 Back
BackMicrobial Cell Structure and Function: Study Guide
Study Guide - Smart Notes

Microbial Cell Structure and Function
Overview of Microbial Cell Structures and Functions
Microbial cells possess a variety of structural features that enable them to interact with their environment, store nutrients, move, and survive harsh conditions. Understanding these structures is fundamental to microbiology.
Cell Envelope/Membrane: Phospholipid bilayers that separate the cytoplasm from the environment and regulate transport.
Cell Walls: Peptidoglycan provides rigidity; thickness varies between Gram-positive (thick) and Gram-negative (thin).
Capsules: Polysaccharide layers that enhance virulence by preventing phagocytosis.
Fimbriae: Protein structures for attachment to surfaces.
Pili: Facilitate conjugation and genetic exchange.
Flagella: Enable motility.
Inclusions: Storage of energy and nutrients.
Endospores: Survival structures resistant to extreme conditions.
Cell Envelope Structure
The cell envelope consists of several layers that protect the cell and mediate interactions with the environment. The composition varies between different microbial groups.
Cytoplasmic Membrane: Present in all cells; composed of a phospholipid bilayer.
Cell Wall: Most bacteria have a cell wall; provides structural support.
Outer Membrane: Found only in Gram-negative bacteria.
S-layers: Protein or glycoprotein layers found in some bacteria and archaea.
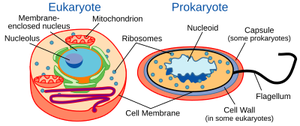
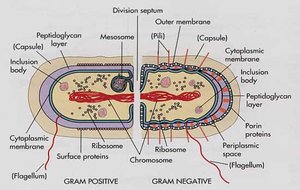
Gram-Positive and Gram-Negative Cell Envelopes
Bacteria are classified based on their cell wall structure, which affects their staining properties and susceptibility to antibiotics.
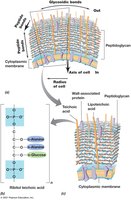
Gram-Positive: Thick peptidoglycan layer, no outer membrane.
Gram-Negative: Thin peptidoglycan layer, outer membrane present.
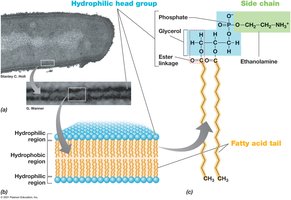
Cytoplasmic Membrane Structure and Function
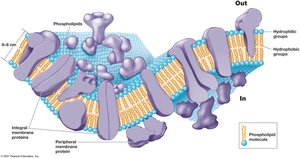
The cytoplasmic membrane is a selectively permeable barrier composed of a phospholipid bilayer with embedded proteins. It is essential for nutrient transport, waste removal, and energy metabolism.
Phospholipid Bilayer: Hydrophobic fatty acid tails face inward; hydrophilic head groups face outward.
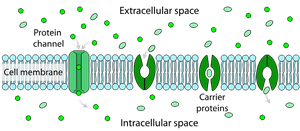
Membrane Proteins: Integral, transmembrane, and peripheral proteins facilitate transport and metabolic reactions.
Selective Permeability: Allows specific molecules to enter or exit the cell.
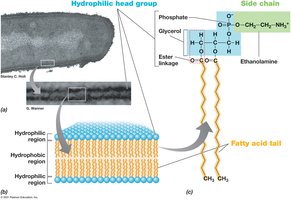
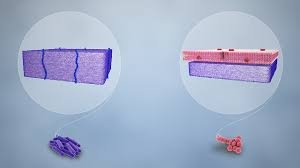
Variations in Cytoplasmic Membranes Across Domains
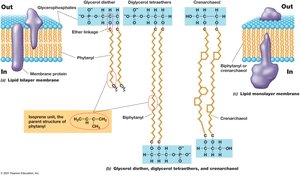
Membrane composition differs between Bacteria, Archaea, and Eukarya, affecting their stability and function.
Bacteria/Eukarya: Ester linkages in phospholipids; fatty acids as side chains.
Archaea: Ether linkages; isoprene-based side chains; can form monolayers for increased stability.
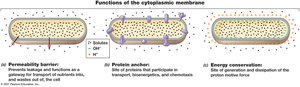
Functions of the Cytoplasmic Membrane
The cytoplasmic membrane serves as a permeability barrier, protein anchor, and site of energy conservation.
Permeability Barrier: Prevents leakage and controls entry/exit of substances.
Protein Anchor: Holds proteins involved in transport and chemotaxis.
Energy Conservation: Site of proton motive force generation and dissipation.
