 Back
BackMicrobial Genetics and DNA Replication: Core Concepts and Mechanisms
Study Guide - Smart Notes

Microbial Genetics and DNA Replication
Serial Dilution and Viable Plate Counts
Serial dilution is a fundamental microbiological technique used to estimate the concentration of viable microorganisms in a sample. It involves stepwise dilution of a bacterial culture to obtain countable colonies on agar plates.
Serial Dilution: A process where a sample is diluted in a series of steps, typically by factors of 10, to reduce the concentration of cells for easier counting.
Viable Plate Count: After dilution, aliquots are plated on agar and incubated. Colonies that form are counted, and the original cell concentration is calculated.
Applications: Used in food safety, water quality testing, and research to quantify microbial populations.
Example: 1 ml of bacteria added to 9 ml broth yields a 1:10 (10-1) dilution; further dilutions can reach 1:100,000 or more.


Microbial Genetics: Key Concepts
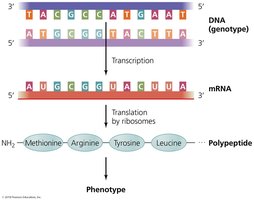
Genotype and Phenotype
Genetics is the study of heredity and variation in organisms. In microbiology, understanding genotype and phenotype is crucial for manipulating and studying microorganisms.
Genotype: The genetic makeup (DNA sequence) of an organism.
Phenotype: The observable physical or biochemical characteristics determined by the genotype.
Example: A DNA sequence encoding antibiotic resistance (genotype) results in a bacterium surviving antibiotic treatment (phenotype).

