica alternates between two flagellin genes.
Slipped-strand mispairing: Short sequence repeats cause DNA polymerase to slip during replication, adding or deleting repeats and altering gene expression. Example: Neisseria gonorrhoeae varies its outer membrane proteins.
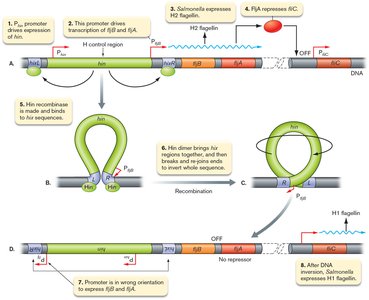
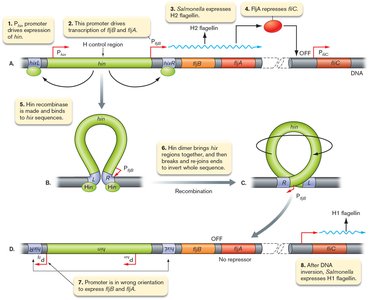
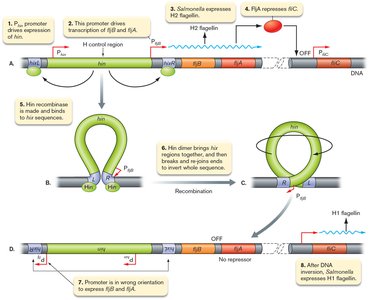
Example: Salmonella enterica uses gene inversion to switch between H1 and H2 flagellin, avoiding immune detection.
Summary Table: Key Regulatory Mechanisms
Mechanism | Key Feature | Example |
|---|---|---|
Sigma factors | Direct RNA polymerase to specific promoters | RpoS in stress response |
Riboswitches | RNA structure changes in response to metabolites | Thiamine pyrophosphate riboswitch |
sRNAs/asRNAs | Post-transcriptional regulation of mRNA | OxyS, RyhB in E. coli |
Second messengers | Small molecules modulate regulatory proteins | cAMP in lac operon |
Quorum sensing | Cell-density dependent gene regulation | Lux system in Vibrio fischeri |
DNA rearrangement | Phase variation by gene inversion or mispairing | Flagellin switching in Salmonella |
