 Back
BackMicrobial Genetics: Structure, Replication, Expression, and Regulation
Study Guide - Smart Notes

Microbial Genetics
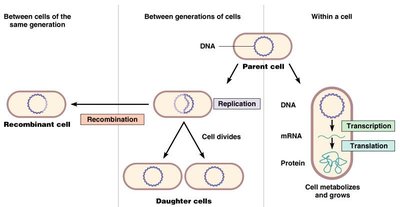
Flow of Genetic Information
Microbial genetics explores how genetic information is stored, replicated, expressed, and regulated in microorganisms. The flow of genetic information involves the processes of DNA replication, transcription, and translation, as well as genetic recombination between cells.
Replication: The process by which DNA is copied to produce identical daughter cells during cell division.
Transcription: The synthesis of RNA from a DNA template, resulting in messenger RNA (mRNA).
Translation: The process by which ribosomes synthesize proteins using the sequence encoded in mRNA.
Recombination: The exchange of genetic material between different cells, leading to genetic diversity.
Structure of DNA
Nucleotides and DNA Double Helix
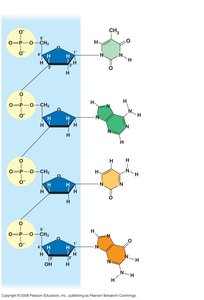
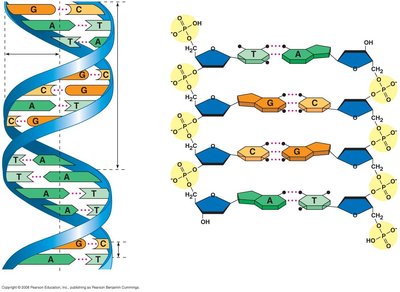
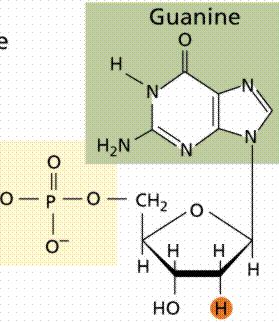
DNA is a polymer composed of nucleotides, each consisting of a deoxyribose sugar, a phosphate group, and a nitrogenous base. The sequence of these bases encodes genetic information.
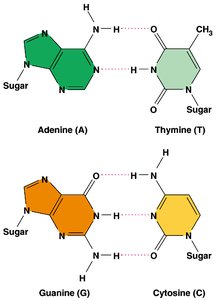
Nitrogenous Bases: Two types—purines (adenine, guanine) and pyrimidines (cytosine, thymine).
Sugar-Phosphate Backbone: The backbone of DNA is formed by alternating sugars and phosphates.
Base Pairing: Adenine pairs with thymine (A=T) via two hydrogen bonds; guanine pairs with cytosine (G≡C) via three hydrogen bonds.
Antiparallel Strands: The two DNA strands run in opposite directions (5' to 3' and 3' to 5').
DNA Replication
Mechanism and Enzymes
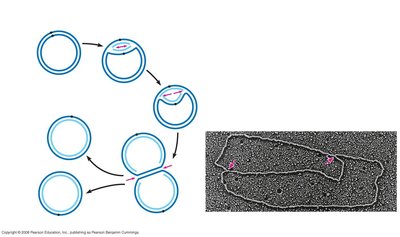
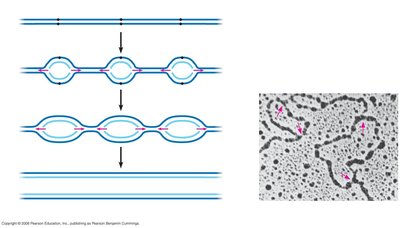
DNA replication is semiconservative, meaning each new DNA molecule consists of one parental and one newly synthesized strand. Replication begins at specific origins and proceeds bidirectionally.
Origins of Replication: Specific sequences where DNA unwinding and synthesis begin.