Importance of Microbial Genetics
Microbial genetics enables the manipulation of microorganisms for beneficial purposes and helps explain the genetic basis of diseases.
Applications: Production of insulin, vaccines (e.g., Hepatitis B), and understanding disease mechanisms.
Ease of Study: Microbial genomes are simpler and more accessible than those of higher organisms.
The Language of Genetics
Gene: A sequence of nucleotides encoding a protein.
Gene Locus: The specific location of a gene on a chromosome.
Genome: The complete set of genetic material in an organism.
Genetic Code: The set of rules by which information encoded in DNA is translated into proteins.
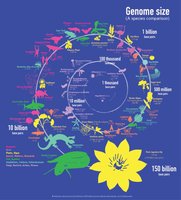
Genome Size and Complexity
Genome size does not always correlate with organism complexity. Microbial genomes are typically smaller than those of multicellular organisms.
Examples: E. coli (~4,288 genes), humans (~25,000 genes), corn (~50,000 genes).
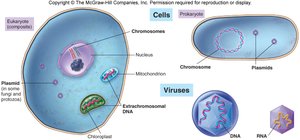
Genetic Material Organization
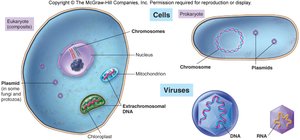
Genetic material is packaged differently in prokaryotes, eukaryotes, and viruses.
Prokaryotes: Circular chromosomes, plasmids, no nucleus.
Eukaryotes: Linear chromosomes in a nucleus, extranuclear DNA in mitochondria/chloroplasts.
Viruses: DNA or RNA, single or double-stranded.
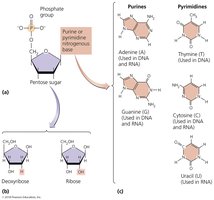
Nucleic Acids: Structure and Function
Nucleotides and Nucleic Acids
Nucleic acids (DNA and RNA) are polymers of nucleotides, each consisting of a phosphate group, a pentose sugar, and a nitrogenous base.
DNA: Contains deoxyribose sugar; bases are adenine (A), thymine (T), cytosine (C), guanine (G).
RNA: Contains ribose sugar; uracil (U) replaces thymine.
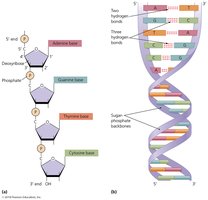
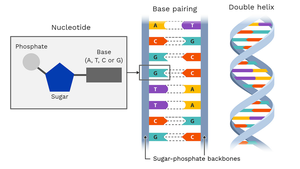
DNA Double Helix and Base Pairing
DNA is typically double-stranded, with complementary base pairing and antiparallel orientation.
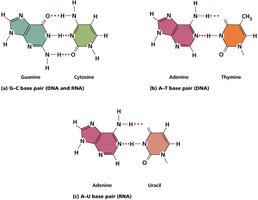
Base Pairing: A pairs with T (2 hydrogen bonds), C pairs with G (3 hydrogen bonds).
Antiparallel Strands: One strand runs 5' to 3', the other 3' to 5'.
Stability: More C-G pairs increase DNA stability due to extra hydrogen bonds.
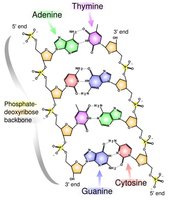
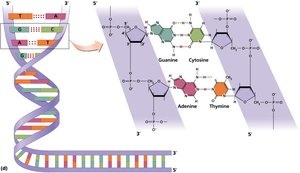
Sugar-Phosphate Backbone
The backbone of DNA is formed by covalent bonds between the 5' and 3' carbons of the sugar molecules, creating a stable structure.
Covalent Bonds: Link the sugar of one nucleotide to the phosphate of the next.
Antiparallel Arrangement: Essential for replication and function.
Prokaryotic and Eukaryotic Genomes
Prokaryotic Genomes
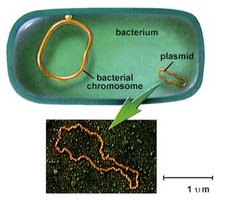
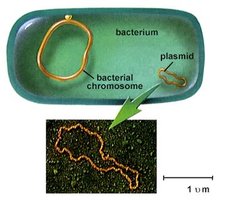
Prokaryotes (bacteria) have a single, circular chromosome located in the nucleoid region, and often contain plasmids.
Nucleoid: Region in the cytoplasm where DNA is concentrated; not membrane-bound.
Plasmids: Small, circular DNA molecules that replicate independently and can carry advantageous genes (e.g., antibiotic resistance).
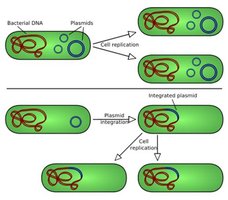
Plasmid Functions and Manipulation
Fertility (F) Plasmids: Enable conjugation (gene transfer between bacteria).
Resistance (R) Plasmids: Carry antibiotic resistance genes.
Bacteriocin (Col) Plasmids: Encode proteins that kill other bacteria.
Virulence Plasmids: Carry genes that enhance pathogenicity.
Genetic Engineering: Plasmids can be manipulated to alter bacterial phenotypes for research or biotechnology.

Eukaryotic Genomes
Nuclear DNA: Located in the nucleus, organized into linear chromosomes.
Extranuclear DNA: Found in mitochondria and chloroplasts, inherited maternally.
DNA Replication
Key Features of DNA Replication
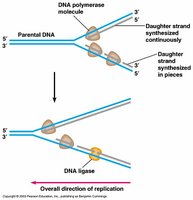
DNA replication is an anabolic, semiconservative process requiring energy and enzymes.
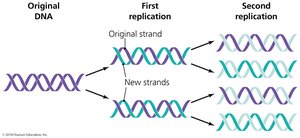
Semiconservative Replication: Each new DNA molecule consists of one original (parental) and one new (daughter) strand.
Enzymes: DNA polymerases, helicase, primase, ligase, and others are involved.
Directionality: DNA polymerase synthesizes new DNA only in the 5' to 3' direction.
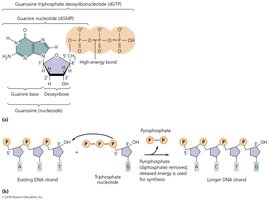
Steps in DNA Replication
Initiation: DNA helicase unwinds the double helix at the origin of replication, creating a replication fork.
Elongation: DNA polymerase adds nucleotides to the growing DNA strand, using the original strand as a template.
Termination: Replication ends when the forks meet, and ligase joins DNA fragments.
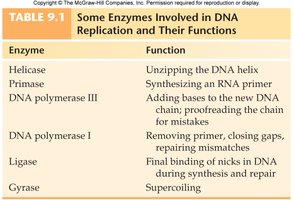