Bacterial Cell Walls: Peptidoglycan Structure
Peptidoglycan is a rigid polysaccharide layer providing strength to bacterial cell walls. Its structure and cross-linking differ between Gram-positive and Gram-negative bacteria.
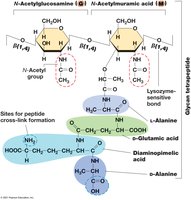
Glycan Tetrapeptide: Alternating N-acetylglucosamine and N-acetylmuramic acid joined by β-1,4 linkages.
Peptide Cross-links: Amino acids vary; cross-linking provides rigidity.
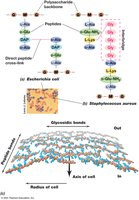
Gram-Negative: Single layer, direct cross-links.
Gram-Positive: Multiple layers, peptide interbridges (e.g., five glycines in Staphylococcus aureus).
Archaeal Cell Walls
Archaeal cell walls lack peptidoglycan and outer membranes. Some have S-layers, while methanogens possess pseudomurein, which is structurally similar to peptidoglycan but resistant to lysozyme and penicillin.
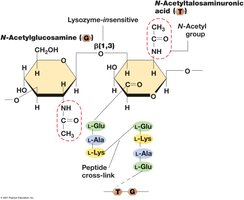
Pseudomurein: Alternating N-acetylglucosamine and N-acetyltalosaminuronic acid joined by β-1,3 linkages.
S-layers: Protein shells providing structural support.
Gram-Negative Outer Membrane and Lipopolysaccharide (LPS)
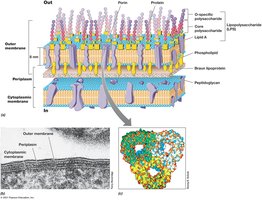
The outer membrane of Gram-negative bacteria contains lipopolysaccharide (LPS), which is important for surface recognition, virulence, and structural integrity.
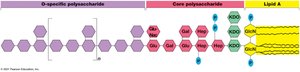
LPS Structure: Composed of O-polysaccharide, core polysaccharide, and lipid A (endotoxin).
Porins: Protein channels for solute transport.
Periplasm: Space between cytoplasmic and outer membranes, containing extracellular proteins.
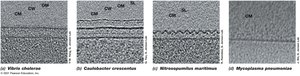
Diversity of Cell Envelope Structure: S-Layers and Alternative Structures
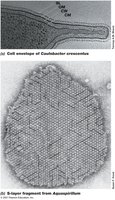
S-layers are paracrystalline protein or glycoprotein layers found in some Bacteria and Archaea. Some microbes lack cell walls but have tough membranes containing sterols.
S-layers: Provide strength, shape, and protection.
Alternative Structures: Mycoplasma (Bacteria) and Thermoplasma (Archaea) lack cell walls.

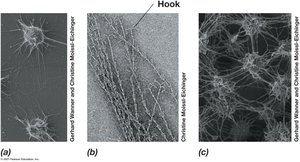
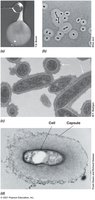
Cell Surface Structures: Capsules, Slime Layers, Pili, and Fimbriae
Microbial cells may possess additional surface structures that aid in attachment, biofilm formation, infectivity, and genetic exchange.
Capsules: Tightly attached polysaccharide layers; visible with India ink.
Slime Layers: Loosely attached, easily deformed.
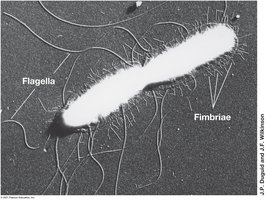
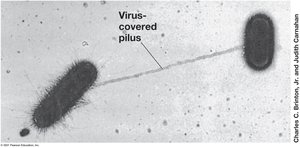
Pili: Thin protein filaments for attachment, conjugation, and motility.
Fimbriae: Short pili for attachment; produced by many bacteria.
Hami: Archaeal grappling hooks for biofilm formation.

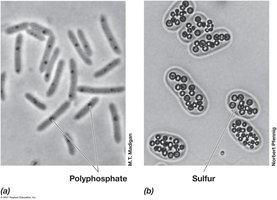
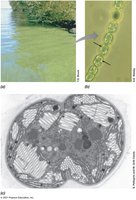
Cell Inclusions: Storage and Special Functions
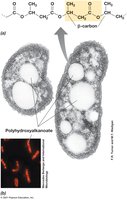
Cell inclusions serve as energy reserves, carbon or phosphorus reservoirs, and confer special functions such as buoyancy and magnetotaxis.
