 Back
BackIsolation and Identification of Enterobacteriaceae and Pseudomonas: Laboratory Methods and Clinical Relevance
Study Guide - Smart Notes

Enterobacteriaceae
Background
Members of the family Enterobacteriaceae are Gram-negative rods commonly found as normal flora in the intestines of humans and animals. They are classified based on their ability to ferment lactose and glucose, which is important for their identification in clinical and environmental samples.
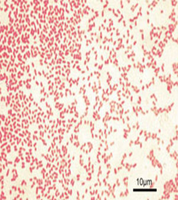
Gram-negative rods: Appear pink/red after Gram staining due to their thin peptidoglycan layer.
Normal flora: Includes Escherichia coli (lactose positive).
Human pathogens: Includes Salmonella spp., Shigella spp., and Vibrio cholerae (lactose negative).
Fermentation: All enterics ferment glucose; some ferment lactose.
Isolation of Enteric Bacteria
Selective and Differential Media
Selective and differential media are used to isolate and differentiate enteric bacteria based on their growth characteristics and metabolic properties.
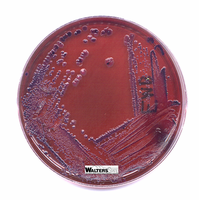
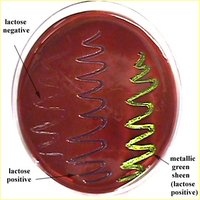
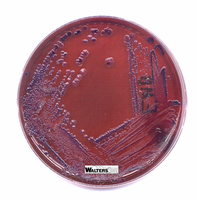
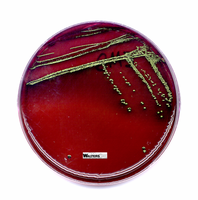
EMB (Eosin Methylene Blue) Agar: Selective for Gram-negative rods; differential for lactose fermentation. Lactose fermenters produce dark colonies with a metallic sheen, while non-fermenters remain colorless or pale.
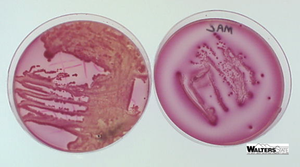
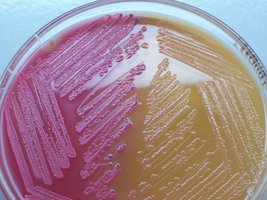
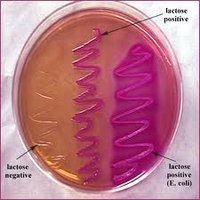
MacConkey Agar: Selective for Gram-negative rods due to bile salts and crystal violet; differential for lactose fermentation. Lactose fermenters appear pink/red, non-fermenters are colorless.
Interpretation of Results
Lactose fermenters: E. coli and Enterobacter produce colored colonies on EMB and MacConkey agar.
Lactose non-fermenters: Salmonella, Shigella, and Vibrio produce colorless colonies.
Identification of Enteric Bacteria
Triple Sugar Iron (TSI) Agar Test
The TSI test differentiates enteric bacteria based on their ability to ferment glucose, lactose, and sucrose, produce gas, and generate hydrogen sulfide (H2S).
Indicator: Phenol red detects acid production.
Gas production: Detected by splitting of agar.
H2S production: Detected by blackening of the medium.
Possible TSI reactions:
Yellow butt/red slant: Glucose fermentation only.
Yellow butt/yellow slant: Lactose and/or sucrose fermentation.
Red butt/red slant: No sugar fermentation.
Black precipitate: H2S production.
Cracks or bubbles: Gas production.




IMViC Tests
The IMViC series (Indole, Methyl Red, Voges-Proskauer, Citrate) is used to further differentiate enteric bacteria, especially within the coliform group.
Indole Test: Detects tryptophanase activity (red color after Kovac's reagent indicates positive).
Methyl Red Test: Detects mixed acid fermentation (red color is positive).
Voges-Proskauer Test: Detects butanediol fermentation (red color is positive).
Citrate Utilization Test: Detects ability to use citrate as sole carbon source (blue color is positive).

Enterotube System
The Enterotube is a rapid, multi-test system for identifying enteric bacteria. It contains 12 different media and allows for 15 biochemical tests from a single colony.
Example: Used in clinical labs for quick identification of unknown Gram-negative rods.
Pseudomonas
Background
Pseudomonas species are Gram-negative rods, primarily free-living in soil and water. They are notable for their metabolic diversity and resistance to many antibiotics and disinfectants.
No glucose fermentation: Distinguishes them from Enterobacteriaceae.
Oxidase positive: Key diagnostic feature.
P. aeruginosa: Major opportunistic pathogen, produces pigments (pyocyanin, fluorescein).
Identification of Pseudomonas aeruginosa
Oxidase test: Turns black within 20–30 minutes if positive.
Growth on Pseudosel agar (PSA): Selective for P. aeruginosa, stimulates pigment production.
Mueller Hinton agar: Selective for P. aeruginosa, colonies appear green.
Characteristic odor: Fruity, grape-like smell.
Waterborne Pathogens and Coliforms
Diseases from Contaminated Water
Typhoid Fever: Caused by Salmonella typhi, spread via feces; symptoms include high fever, diarrhea, headache.
Cholera: Caused by Vibrio cholerae, produces exotoxin; symptoms include rice-water stool, severe dehydration.
Bacillary Dysentery (Shigellosis): Caused by Shigella spp., spread person-to-person; symptoms include multiple bowel movements, fever, cramps.
Coliforms as Indicators
Coliforms are Gram-negative, non-endospore forming, lactose-fermenting bacilli used as indicators of fecal contamination in water.
Include Citrobacter, Enterobacter, Hafnia, Klebsiella, and Escherichia.
Ferment lactose to produce acid and gas.
Survive outside the body, making them reliable indicators.
Water Testing Methods
Most Probable Number (MPN) Testing
The MPN method estimates the number of coliform bacteria in a water sample through a series of fermentation tests.
Presumptive Test: Inoculate lactose broth tubes with water samples; look for acid and gas production.
Confirmed Test: Streak positive tubes onto EMB agar; confirm Gram-negative, lactose-fermenting colonies.
Completed Test: Further confirm identity by Gram stain and additional biochemical tests.
Membrane Filter Technique
This technique provides a direct count of bacteria in water. The sample is filtered, and the membrane is placed on differential media to count colonies.
Allows for direct enumeration of live bacteria.
Commonly used for routine water quality monitoring.
Summary Table: Key Biochemical Tests for Enteric Bacteria
Test | Purpose | Positive Result | Negative Result |
|---|---|---|---|
EMB Agar | Lactose fermentation | Dark colonies, metallic sheen | Colorless colonies |
MacConkey Agar | Lactose fermentation | Pink/red colonies | Colorless colonies |
TSI Agar | Sugar fermentation, H2S, gas | Yellow (acid), black (H2S), cracks (gas) | Red (alkaline), no black, no cracks |
Indole | Tryptophanase activity | Red ring | No color change |
Methyl Red | Mixed acid fermentation | Red color | Yellow color |
Voges-Proskauer | Butanediol fermentation | Red color | No color change |
Citrate | Citrate utilization | Blue color | Green color |
Key Equations and Concepts
Fermentation equation (generalized):
Most Probable Number (MPN): Statistical estimation based on number of positive tubes at each dilution.
Additional info: The IMViC tests are especially useful for distinguishing E. coli (Indole+, Methyl Red+, VP-, Citrate-) from Enterobacter (Indole-, Methyl Red-, VP+, Citrate+).