Enzymes Involved in DNA Replication
Enzyme | Function |
|---|---|
Helicase | Unzipping the DNA helix |
Primase | Synthesizing an RNA primer |
DNA polymerase III | Adding bases to the new DNA chain; proofreading |
DNA polymerase I | Removing primer, closing gaps, repairing mismatches |
Ligase | Final binding of nicks in DNA during synthesis and repair |
Gyrase | Supercoiling |
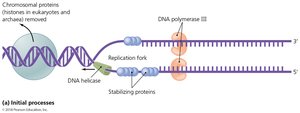
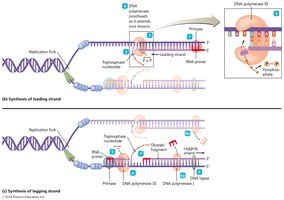
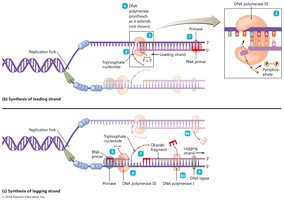
Leading and Lagging Strand Synthesis
Leading Strand: Synthesized continuously in the 5' to 3' direction.
Lagging Strand: Synthesized discontinuously as Okazaki fragments, later joined by DNA ligase.
Primase: Synthesizes short RNA primers to initiate DNA synthesis.
Central Dogma of Biology
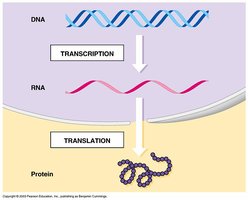
From DNA to Protein
The central dogma describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein.
Transcription: Synthesis of RNA from a DNA template.
Translation: Synthesis of proteins using mRNA as a template.
Transcription in Prokaryotes and Eukaryotes
Prokaryotes: Transcription occurs in the nucleoid; genes often organized in operons.
Eukaryotes: Transcription occurs in the nucleus, mitochondria, and chloroplasts; genes are not grouped in operons.
Types of RNA: mRNA (messenger), rRNA (ribosomal), tRNA (transfer).
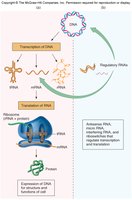
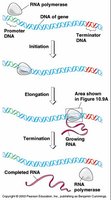
Steps of Transcription
Initiation: RNA polymerase binds to promoter regions, aided by transcription factors.
Elongation: RNA polymerase synthesizes RNA in the 5' to 3' direction.
Termination: RNA transcript is released, and polymerase detaches from DNA.
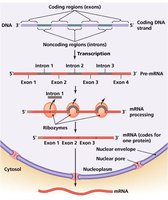
Gene Structure in Eukaryotes: Introns and Exons
Introns: Non-coding sequences removed during RNA processing.
Exons: Coding sequences that are spliced together to form mature mRNA.
Splicing: Performed by spliceosomes, which recognize exon-intron boundaries.
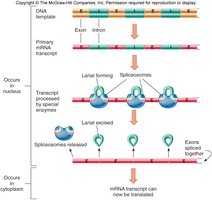
Gene Expression and Cell Differentiation
Although all cells in an organism have the same genetic code, gene expression varies, leading to different cell types and functions.
*Additional info: The notes above provide a comprehensive overview of microbial genetics, DNA structure, replication, and the central dogma, with emphasis on prokaryotic systems relevant to microbiology.*