Carbon Storage Polymers: Poly-β-hydroxybutyric acid (PHB), poly-β-hydroxyalkanoate (PHA), and glycogen.
Polyphosphate Granules: Inorganic phosphate storage.
Sulfur Granules: Elemental sulfur storage.
Carbonate Minerals: Biomineralization of barium, strontium, magnesium.
Gas Vesicles: Provide buoyancy.
Magnetosomes: Magnetic iron oxides for orientation in magnetic fields.
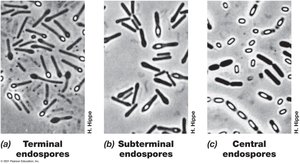
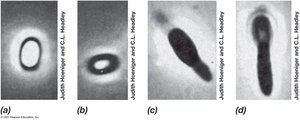
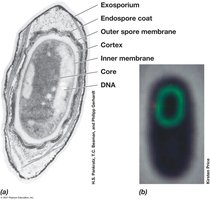
Endospores: Survival Structures
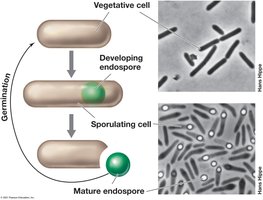
Endospores are highly differentiated, dormant cells formed by some Gram-positive bacteria. They are resistant to heat, radiation, chemicals, and desiccation, enabling survival in harsh environments.
Formation: Sporulation occurs in response to nutrient limitation.
Structure: Contains dipicolinic acid, Ca2+, and small acid-soluble spore proteins (SASPs).
Germination: Activation, germination, and outgrowth upon nutrient availability.
Flagella, Archaella, and Swimming Motility
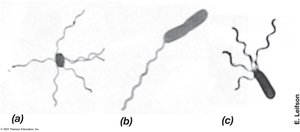
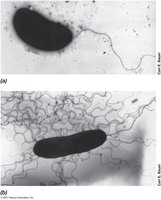
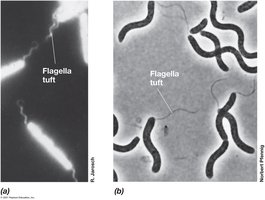
Flagella and archaella are motility structures that enable swimming in liquid environments. Their arrangement and synthesis are genetically controlled.
Flagella: Long, thin appendages; arrangements include polar, lophotrichous, amphitrichous, and peritrichous.
Motility: Rotational speed depends on proton motive force.
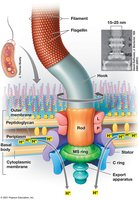
Synthesis: Filament grows from tip; MS ring made first.
Surface Motility: Twitching and Gliding
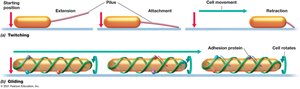
Some bacteria move across surfaces using twitching or gliding motility, which is slower than swimming.
Twitching Motility: Pili extend, attach, and retract to pull the cell forward; powered by ATP hydrolysis.
Gliding Motility: Smooth, continuous motion along the cell axis; involves intracellular protein tracks and adhesion proteins.

Chemotaxis and Other Forms of Taxis
Taxis refers to directed movement in response to environmental stimuli. Chemotaxis is movement toward or away from chemicals, while other forms include phototaxis, aerotaxis, osmotaxis, and hydrotaxis.
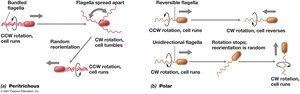
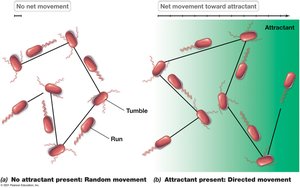
Run and Tumble: Peritrichous bacteria alternate between smooth runs and tumbles; direction is biased by attractant concentration.
Polarly Flagellated Bacteria: May reverse flagellar rotation or rely on Brownian motion.
Measuring Chemotaxis: Capillary tube assays quantify movement toward attractants or repellents.
Other Taxis: Aerotaxis (oxygen), phototaxis (light), osmotaxis (ionic strength), hydrotaxis (water), magnetotaxis (magnetic fields).
Endosymbiotic Origin of Organelles
The endosymbiotic hypothesis proposes that mitochondria and chloroplasts originated from bacterial cells that formed symbiotic relationships with early eukaryotes. Evidence includes the presence of circular DNA and bacterial-like ribosomes in these organelles.
Mitochondria: Descended from respiratory bacteria.
Chloroplasts: Descended from phototrophic bacteria.
Eukarya: Originated from symbiotic fusion of archaeal host and mitochondrial endosymbiont.