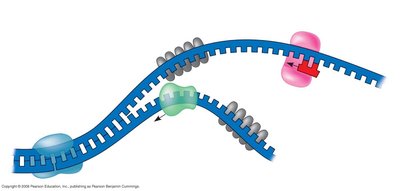
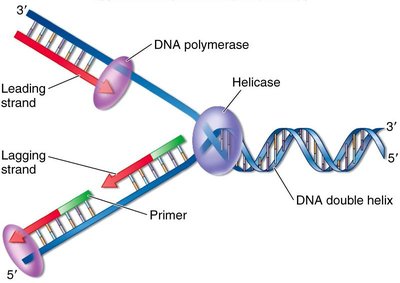
Replication Fork: The Y-shaped region where new DNA strands are synthesized.
Key Enzymes:
Helicase: Unwinds the DNA double helix.
Single-Strand Binding Proteins: Stabilize unwound DNA.
Topoisomerase: Relieves supercoiling ahead of the fork.
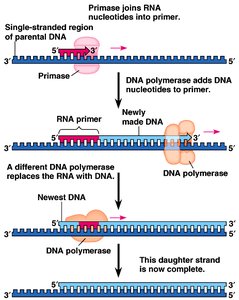
Primase: Synthesizes short RNA primers.
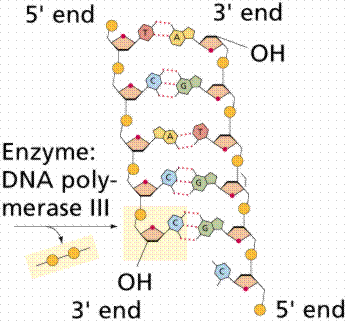
DNA Polymerase: Adds nucleotides to the 3' end of the primer; synthesizes new DNA in the 5' to 3' direction.
Leading and Lagging Strands: The leading strand is synthesized continuously; the lagging strand is synthesized in Okazaki fragments.

Gene Expression: Transcription and Translation
Transcription
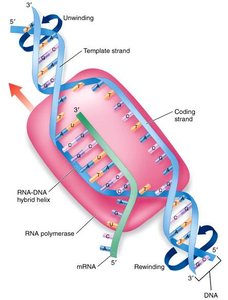
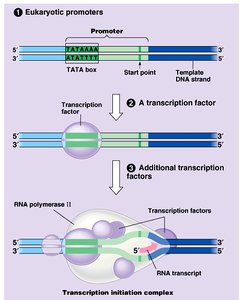
Transcription is the process by which RNA is synthesized from a DNA template. In prokaryotes, RNA polymerase binds directly to the promoter(Pribnow box, TATAAT); in eukaryotes, transcription factors recognize the promoter region (TATA box).
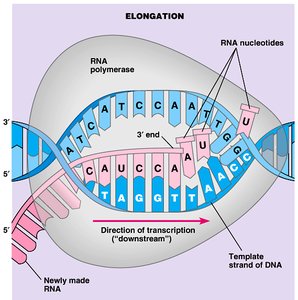
Stages of Transcription: Initiation, elongation, and termination.
RNA Polymerase: Synthesizes RNA in the 5' to 3' direction, reading the DNA template 3' to 5'.
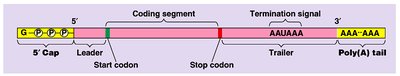
mRNA Processing (Eukaryotes): Addition of a 5' cap, poly-A tail, and splicing to remove introns.
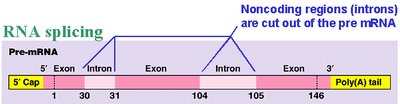
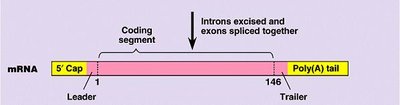
Translation
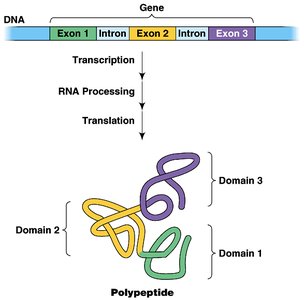
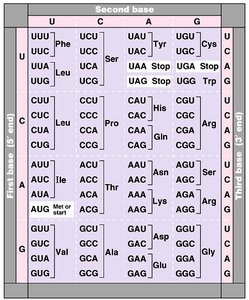
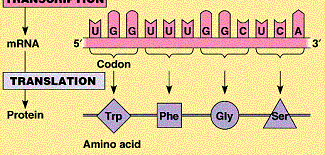
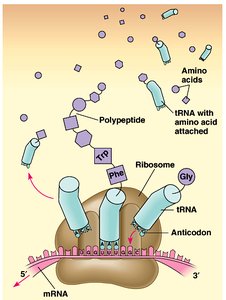
Translation is the synthesis of proteins from mRNA by ribosomes. The genetic code is read in sets of three nucleotides (codons), each specifying an amino acid.
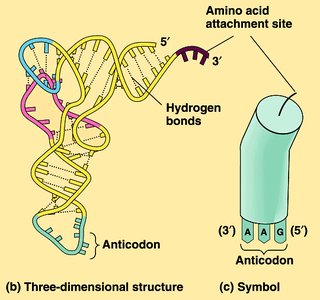
tRNA: Transfer RNA molecules bring amino acids to the ribosome, matching codons with anticodons.
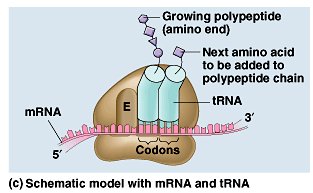
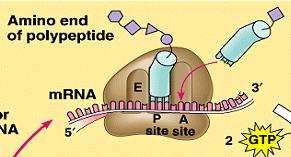
Ribosome Structure: Ribosomes have binding sites for mRNA and tRNAs (A, P, and E sites).
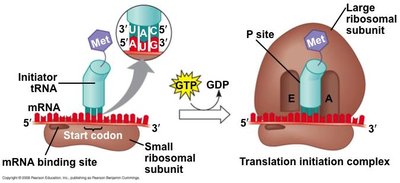
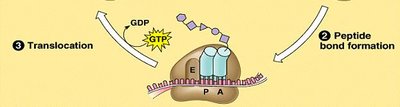
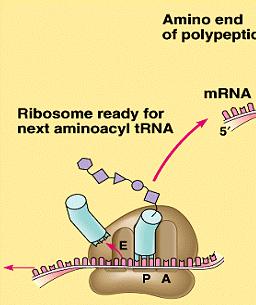
Stages of Translation: Initiation (assembly of the ribosome on mRNA), elongation (addition of amino acids), and termination (release of the completed polypeptide).
Start and Stop Codons: AUG (methionine) is the start codon; UAA, UAG, and UGA are stop codons.
Mutations and Genetic Variation
Types of Mutations
Mutations are changes in the DNA sequence that can affect protein structure and function. They are a source of genetic variation in microbial populations.
Point Mutations: Change in a single nucleotide pair.
Silent Mutation: No effect on amino acid sequence.
Missense Mutation: Changes one amino acid in the protein.
Nonsense Mutation: Converts a codon to a stop codon, truncating the protein.
Frameshift Mutations: Insertions or deletions that alter the reading frame, often resulting in nonfunctional proteins.
Mutagens: Physical or chemical agents that increase mutation rates.
Genetic Recombination in Bacteria
Mechanisms of Recombination
Genetic recombination increases genetic diversity in bacteria and can occur through several mechanisms:
Transformation: Uptake of naked DNA from the environment.
Conjugation: Direct transfer of DNA between bacteria via pili (F factor or plasmids).
Transduction: Transfer of bacterial genes by bacteriophages (viruses).
Transposons: Mobile genetic elements that can move within and between DNA molecules.
Regulation of Bacterial Gene Expression
Operon Model and Gene Regulation
Bacteria regulate gene expression to conserve energy and resources, producing proteins only when needed. The operon model describes how groups of genes are regulated together.
Constitutive Genes: Expressed continuously (e.g., genes for essential metabolic enzymes).
Regulated Genes: Expressed only under certain conditions (e.g., lac operon for lactose metabolism).
Repressible Operons: Usually on; can be turned off by a repressor (e.g., trp operon).
Inducible Operons: Usually off; can be turned on by an inducer (e.g., lac operon).
Positive Regulation: Activator proteins (e.g., CAP) enhance transcription in response to environmental signals.
Example: The lac operon is induced in the presence of lactose and low glucose, allowing bacteria to metabolize lactose only when it is available and preferred carbon sources are absent.
Additional info: This guide covers the core concepts of microbial genetics, including DNA structure, replication, gene expression, mutation, recombination, and gene regulation, as relevant to a college-level microbiology course.
